
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 9,629,409 – 9,629,597 |
| Length | 188 |
| Max. P | 0.982410 |

| Location | 9,629,409 – 9,629,502 |
|---|---|
| Length | 93 |
| Sequences | 6 |
| Columns | 111 |
| Reading direction | reverse |
| Mean pairwise identity | 77.47 |
| Mean single sequence MFE | -35.22 |
| Consensus MFE | -17.62 |
| Energy contribution | -18.23 |
| Covariance contribution | 0.61 |
| Combinations/Pair | 1.26 |
| Mean z-score | -2.85 |
| Structure conservation index | 0.50 |
| SVM decision value | 1.91 |
| SVM RNA-class probability | 0.982410 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 9629409 93 - 22224390 GGACUCAAAUCGUUUUACGCUGUCGCCGCAUGUGGGCAUUAAGUUAAAUGCCGGCGUAGACGACAGGAACAAUGAUGAUGG----GU----CCUGCCCAUG---------- (((((((.((((((...(.((((((.(..((((.((((((......)))))).)))).).)))))))...))))))..)))----))----))........---------- ( -37.90) >DroGri_CAF1 7424 95 - 1 AGAUUCAAAACGUUUUACGCUGUCGCCGCAUGA-UGCAUUACGUUAAAUGGCAACGUUGAUGUCG---ACGCUGACGAUG------------ACACCGAUGACACCGACUC .................((.((((((((..(((-((.....)))))..))))...((((.(((((---.((....)).))------------))).)))))))).)).... ( -23.30) >DroSec_CAF1 5002 93 - 1 GGACUCAAAUCGUUUUACGCUGUCGCCGCAUGUGGGCAUUAAGUUAAAUGCCGGCGUAGACGACAGGAACAAUGAUGAUGG----GU----CCUGCCCAUG---------- (((((((.((((((...(.((((((.(..((((.((((((......)))))).)))).).)))))))...))))))..)))----))----))........---------- ( -37.90) >DroSim_CAF1 5172 93 - 1 GGACUCAAAUCGUUUUACGCUGUCGCCGCAUGUGGGCAUUAAGUUAAAUGCCGGCGUAGACGACAGGAACAAUGAUGAUGG----GU----CCUGCCCAUG---------- (((((((.((((((...(.((((((.(..((((.((((((......)))))).)))).).)))))))...))))))..)))----))----))........---------- ( -37.90) >DroWil_CAF1 54029 106 - 1 GGACUCAAAACGUUUUACGUUGUCGCCGCAUGUG-GCAUUAAGUUAAAUGGCAACGUAGACGAUGAUGUCGAUGUCGAUGUCGUUGUCGUUGUCGUCGAGGAUGAGG---- ...(((..(((....(((((((((((((....))-))(((......))))))))))))(((((((((((((....))))))))))))))))(((......)))))).---- ( -39.30) >DroYak_CAF1 5856 93 - 1 GGACUCAAAUCGUUUUACGCUGUCGCCGCAUGUGGGCAUUAAGUUAAAUGCCGGCGUAGACGACAGGAACAAUGAUGAUGG----GU----UCUGCCCAUG---------- (((((((.((((((...(.((((((.(..((((.((((((......)))))).)))).).)))))))...))))))..)))----))----))........---------- ( -35.00) >consensus GGACUCAAAUCGUUUUACGCUGUCGCCGCAUGUGGGCAUUAAGUUAAAUGCCGGCGUAGACGACAGGAACAAUGAUGAUGG____GU____CCUGCCCAUG__________ ........((((((.....((((((.(..((((.((((((......)))))).)))).).)))))).......))))))................................ (-17.62 = -18.23 + 0.61)
