
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 1,122,293 – 1,122,389 |
| Length | 96 |
| Max. P | 0.999916 |

| Location | 1,122,293 – 1,122,389 |
|---|---|
| Length | 96 |
| Sequences | 5 |
| Columns | 102 |
| Reading direction | forward |
| Mean pairwise identity | 88.55 |
| Mean single sequence MFE | -26.68 |
| Consensus MFE | -26.22 |
| Energy contribution | -25.94 |
| Covariance contribution | -0.28 |
| Combinations/Pair | 1.19 |
| Mean z-score | -2.67 |
| Structure conservation index | 0.98 |
| SVM decision value | 4.53 |
| SVM RNA-class probability | 0.999916 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 1122293 96 + 22224390 UGGGUGACUAAUAGAAAAUCUCGCUGCUUGCUGGCGAAUUUUUCUUAGUCGAGCUUAAACGAGAAAUAUCGACGACUCCCACCCACAAUACCCAC--A---- ((((((......((((((..(((((.......)))))..))))))..(((((.(((....))).....)))))......))))))..........--.---- ( -27.30) >DroSec_CAF1 19035 93 + 1 UGGGUGACUAACAGAAAAACUCGCUGCUUGCUAGCAAAUUUUUCUUAGUCGAGCUUAAACGAGAAAUAUCGACGACUCCCACCCACAAUACCC--------- ((((((......(((((((...((((.....))))...)))))))..(((((.(((....))).....)))))......))))))........--------- ( -23.10) >DroSim_CAF1 19972 96 + 1 UGGGUGACUAAUAGAAAAACUCGCUGCUUGCUAGCGAAUUUUUCUUAGUCGAGCUUAAACGAGAAAUAUCGACGACUCCCACCCACAAUACCCAC--A---- ((((((......(((((((.((((((.....)))))).)))))))..(((((.(((....))).....)))))......))))))..........--.---- ( -28.80) >DroEre_CAF1 20129 100 + 1 UGGGUGACUAAUAGAAAAACUUGCCGCUCGCUGGCGAAUUUUUCUUAGUCGAGCUUGAACGAGAAAUAUCGACGACUCCCACCCACAAGUCCCAAACACC-- ((((((((((..(((((((.((((((.....)))))).))))))))))))(((.(((..(((......))).))))))..)))))...............-- ( -29.30) >DroYak_CAF1 19489 102 + 1 UGGGUGACUAAUGGAAAAACUUGCUGCUUGCUGGUGAAUUUGUCUUAGUCGAGCUUAAACGAGAAAUAUCGACGAUUCCCACCCACAAGUCCCAAGCAACCA (((....))).(((.....((((..(((((..((((.....(((...(((((.(((....))).....))))))))...))))..)))))..))))...))) ( -24.90) >consensus UGGGUGACUAAUAGAAAAACUCGCUGCUUGCUGGCGAAUUUUUCUUAGUCGAGCUUAAACGAGAAAUAUCGACGACUCCCACCCACAAUACCCAA__A____ ((((((......(((((((.((((((.....)))))).)))))))..(((((.(((....))).....)))))......))))))................. (-26.22 = -25.94 + -0.28)

