
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 9,156,281 – 9,156,458 |
| Length | 177 |
| Max. P | 0.979872 |

| Location | 9,156,281 – 9,156,388 |
|---|---|
| Length | 107 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 82.92 |
| Mean single sequence MFE | -34.97 |
| Consensus MFE | -25.95 |
| Energy contribution | -27.07 |
| Covariance contribution | 1.12 |
| Combinations/Pair | 1.08 |
| Mean z-score | -2.34 |
| Structure conservation index | 0.74 |
| SVM decision value | 1.52 |
| SVM RNA-class probability | 0.961019 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 9156281 107 + 22224390 GGCGCAGAAAAAAG---GGGG-------UAGGAGGUGGGUUUUU---CGAUAUUGGCCGAAGGGCAUUUUUCAAAUUGAAAAGUGAGCCCCAAAACCGCAAGGUUCUUAACUACCUCUAU .............(---((((-------(((...(.((((((..---........(((....)))(((((((.....))))))))))))))..((((....)))).....)))))))).. ( -36.30) >DroSec_CAF1 8949 105 + 1 GGCGCAGAACAAAG---GGGG--------AGCAGGUGGGUU-UU---CGGUUUAGGCCGAAGGGCAUUUUUCAAAUUGAAAAGAGAGCCCCAAAACCGCAAGGUUCUUAACUACCUCUAU ............((---(((.--------((...(.(((((-((---(.......(((....)))..(((((.....))))))))))))))..((((....)))).....)).))))).. ( -33.90) >DroSim_CAF1 9977 105 + 1 GGCGCAGAAAAAAG---GGGG--------AGGAGGUGGGUU-UU---CGGUAUUGGCCGAAGGGCAUUUUUCAAAUUGAAAAGUGAGCCCCAAAACCGCAAGGUUCUUAACUACCUCUAU ..............---....--------.((((((((...-((---((((....))))))(((((((((((.....)))))))..))))...((((....)))).....)))))))).. ( -36.40) >DroYak_CAF1 9465 120 + 1 GGUGCGGAAAAGGGAUUGGGAAACGGGAUUGGGAUUGGGAUUGGGAAUGGAAUUGGCCGAAGGGCAUUUUUCAAUUUGAAAAGUGAGCCCCAAAACCGCAAGGUGCCUAACUACCUCUAU ..........((((.((.(....).))....((.(((((.(((((..........(((....)))(((((((.....)))))))....))))).(((....))).)))))))..)))).. ( -33.30) >consensus GGCGCAGAAAAAAG___GGGG________AGGAGGUGGGUU_UU___CGGUAUUGGCCGAAGGGCAUUUUUCAAAUUGAAAAGUGAGCCCCAAAACCGCAAGGUUCUUAACUACCUCUAU ..............................((((((((.........(((......)))..(((((((((((.....)))))))..))))...((((....)))).....)))))))).. (-25.95 = -27.07 + 1.12)


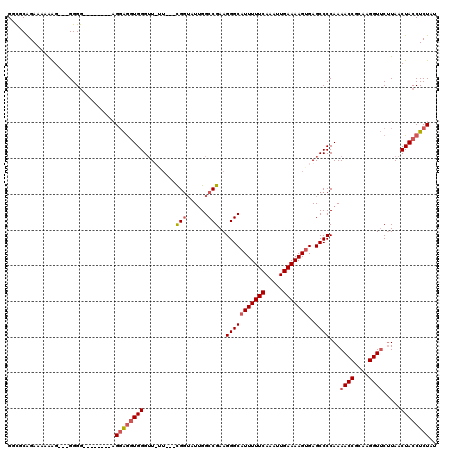
| Location | 9,156,311 – 9,156,421 |
|---|---|
| Length | 110 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 87.43 |
| Mean single sequence MFE | -29.40 |
| Consensus MFE | -24.38 |
| Energy contribution | -24.98 |
| Covariance contribution | 0.60 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.25 |
| Structure conservation index | 0.83 |
| SVM decision value | 1.85 |
| SVM RNA-class probability | 0.979872 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 9156311 110 + 22224390 UUUU---CGAUAUUGGCCGAAGGGCAUUUUUCAAAUUGAAAAGUGAGCCCCAAAACCGCAAGGUUCUUAACUACCUCUAUUUAUUGGCUCACAUAAUAGAUGC-------AAAAGGGUCC ((((---((((.((((((....)))......)))))))))))(((((((....((((....))))....................)))))))......(((.(-------....).))). ( -28.05) >DroSec_CAF1 8978 109 + 1 U-UU---CGGUUUAGGCCGAAGGGCAUUUUUCAAAUUGAAAAGAGAGCCCCAAAACCGCAAGGUUCUUAACUACCUCUAUUUAUUGGCUCACAUAAUAGAUGC-------AAAAGGGUCC .-((---((((....))))))((((.((((((.....))))))...))))...((((....))))........(((.(((((((((.......))))))))).-------...))).... ( -28.20) >DroSim_CAF1 10006 109 + 1 U-UU---CGGUAUUGGCCGAAGGGCAUUUUUCAAAUUGAAAAGUGAGCCCCAAAACCGCAAGGUUCUUAACUACCUCUAUUUAUUGGCUCACAUAUUAGAUGC-------AAAAGGGUCC .-((---((((....))))))((((..(((((.....)))))(((((((....((((....))))....................)))))))...........-------......)))) ( -29.65) >DroEre_CAF1 9298 107 + 1 ----CAAUCC--CAAGCCGAAGGGCAUUUUUCAAUUUGAAAAGUUAGCCCCAAAACCGCAAGGUUCCUAACUACCUCUAUUUAUUGGCUCACAUAAUAGAUGC-------AAAAGGGUCC ----....((--(.((((((.(((((((((((.....)))))))..))))...((((....))))..................))))))..(((.....))).-------....)))... ( -28.00) >DroYak_CAF1 9505 120 + 1 UUGGGAAUGGAAUUGGCCGAAGGGCAUUUUUCAAUUUGAAAAGUGAGCCCCAAAACCGCAAGGUGCCUAACUACCUCUAUUUAUUGGCUCACAUAAUAGAUGCAAAAUGCAAAAGGGUCC (((((..........(((....)))(((((((.....)))))))....))))).(((....)))((((.......((((((...((.....)).))))))(((.....)))...)))).. ( -33.10) >consensus U_UU___CGGUAUUGGCCGAAGGGCAUUUUUCAAAUUGAAAAGUGAGCCCCAAAACCGCAAGGUUCUUAACUACCUCUAUUUAUUGGCUCACAUAAUAGAUGC_______AAAAGGGUCC ...............(((....)))(((((((.....)))))))..((((...((((....))))..........((((((...((.....)).))))))..............)))).. (-24.38 = -24.98 + 0.60)



