
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 9,297,797 – 9,297,900 |
| Length | 103 |
| Max. P | 0.516747 |

| Location | 9,297,797 – 9,297,900 |
|---|---|
| Length | 103 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 74.57 |
| Mean single sequence MFE | -31.02 |
| Consensus MFE | -14.82 |
| Energy contribution | -14.85 |
| Covariance contribution | 0.03 |
| Combinations/Pair | 1.15 |
| Mean z-score | -1.72 |
| Structure conservation index | 0.48 |
| SVM decision value | -0.03 |
| SVM RNA-class probability | 0.516747 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
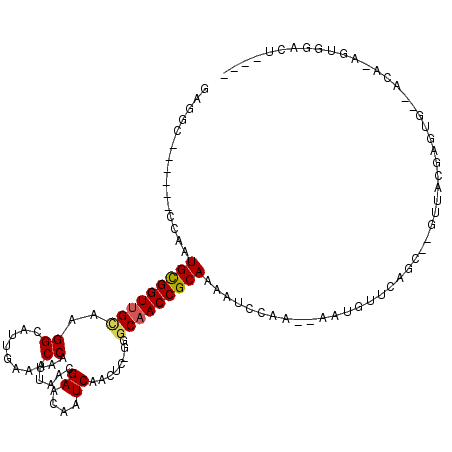
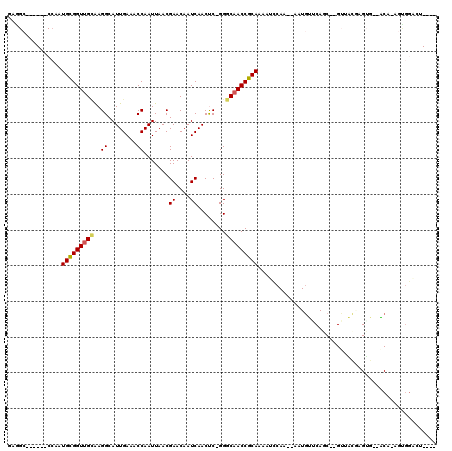
>X_DroMel_CAF1 9297797 103 - 22224390 GAGGC------CCAGUGCGGUUGCAAGGCAUUGAAACCAAUUAACGAACAAUCAACCC-GGGCAACCGCAAAAUCCAA--AUUGUUCAGC--GUUACGAGUG--ACUCCGUGGGUU---- ..(((------(((.(((((((((..((..((((.................)))).))-..)))))))))........--..........--((((....))--))....))))))---- ( -29.23) >DroVir_CAF1 91441 115 - 1 GAAGCUGGCGGCCAAUGUGGUUGUGAGGCAUUGAAACCAAUUAACGAACAAUCAACUCUGGGCAACCGCAAAAUUCAA--AAUGCACAGCCAGUGGCCAGUG--GCC-AGUGGCCUGUGG ...((.((..(((....(((((((..(..((((....))))...)..)))))))......)))..)))).........--....((((((((.(((((...)--)))-).))).))))). ( -42.70) >DroPse_CAF1 86858 105 - 1 GAGGC------CAAAUGCGGUUGCAAGGCAUUGAAACCAAUUAACGAACAAUCAACUC-GGGCAACCGCAAAAUCCAACAAAUUUUCACC--GAUUCUGGGC--ACAAAACAGAGA---- ..((.------....(((((((((..((........))......(((.........))-).)))))))))....))..............--..(((((...--......))))).---- ( -23.70) >DroSim_CAF1 77514 102 - 1 GAGGC------CCAGUGCGGUUGCAAGGCAUUGAAACCAAUUAACGAACAAUCAACCC-GGGCAACCGCAAAAUCCAA--AUUGUUCACC--GUUACAAGUG--ACU-CGCGGGUU---- ..(((------((..(((((((((..((..((((.................)))).))-..)))))))))........--.....((((.--.......)))--)..-...)))))---- ( -29.03) >DroMoj_CAF1 98827 115 - 1 GAGGCUGGCGGCCAAUGCGGUUGUGAGGCAUUGAAACCAAUUAACGAACAAUCAACUCCGGGCAACCGCAAAAUUCAA--AAUGCACAGCCAGUGGCGAGUG--GCC-AGUGCCCUGUCG ..(((.((((((((.(((((((((..((..((((.................))))..))..)))))))))........--........(((...)))...))--)))-...)))..))). ( -38.33) >DroAna_CAF1 86714 98 - 1 -------------AAUGCGGUCGGAAGGCAUUGAAACCAAUUAACGAACAAUCAACUC-GGGCAACCGCAAAAUCCAA--AAUGUUCAGC--GAUAUGCGAGGAGCAAGAAGGAGU---- -------------..(((.(((....))).............................-((....)))))...(((..--..(((((..(--(.....))..)))))....)))..---- ( -23.10) >consensus GAGGC______CCAAUGCGGUUGCAAGGCAUUGAAACCAAUUAACGAACAAUCAACUC_GGGCAACCGCAAAAUCCAA__AAUGUUCAGC__GUUACGAGUG__ACA_AGUGGACU____ ...............(((((((((..((........)).......((....))........))))))))).................................................. (-14.82 = -14.85 + 0.03)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:00:05 2006