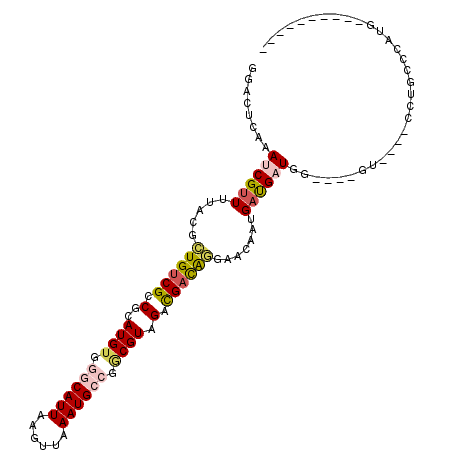


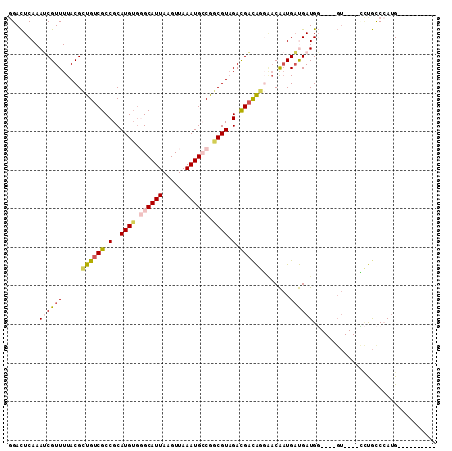
| Location | 9,629,463 – 9,629,565 |
|---|---|
| Length | 102 |
| Sequences | 6 |
| Columns | 102 |
| Reading direction | reverse |
| Mean pairwise identity | 81.79 |
| Mean single sequence MFE | -28.82 |
| Consensus MFE | -18.92 |
| Energy contribution | -20.53 |
| Covariance contribution | 1.61 |
| Combinations/Pair | 1.12 |
| Mean z-score | -1.67 |
| Structure conservation index | 0.66 |
| SVM decision value | 0.14 |
| SVM RNA-class probability | 0.604189 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 9629463 102 - 22224390 ACCGUUACACAUUCCCACACCCACUAUGAACUCCUGUUGCUGCAGCUGGGAAUCUAAUCCCGGGGACUCAAAUCGUUUUACGCUGUCGCCGCAUGUGGGCAU ...................(((((.((((((....)))((.(((((.((((......))))(....)..............))))).))..))))))))... ( -27.10) >DroSec_CAF1 5056 102 - 1 ACCCAUUCCCAUUCCCACACCCACUGGAAACUCCUGUUGCUGCAGCUGGGAAUCUAAUCCCGGGGACUCAAAUCGUUUUACGCUGUCGCCGCAUGUGGGCAU .............((((((......(....)...(((.((.(((((.((((......))))(....)..............))))).)).)))))))))... ( -30.10) >DroSim_CAF1 5226 102 - 1 ACCCAUUCCCAUUCCCACACCCACUGGAAACUCCUGUUGCUGCAGCUGGGAAUCUAAUCCCGGGGACUCAAAUCGUUUUACGCUGUCGCCGCAUGUGGGCAU .............((((((......(....)...(((.((.(((((.((((......))))(....)..............))))).)).)))))))))... ( -30.10) >DroEre_CAF1 5334 94 - 1 --------CCAUUCCCACACCCACUGGGAACCCCUGUUGCUGCAGCUGGGAAUCUAAUCCCGGGGACUCAAAUCGUUUUACGCUGUCGCCGCAUGUGGGCAU --------...((((((.......))))))(((((((.((.(((((.((((......))))(....)..............))))).)).))).).)))... ( -31.60) >DroWil_CAF1 54097 79 - 1 -------------------UCCUCUGGUU-UCUCUGGUGCUCUAUACGAGUAUCUAAUCCA--GGACUCAAAACGUUUUACGUUGUCGCCGCAUGUG-GCAU -------------------....((((..-.....(((((((.....)))))))....)))--)(((....((((.....)))))))((((....))-)).. ( -22.90) >DroYak_CAF1 5910 96 - 1 AU------CCAGACCCACACCCACUGGAAACUCCUGUUGCUGCAGCUGGGAAUCUAAUCCCGGGGACUCAAAUCGUUUUACGCUGUCGCCGCAUGUGGGCAU ..------...(.((((((......(....)...(((.((.(((((.((((......))))(....)..............))))).)).)))))))))).. ( -31.10) >consensus AC______CCAUUCCCACACCCACUGGAAACUCCUGUUGCUGCAGCUGGGAAUCUAAUCCCGGGGACUCAAAUCGUUUUACGCUGUCGCCGCAUGUGGGCAU ...................(((((.((.....))(((.((.(((((.((((......)))).(((((.......)))))..))))).)).))).)))))... (-18.92 = -20.53 + 1.61)