| Location | 9,156,311 – 9,156,421 |
|---|---|
| Length | 110 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 87.43 |
| Mean single sequence MFE | -29.65 |
| Consensus MFE | -24.20 |
| Energy contribution | -24.44 |
| Covariance contribution | 0.24 |
| Combinations/Pair | 1.04 |
| Mean z-score | -1.65 |
| Structure conservation index | 0.82 |
| SVM decision value | 0.49 |
| SVM RNA-class probability | 0.756518 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 9156311 110 - 22224390 GGACCCUUUU-------GCAUCUAUUAUGUGAGCCAAUAAAUAGAGGUAGUUAAGAACCUUGCGGUUUUGGGGCUCACUUUUCAAUUUGAAAAAUGCCCUUCGGCCAAUAUCG---AAAA ((.((.....-------((((.......(((((((...((((.(((((........)))))...))))...)))))))(((((.....))))))))).....)))).......---.... ( -29.90) >DroSec_CAF1 8978 109 - 1 GGACCCUUUU-------GCAUCUAUUAUGUGAGCCAAUAAAUAGAGGUAGUUAAGAACCUUGCGGUUUUGGGGCUCUCUUUUCAAUUUGAAAAAUGCCCUUCGGCCUAAACCG---AA-A ((........-------(((((((((.(((......)))))))))(((........))).)))((((..(((((....(((((.....)))))..)))))..))))....)).---..-. ( -26.90) >DroSim_CAF1 10006 109 - 1 GGACCCUUUU-------GCAUCUAAUAUGUGAGCCAAUAAAUAGAGGUAGUUAAGAACCUUGCGGUUUUGGGGCUCACUUUUCAAUUUGAAAAAUGCCCUUCGGCCAAUACCG---AA-A ..........-------((((.......(((((((...((((.(((((........)))))...))))...)))))))(((((.....)))))))))..(((((......)))---))-. ( -30.40) >DroEre_CAF1 9298 107 - 1 GGACCCUUUU-------GCAUCUAUUAUGUGAGCCAAUAAAUAGAGGUAGUUAGGAACCUUGCGGUUUUGGGGCUAACUUUUCAAAUUGAAAAAUGCCCUUCGGCUUG--GGAUUG---- ....(((.((-------((.((((((.(((......))))))))).))))..)))..((..((.(....(((((....(((((.....)))))..))))).).))..)--).....---- ( -28.50) >DroYak_CAF1 9505 120 - 1 GGACCCUUUUGCAUUUUGCAUCUAUUAUGUGAGCCAAUAAAUAGAGGUAGUUAGGCACCUUGCGGUUUUGGGGCUCACUUUUCAAAUUGAAAAAUGCCCUUCGGCCAAUUCCAUUCCCAA ((.((.....(((((((.((........(((((((...((((.(((((.(.....))))))...))))...))))))).........)).))))))).....)))).............. ( -32.53) >consensus GGACCCUUUU_______GCAUCUAUUAUGUGAGCCAAUAAAUAGAGGUAGUUAAGAACCUUGCGGUUUUGGGGCUCACUUUUCAAUUUGAAAAAUGCCCUUCGGCCAAUACCG___AA_A ((.((.......................(((((((...((((.(((((........)))))...))))...)))))))(((((.....))))).........)))).............. (-24.20 = -24.44 + 0.24)


