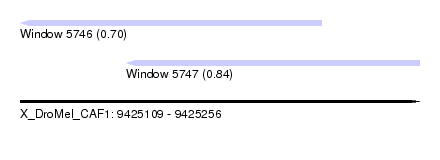
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 9,425,109 – 9,425,256 |
| Length | 147 |
| Max. P | 0.842964 |

| Location | 9,425,109 – 9,425,220 |
|---|---|
| Length | 111 |
| Sequences | 4 |
| Columns | 116 |
| Reading direction | reverse |
| Mean pairwise identity | 88.69 |
| Mean single sequence MFE | -26.23 |
| Consensus MFE | -20.44 |
| Energy contribution | -20.31 |
| Covariance contribution | -0.13 |
| Combinations/Pair | 1.12 |
| Mean z-score | -1.67 |
| Structure conservation index | 0.78 |
| SVM decision value | 0.35 |
| SVM RNA-class probability | 0.699648 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 9425109 111 - 22224390 GAAACUGCGCACUAAAG----AGUUUGCAAAACGUAUUUUGUUGUUGUGUCGCUCACCAGCGAAUCCAAUAACUAUACUAUGCGAUCUGAGUUUGCGGUAUAA-AAUACAAAAAAA ...((((((.(((..((----(..(((((....((((...(((((((..(((((....)))))...))))))).))))..)))))))).))).))))))....-............ ( -29.60) >DroSec_CAF1 6886 108 - 1 GAAACUGCGCAUCCAAG----AGUUUGCAAAACGUAUUUUGUUGUUGUGUCGCUCACCAGCGAAUCCAAUAACUAUACUAUGCGAUCUGAGUUUGCGGUAUAA-AAUACAA---AA ...((((((.(..((.(----(..(((((....((((...(((((((..(((((....)))))...))))))).))))..)))))))))..).))))))....-.......---.. ( -27.40) >DroEre_CAF1 7079 112 - 1 CAAACUGCGUGGAAAAGACAAAGUGUGCAAUACGUAUUUUGUUGUUGUGUUGCUCGCUUGCGAAUCCAAUAACUAUACUAUGCGAUCUGAGUUUGCGGUAUAA-AAUACAA---AA .........((((...(...(((((.((((((((...........)))))))).))))).)...)))).....((((((..((((.......)))))))))).-.......---.. ( -26.00) >DroYak_CAF1 7359 108 - 1 GAAACUGCGCAGCAAAG----AGUUUGCAAAACGUAUU-UGUUGUUGUGUUGCUCGCUAACGAAUCCAAUAACUAUACUAUGCGAUCUGAGUUUGCGGCAUAAAAAUACAA---AA ....(((((.(((..((----(..(((((....((((.-.(((((((.(....(((....)))..)))))))).))))..))))))))..)))))))).............---.. ( -21.90) >consensus GAAACUGCGCAGCAAAG____AGUUUGCAAAACGUAUUUUGUUGUUGUGUCGCUCACCAGCGAAUCCAAUAACUAUACUAUGCGAUCUGAGUUUGCGGUAUAA_AAUACAA___AA ...((((((((..............)))....(((((...(((((((..((((......))))...)))))))......)))))..........)))))................. (-20.44 = -20.31 + -0.13)

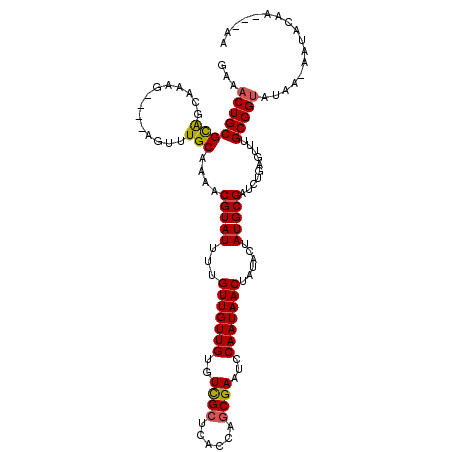

| Location | 9,425,148 – 9,425,256 |
|---|---|
| Length | 108 |
| Sequences | 4 |
| Columns | 112 |
| Reading direction | reverse |
| Mean pairwise identity | 87.42 |
| Mean single sequence MFE | -33.35 |
| Consensus MFE | -25.74 |
| Energy contribution | -25.43 |
| Covariance contribution | -0.31 |
| Combinations/Pair | 1.14 |
| Mean z-score | -1.96 |
| Structure conservation index | 0.77 |
| SVM decision value | 0.76 |
| SVM RNA-class probability | 0.842964 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 9425148 108 - 22224390 ACGCAGUGCUGCACAACGCUGCUGCGAAAGUGUUGCGAAACUGCGCACUAAAG----AGUUUGCAAAACGUAUUUUGUUGUUGUGUCGCUCACCAGCGAAUCCAAUAACUAU .((((((((........)).))))))...((.((((.(((((.(........)----))))))))).)).......(((((((..(((((....)))))...)))))))... ( -31.90) >DroSec_CAF1 6922 108 - 1 AUGCAGUGCUGCACAACGUGGCGGCAAAAGUGUUGCGAAACUGCGCAUCCAAG----AGUUUGCAAAACGUAUUUUGUUGUUGUGUCGCUCACCAGCGAAUCCAAUAACUAU ..(((((((..(.....)..)).((((.....))))...)))))(((..(...----.)..)))............(((((((..(((((....)))))...)))))))... ( -31.80) >DroEre_CAF1 7115 112 - 1 AUGCAGUGUUGCACAACGCUGCGGAGAAAGUGUUGCCAAACUGCGUGGAAAAGACAAAGUGUGCAAUACGUAUUUUGUUGUUGUGUUGCUCGCUUGCGAAUCCAAUAACUAU (((((((.(((..(((((((........))))))).))))))))))(((...(...(((((.((((((((...........)))))))).))))).)...)))......... ( -36.40) >DroYak_CAF1 7396 107 - 1 ACGCAGUGCUGCACAACGCUGCGGCGAAAGUGUUGCGAAACUGCGCAGCAAAG----AGUUUGCAAAACGUAUU-UGUUGUUGUGUUGCUCGCUAACGAAUCCAAUAACUAU .(((((((........)))))))((....))(((((((....(((((((((..----.(..(((.....)))..-).)))))))))...)))).)))............... ( -33.30) >consensus ACGCAGUGCUGCACAACGCUGCGGCGAAAGUGUUGCGAAACUGCGCAGCAAAG____AGUUUGCAAAACGUAUUUUGUUGUUGUGUCGCUCACCAGCGAAUCCAAUAACUAU .((((((....(.(((((((........))))))).)..)))))).............((((...)))).......(((((((..((((......))))...)))))))... (-25.74 = -25.43 + -0.31)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:01:12 2006