| Location | 9,629,502 – 9,629,597 |
|---|---|
| Length | 95 |
| Sequences | 5 |
| Columns | 103 |
| Reading direction | reverse |
| Mean pairwise identity | 80.43 |
| Mean single sequence MFE | -21.08 |
| Consensus MFE | -13.48 |
| Energy contribution | -13.52 |
| Covariance contribution | 0.04 |
| Combinations/Pair | 1.07 |
| Mean z-score | -2.06 |
| Structure conservation index | 0.64 |
| SVM decision value | 0.84 |
| SVM RNA-class probability | 0.863804 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 9629502 95 - 22224390 CUCCA--ACCCAUUUGAGUUGACCCCAUACCC------AUACCGUUACACAUUCCCACACCCACUAUGAACUCCUGUUGCUGCAGCUGGGAAUCUAAUCCCGG ...((--((.(....).))))...........------...(((......(((((((...........(((....)))((....))))))))).......))) ( -14.12) >DroSec_CAF1 5095 95 - 1 CCCCA--ACCCAUCUGGGUUGACCCCAUACCC------AUACCCAUUCCCAUUCCCACACCCACUGGAAACUCCUGUUGCUGCAGCUGGGAAUCUAAUCCCGG .((((--((((....))))))...........------......(((((((((((..........))))....((((....)))).)))))))........)) ( -22.20) >DroSim_CAF1 5265 101 - 1 CCCCA--ACCCAUCUGGGUUGACCCCAUACCCAUACCCAUACCCAUUCCCAUUCCCACACCCACUGGAAACUCCUGUUGCUGCAGCUGGGAAUCUAAUCCCGG .((((--((((....)))))).......................(((((((((((..........))))....((((....)))).)))))))........)) ( -22.20) >DroEre_CAF1 5373 78 - 1 CCCCAA-GCCCAUCGGGGUUGACC------------------------CCAUUCCCACACCCACUGGGAACCCCUGUUGCUGCAGCUGGGAAUCUAAUCCCGG ......-......(((((((((.(------------------------(((((((((.......))))))...((((....)))).))))..)).))))))). ( -25.30) >DroYak_CAF1 5949 88 - 1 CCCAACCACCCAUCUGGGUUGACC---UACCC------ACAU------CCAGACCCACACCCACUGGAAACUCCUGUUGCUGCAGCUGGGAAUCUAAUCCCGG ..............(((((.....---.))))------)..(------((((...........))))).....((((....))))((((((......)))))) ( -21.60) >consensus CCCCA__ACCCAUCUGGGUUGACCCCAUACCC______AUACC__U_CCCAUUCCCACACCCACUGGAAACUCCUGUUGCUGCAGCUGGGAAUCUAAUCCCGG .......((((....))))......................................................((((....))))((((((......)))))) (-13.48 = -13.52 + 0.04)


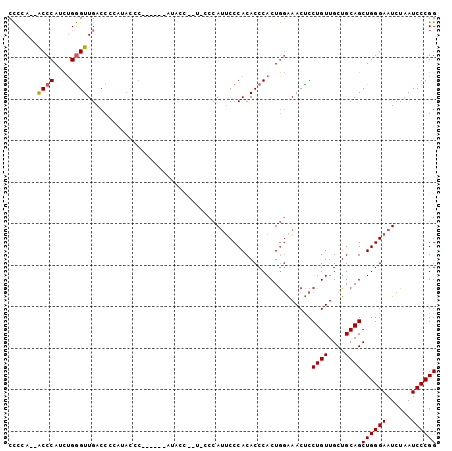
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:03:19 2006