
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 9,488,240 – 9,488,330 |
| Length | 90 |
| Max. P | 0.858010 |

| Location | 9,488,240 – 9,488,330 |
|---|---|
| Length | 90 |
| Sequences | 5 |
| Columns | 94 |
| Reading direction | forward |
| Mean pairwise identity | 68.19 |
| Mean single sequence MFE | -14.26 |
| Consensus MFE | -6.00 |
| Energy contribution | -6.52 |
| Covariance contribution | 0.52 |
| Combinations/Pair | 1.25 |
| Mean z-score | -1.75 |
| Structure conservation index | 0.42 |
| SVM decision value | 0.24 |
| SVM RNA-class probability | 0.650109 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 9488240 90 + 22224390 AACCACGGAUAGUUGGAUGCGCGUAUACUGAUUCACAGAUACGACAA-AA-UGGCAAAACCAAAA--AAAAAAAACAUAAAUGCGCAUAUGGCA ...........((((.((((((((((.(((.....))))))).....-..-(((.....)))...--...............)))))).)))). ( -18.80) >DroGri_CAF1 41902 73 + 1 ------------UUAAAUGCACA---AUAGAUG--CAUAUGCGAUAAGAAGCAAUU-ACA-GACA--CAGACACACACACAUGCACAUAUAGCA ------------.....(((...---(((..((--(((.(((........)))...-...-....--..(.....)....)))))..))).))) ( -7.70) >DroSim_CAF1 31307 87 + 1 AACCACGGAUAGUUGGAUGCGCGUAUACUGAUUCACAGAUACGACAAAAA-UGGCAGAAAAAAACAAAAA------AAAAAUCCGCAUAUGGCA ..((((((((.((.....))((((((.(((.....)))))))).).....-...................------....)))))....))).. ( -13.80) >DroEre_CAF1 31465 80 + 1 AACCACGGAUUGUUGGAUGCGCGUAUACUGAUUCACAGAUACGACAAAAG------------AAA--AAAAAAAACAUAAAUGCGCAUAUGGCA ..........(((((.((((((((((.(((.....)))))))..(....)------------...--...............)))))).))))) ( -14.70) >DroYak_CAF1 31677 89 + 1 AACCACGGAUAGUUGGAUGCGCGUAUACUGAUUCACAGAUACGAUAAAAAACAGCA-AAA--AAA--ACAAAAAACAUAUAUGCGCAUAUGGCA ...........((((.((((((((((((((.....)))....(........)....-...--...--..........))))))))))).)))). ( -16.30) >consensus AACCACGGAUAGUUGGAUGCGCGUAUACUGAUUCACAGAUACGACAAAAA_CAGCA_AAA__AAA__AAAAAAAACAUAAAUGCGCAUAUGGCA ...........((((.((((((((........................................................)))))))).)))). ( -6.00 = -6.52 + 0.52)

| Location | 9,488,240 – 9,488,330 |
|---|---|
| Length | 90 |
| Sequences | 5 |
| Columns | 94 |
| Reading direction | reverse |
| Mean pairwise identity | 68.19 |
| Mean single sequence MFE | -17.87 |
| Consensus MFE | -6.02 |
| Energy contribution | -7.66 |
| Covariance contribution | 1.64 |
| Combinations/Pair | 1.25 |
| Mean z-score | -2.02 |
| Structure conservation index | 0.34 |
| SVM decision value | 0.82 |
| SVM RNA-class probability | 0.858010 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
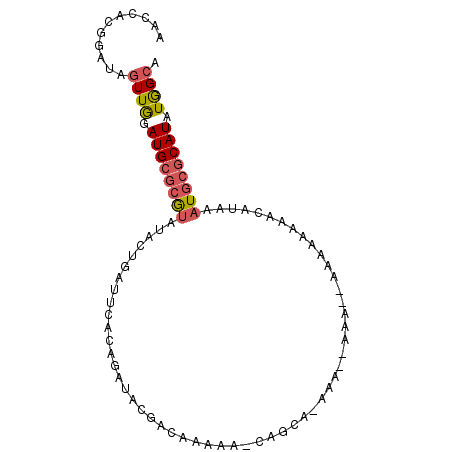
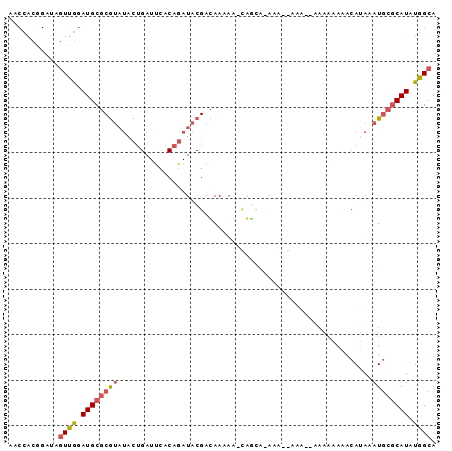
>X_DroMel_CAF1 9488240 90 - 22224390 UGCCAUAUGCGCAUUUAUGUUUUUUUU--UUUUGGUUUUGCCA-UU-UUGUCGUAUCUGUGAAUCAGUAUACGCGCAUCCAACUAUCCGUGGUU .(((((((((((...............--...(((.....)))-..-.....(((((((.....))).))))))))))..........))))). ( -20.60) >DroGri_CAF1 41902 73 - 1 UGCUAUAUGUGCAUGUGUGUGUGUCUG--UGUC-UGU-AAUUGCUUCUUAUCGCAUAUG--CAUCUAU---UGUGCAUUUAA------------ (((.(((.(((((((((((.......(--((..-...-...))).......))))))))--))).)))---...))).....------------ ( -19.04) >DroSim_CAF1 31307 87 - 1 UGCCAUAUGCGGAUUUUU------UUUUUGUUUUUUUCUGCCA-UUUUUGUCGUAUCUGUGAAUCAGUAUACGCGCAUCCAACUAUCCGUGGUU .(((((..(((((.....------............)))))..-....(((((((((((.....))).))))).)))...........))))). ( -17.43) >DroEre_CAF1 31465 80 - 1 UGCCAUAUGCGCAUUUAUGUUUUUUUU--UUU------------CUUUUGUCGUAUCUGUGAAUCAGUAUACGCGCAUCCAACAAUCCGUGGUU .(((((((((((...............--...------------(....)..(((((((.....))).))))))))))..........))))). ( -15.40) >DroYak_CAF1 31677 89 - 1 UGCCAUAUGCGCAUAUAUGUUUUUUGU--UUU--UUU-UGCUGUUUUUUAUCGUAUCUGUGAAUCAGUAUACGCGCAUCCAACUAUCCGUGGUU .(((((((((((.............((--...--...-.))...........(((((((.....))).))))))))))..........))))). ( -16.90) >consensus UGCCAUAUGCGCAUUUAUGUUUUUUUU__UUU__UUU_UGCCA_UUUUUGUCGUAUCUGUGAAUCAGUAUACGCGCAUCCAACUAUCCGUGGUU .(((((((((((.............................((.....))..(((((((.....))).))))))))))..........))))). ( -6.02 = -7.66 + 1.64)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:01:58 2006