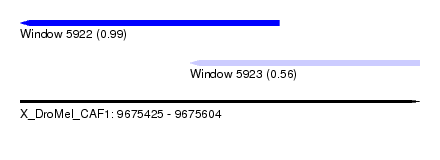
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 9,675,425 – 9,675,604 |
| Length | 179 |
| Max. P | 0.986843 |

| Location | 9,675,425 – 9,675,541 |
|---|---|
| Length | 116 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 82.91 |
| Mean single sequence MFE | -31.20 |
| Consensus MFE | -20.49 |
| Energy contribution | -22.04 |
| Covariance contribution | 1.55 |
| Combinations/Pair | 1.00 |
| Mean z-score | -3.06 |
| Structure conservation index | 0.66 |
| SVM decision value | 2.06 |
| SVM RNA-class probability | 0.986843 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
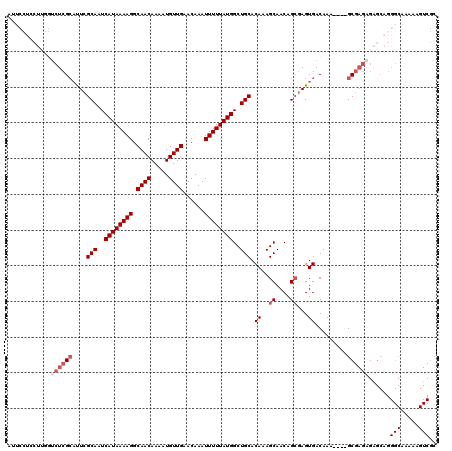
>X_DroMel_CAF1 9675425 116 - 22224390 AUUCCUUCUCGGUCUCGCAUUCGCAAUCAUAAAAGGCAACAAAAUGUUGAACAAAUUUUUAUGGCUGCACAAAGCAACAGCGAGUGACAAA----GCGAGAGAGCAGGGCAAAAAGUCGC ..((((.(((..((((((....(((..((((((((.((((.....))))......))))))))..))).((..((....))...)).....----))))))))).))))........... ( -33.70) >DroAna_CAF1 38469 103 - 1 -------------CUCG--CUCGCAAUCAUAAAAGGCAACAAAAUGUUGAACAAAUUUUUAUGGCUGCACAAAGCAACAG--AGUGACAAAGAAGGCGAGAGGGCAGGGCGAGAAGUCAG -------------((((--((((((..((((((((.((((.....))))......))))))))..)))............--.............((......)).)))))))....... ( -27.90) >DroSim_CAF1 22570 116 - 1 AUUCCUCCUUGGUCUCGCAUUCGCAAUCAUAAAAGGCAACAAAAUGUUGAACAAAUUUUUAUGGCUGCACAAAGCAACAGCGAGUGACAAA----GCGAGAGAGCAGGGCAAAAAGUCGC ......(((((.((((.(((((((...((((((((.((((.....))))......))))))))(((......)))....))))))).....----....)))).)))))........... ( -33.40) >DroEre_CAF1 17934 116 - 1 AUUCCUCCUUGCUCUCGCAUUCGCAAUCAUAAAAGGCAACAAAAUGUUGAACAAAUUUUUAUGGCUGCACAAAGCAACAGCGAGUGACAAA----GCGAGAGAGCAGGGCAAAAAGUCGC ......((((((((((.(((((((...((((((((.((((.....))))......))))))))(((......)))....))))))).....----....))))))))))........... ( -37.90) >DroYak_CAF1 22129 95 - 1 AUUCCUCCUUCGUCUCGCAUUCGCAAUCAUAAAAGGCAACAAAAUGUUGAACAAAUUUUUAUGGCUGCACAAAGCAACAGCGAGUGACAAA----GC---------------------GC ..........(((....(((((((...((((((((.((((.....))))......))))))))(((......)))....))))))).....----))---------------------). ( -23.10) >consensus AUUCCUCCUUGGUCUCGCAUUCGCAAUCAUAAAAGGCAACAAAAUGUUGAACAAAUUUUUAUGGCUGCACAAAGCAACAGCGAGUGACAAA____GCGAGAGAGCAGGGCAAAAAGUCGC ............((((((....(((..((((((((.((((.....))))......))))))))..))).((..((....))...)).........))))))......(((.....))).. (-20.49 = -22.04 + 1.55)