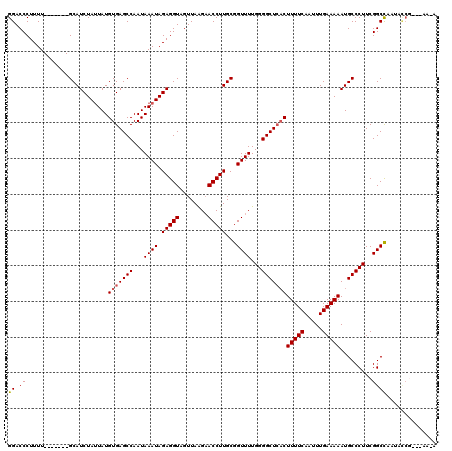
| Location | 9,156,348 – 9,156,458 |
|---|---|
| Length | 110 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 90.37 |
| Mean single sequence MFE | -27.37 |
| Consensus MFE | -21.48 |
| Energy contribution | -22.48 |
| Covariance contribution | 1.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.48 |
| Structure conservation index | 0.78 |
| SVM decision value | 0.11 |
| SVM RNA-class probability | 0.586032 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 9156348 110 + 22224390 AAGUGAGCCCCAAAACCGCAAGGUUCUUAACUACCUCUAUUUAUUGGCUCACAUAAUAGAUGC-------AAAAGGGUCCACUG--UCCU-CGCAAGGGCUCCUCCAUUCGGUGCACUGA ..(((((((....((((....))))....................)))))))....(((.(((-------(..(((((((..((--(...-.))).)))).)))((....))))))))). ( -29.95) >DroSec_CAF1 9014 111 + 1 AAGAGAGCCCCAAAACCGCAAGGUUCUUAACUACCUCUAUUUAUUGGCUCACAUAAUAGAUGC-------AAAAGGGUCCACUG--UCCACCUCACGAGCUCCUCCAUUCGGUGCACUGC .((.(..(((...((((....))))..........((((((...((.....)).))))))...-------....)))..).))(--(.((((....(((...))).....)))).))... ( -23.10) >DroSim_CAF1 10042 111 + 1 AAGUGAGCCCCAAAACCGCAAGGUUCUUAACUACCUCUAUUUAUUGGCUCACAUAUUAGAUGC-------AAAAGGGUCCACUG--UCCACCUCACGAGCUCCUCCAUUCGGUGCACUGC .((((..(((...((((....)))).....................((((........)).))-------....)))..))))(--(.((((....(((...))).....)))).))... ( -25.30) >DroEre_CAF1 9332 111 + 1 AAGUUAGCCCCAAAACCGCAAGGUUCCUAACUACCUCUAUUUAUUGGCUCACAUAAUAGAUGC-------AAAAGGGUCCACUGAGUCCU-UCCACGGGCUCCUCCAUU-GGUGCACUGA .((((((......((((....)))).))))))...((((((...((.....)).))))))(((-------(....((......((((((.-.....))))))..))...-..)))).... ( -27.90) >DroYak_CAF1 9545 116 + 1 AAGUGAGCCCCAAAACCGCAAGGUGCCUAACUACCUCUAUUUAUUGGCUCACAUAAUAGAUGCAAAAUGCAAAAGGGUCCACUG--UCCU-UCCACGGGCUCCUCCAUU-GAUGCACUGA ..(((((((.....(((....))).....................)))))))....(((.((((.((((....(((((((....--....-.....)))).))).))))-..))))))). ( -30.59) >consensus AAGUGAGCCCCAAAACCGCAAGGUUCUUAACUACCUCUAUUUAUUGGCUCACAUAAUAGAUGC_______AAAAGGGUCCACUG__UCCU_CCCACGGGCUCCUCCAUUCGGUGCACUGA .((((.((((...((((....))))..........((((((...((.....)).))))))..............)))).))))...............((.((.......)).))..... (-21.48 = -22.48 + 1.00)


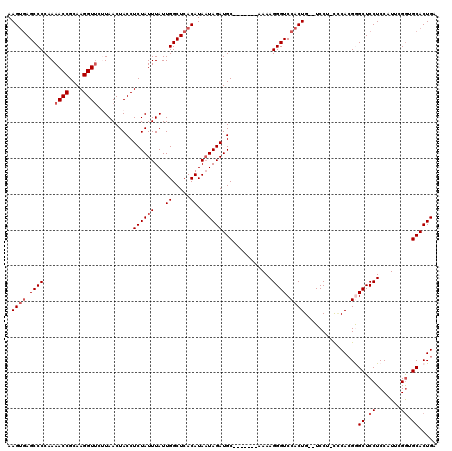
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:59:11 2006