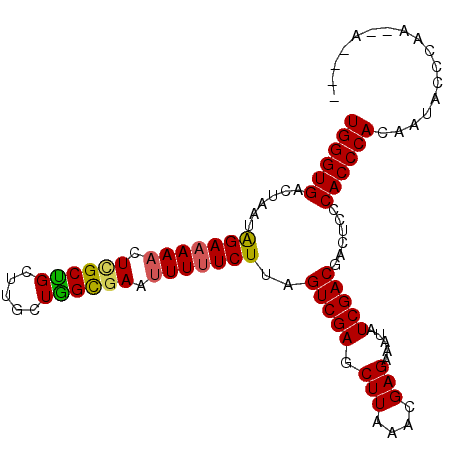
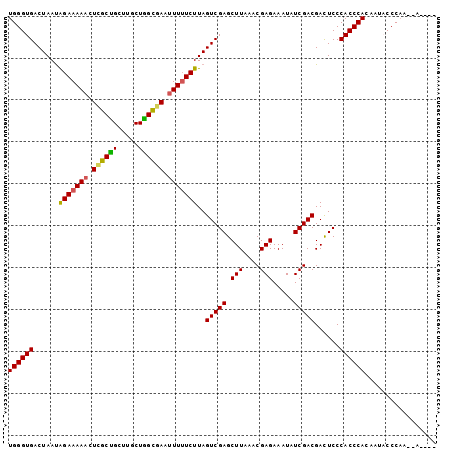
| Location | 1,122,293 – 1,122,389 |
|---|---|
| Length | 96 |
| Sequences | 5 |
| Columns | 102 |
| Reading direction | reverse |
| Mean pairwise identity | 88.55 |
| Mean single sequence MFE | -30.20 |
| Consensus MFE | -26.88 |
| Energy contribution | -27.92 |
| Covariance contribution | 1.04 |
| Combinations/Pair | 1.12 |
| Mean z-score | -1.93 |
| Structure conservation index | 0.89 |
| SVM decision value | 1.86 |
| SVM RNA-class probability | 0.980394 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 1122293 96 - 22224390 ----U--GUGGGUAUUGUGGGUGGGAGUCGUCGAUAUUUCUCGUUUAAGCUCGACUAAGAAAAAUUCGCCAGCAAGCAGCGAGAUUUUCUAUUAGUCACCCA ----.--.(((((.....((((..(((.((..((....)).)))))..))))(((((((((((.(((((..(....).))))).))))))..)))))))))) ( -28.00) >DroSec_CAF1 19035 93 - 1 ---------GGGUAUUGUGGGUGGGAGUCGUCGAUAUUUCUCGUUUAAGCUCGACUAAGAAAAAUUUGCUAGCAAGCAGCGAGUUUUUCUGUUAGUCACCCA ---------((((.....((((..(((.((..((....)).)))))..))))((((((((((((((((((.(....))))))))))))))..))))))))). ( -32.30) >DroSim_CAF1 19972 96 - 1 ----U--GUGGGUAUUGUGGGUGGGAGUCGUCGAUAUUUCUCGUUUAAGCUCGACUAAGAAAAAUUCGCUAGCAAGCAGCGAGUUUUUCUAUUAGUCACCCA ----.--.(((((.....((((..(((.((..((....)).)))))..))))((((((((((((((((((.(....))))))))))))))..)))))))))) ( -34.50) >DroEre_CAF1 20129 100 - 1 --GGUGUUUGGGACUUGUGGGUGGGAGUCGUCGAUAUUUCUCGUUCAAGCUCGACUAAGAAAAAUUCGCCAGCGAGCGGCAAGUUUUUCUAUUAGUCACCCA --......((((.....((..((((((..........))))))..)).....((((((((((((((.(((.......))).)))))))))..))))).)))) ( -31.20) >DroYak_CAF1 19489 102 - 1 UGGUUGCUUGGGACUUGUGGGUGGGAAUCGUCGAUAUUUCUCGUUUAAGCUCGACUAAGACAAAUUCACCAGCAAGCAGCAAGUUUUUCCAUUAGUCACCCA .(((.(((((((((((((.(.(((((((.(((((....))(((........)))....)))..)))).))).).....)))))))...)))..))).))).. ( -25.00) >consensus ____U__GUGGGUAUUGUGGGUGGGAGUCGUCGAUAUUUCUCGUUUAAGCUCGACUAAGAAAAAUUCGCCAGCAAGCAGCGAGUUUUUCUAUUAGUCACCCA .........(((......((((..(((.((..((....)).)))))..))))((((((((((((((((((.......)))))))))))))..))))).))). (-26.88 = -27.92 + 1.04)

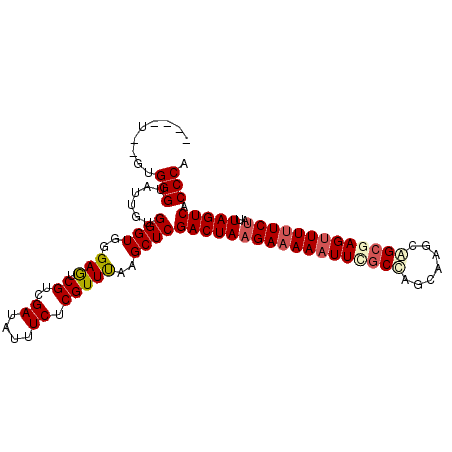
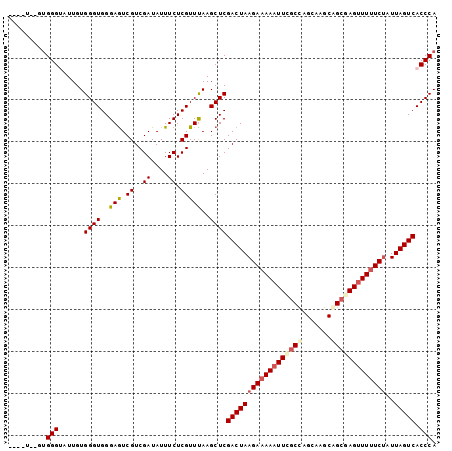
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 09:42:45 2006