| Location | 9,675,501 – 9,675,604 |
|---|---|
| Length | 103 |
| Sequences | 4 |
| Columns | 104 |
| Reading direction | reverse |
| Mean pairwise identity | 86.00 |
| Mean single sequence MFE | -15.13 |
| Consensus MFE | -11.44 |
| Energy contribution | -11.50 |
| Covariance contribution | 0.06 |
| Combinations/Pair | 1.06 |
| Mean z-score | -1.46 |
| Structure conservation index | 0.76 |
| SVM decision value | 0.06 |
| SVM RNA-class probability | 0.562442 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
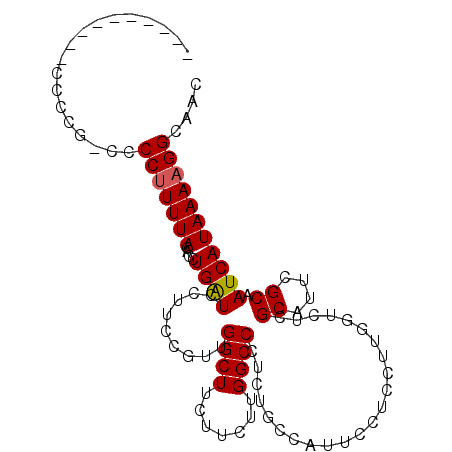
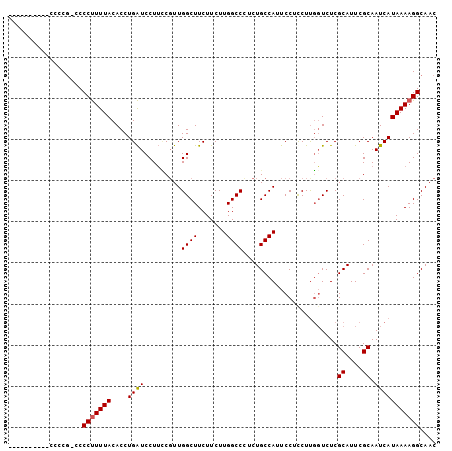
>X_DroMel_CAF1 9675501 103 - 22224390 CACCCCUCAUCCCCG-CCCCUUUUACACCUGAUCCUUCCGUUGGCUUCUUCUUGGCCCUCUGCCAUUCCUUCUCGGUCUCGCAUUCGCAAUCAUAAAAGGCAAC ...............-..(((((((....((((......(..((((.......))))..)(((....((.....))....)))......))))))))))).... ( -14.60) >DroSim_CAF1 22646 91 - 1 ------------CCG-CCCCUUUUACACCUGAUCCUUCCGUUGGCUUCUUCUUGGCCCUCUGCCAUUCCUCCUUGGUCUCGCAUUCGCAAUCAUAAAAGGCAAC ------------...-..(((((((....((((......(..((((.......))))..).((((........))))...((....)).))))))))))).... ( -15.50) >DroEre_CAF1 18010 94 - 1 ----------CCCCACCCCCCUUUACACCUGGUCCUGUCUUUGGCUUCUUCUUGGCCUUCUGCCAUUCCUCCUUGCUCUCGCAUUCGCAAUCAUAAAAGGCAAC ----------.........................((((((((((.......((((.....)))).........)))...((....)).......))))))).. ( -13.59) >DroYak_CAF1 22184 101 - 1 CACUCCC---CCCCGCCCCCUUUUACACCUGAUCCUUGCGUUGGCUUCUUCUUGGCCUUCUGCCAUUCCUCCUUCGUCUCGCAUUCGCAAUCAUAAAAGGCAAC .......---........(((((((....((.....((((..(((.......((((.....))))..........))).))))....))....))))))).... ( -16.83) >consensus __________CCCCG_CCCCUUUUACACCUGAUCCUUCCGUUGGCUUCUUCUUGGCCCUCUGCCAUUCCUCCUUGGUCUCGCAUUCGCAAUCAUAAAAGGCAAC ..................(((((((....((((.........((((.......)))).......................((....)).))))))))))).... (-11.44 = -11.50 + 0.06)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:03:54 2006