
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 9,717,014 – 9,717,105 |
| Length | 91 |
| Max. P | 0.640990 |

| Location | 9,717,014 – 9,717,105 |
|---|---|
| Length | 91 |
| Sequences | 4 |
| Columns | 91 |
| Reading direction | reverse |
| Mean pairwise identity | 87.66 |
| Mean single sequence MFE | -30.53 |
| Consensus MFE | -22.98 |
| Energy contribution | -23.85 |
| Covariance contribution | 0.87 |
| Combinations/Pair | 1.12 |
| Mean z-score | -1.59 |
| Structure conservation index | 0.75 |
| SVM decision value | 0.22 |
| SVM RNA-class probability | 0.640990 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
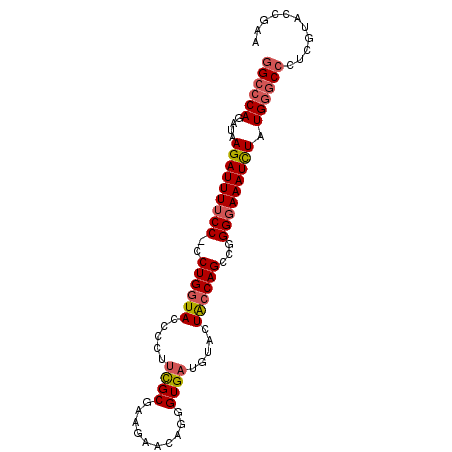
>X_DroMel_CAF1 9717014 91 - 22224390 GGCCCAGAUAAGAUUUUCCCCCUGGUACCCCUUUGCGCAGAACAGGGUGAUGUACUACCAGCCGGGGAAAUCUAUGGCCCCUCGUACCGAU ((((......((((((((((.((((((.....(..(.(......).)..).....))))))..))))))))))..))))............ ( -29.60) >DroSec_CAF1 5056 90 - 1 GGCCCAGAUAAGAUUUUCC-CCUGGUACCCCUUCGCGAAGAACAGGGUGAUGUACUACCAGCCGGGGAAAUUUAUGGGCCCCCGUACCGAA ((((((....(((((((((-((((((((((.(((.....)))..))))........)))))..)))))))))).))))))........... ( -33.70) >DroSim_CAF1 4214 90 - 1 GGCCCAGAUAAGAUUUUCC-CCUGGUACCCCUUCGCGAAGAACAGGGUGAUGUACUGCCAGCCGGGGAAAUCUAUGUGCCCUCGUACCGAA (((.((....(((((((((-((((((((((.(((.....)))..))))........)))))..)))))))))).)).)))........... ( -29.50) >DroYak_CAF1 3427 89 - 1 GGCCCAGAUAAGAUUUUCC-CCUGGUAGCCCA-UGCGAGAAACAGGGUGCUGUACUAGCAGUCGGGUAAAUCUAUGGGCGCUUCAACCGAA .(((((....((((((.((-(((.(((.....-))).)).....((.(((((...))))).))))).)))))).)))))............ ( -29.30) >consensus GGCCCAGAUAAGAUUUUCC_CCUGGUACCCCUUCGCGAAGAACAGGGUGAUGUACUACCAGCCGGGGAAAUCUAUGGGCCCUCGUACCGAA ((((((....(((((((((..((((((.....((((..........)))).....))))))...))))))))).))))))........... (-22.98 = -23.85 + 0.87)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:04:18 2006