
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 9,154,376 – 9,154,550 |
| Length | 174 |
| Max. P | 0.966286 |

| Location | 9,154,376 – 9,154,489 |
|---|---|
| Length | 113 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 88.71 |
| Mean single sequence MFE | -33.60 |
| Consensus MFE | -25.14 |
| Energy contribution | -25.22 |
| Covariance contribution | 0.08 |
| Combinations/Pair | 1.08 |
| Mean z-score | -2.06 |
| Structure conservation index | 0.75 |
| SVM decision value | 0.75 |
| SVM RNA-class probability | 0.840870 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 9154376 113 + 22224390 ------UAGCUUGAUGUAUCAGCCCGAAUCUGCUGCACGUUUGGGUAAAACCCGGGCGUAAACACUUUAUUUUUGCGCCAAGCUCAUUUGU-UUAUAUGGCGCAAUCGGGGAAUAUAAAC ------........(((((...(((((.........(((((((((.....))))))))).............(((((((((((......))-)....)))))))))))))..)))))... ( -35.70) >DroSec_CAF1 7047 120 + 1 UCUGGCUAGCUUGAUGUAUCAGCCAGAAUCUGCUGCACGUUUGGGUAAAACCCGGGCGUAAACACUUUAUUUUUGCGCCAAGCUCAUUUGUUUUAUAUGGCGCAAUCGGGGAAUAUAAAC (((((((.((.....))...))))))).(((.(...(((((((((.....))))))))).............(((((((((((......))).....))))))))..).)))........ ( -39.00) >DroSim_CAF1 8130 115 + 1 ----GCUAGCUUGAUGUAUCAGCCAGAAUCUGCUGCACGUUUGGGUAAAACCCGGGCGUAAACACUUUAUUUUUGCGCCAAGCUCAUUUGU-UUAUAUGGCGCAAUCGGGGAAUAUAAAC ----.....((((((((..((((........)))).(((((((((.....)))))))))..)))........(((((((((((......))-)....))))))))))))).......... ( -33.40) >DroEre_CAF1 7198 116 + 1 UUUGCUGAGC--CAAGUAUCUGCCAGAAUCUGCUGCACGUUUGGGUAAAACCCGGGCGUAAACACUUUAUUUUUGCGCCAAGCUCAUUUGU-UUAUAUGGCGCAAUCGCGGAAUAUAAA- ....(((.((--.........)))))..(((((...(((((((((.....))))))))).............(((((((((((......))-)....))))))))..))))).......- ( -35.50) >DroYak_CAF1 7299 103 + 1 ----------------UAUCAGUCUGAAUCUGCUGCACGUUUGGGUAAAACCCGGGCGUAAACAAUUUAUUUUUGCGCCAAGCUCAUUUGU-UUAUAUGGCACAAUCGGGAAAUAUAAAC ----------------......(((((.........(((((((((.....))))))))).............(((.(((((((......))-)....)))).)))))))).......... ( -24.40) >consensus ______UAGCUUGAUGUAUCAGCCAGAAUCUGCUGCACGUUUGGGUAAAACCCGGGCGUAAACACUUUAUUUUUGCGCCAAGCUCAUUUGU_UUAUAUGGCGCAAUCGGGGAAUAUAAAC ...................((((........)))).(((((((((.....)))))))))......(((....((((((((.((......))......))))))))....)))........ (-25.14 = -25.22 + 0.08)



| Location | 9,154,410 – 9,154,528 |
|---|---|
| Length | 118 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 92.93 |
| Mean single sequence MFE | -30.74 |
| Consensus MFE | -25.36 |
| Energy contribution | -25.96 |
| Covariance contribution | 0.60 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.00 |
| Structure conservation index | 0.83 |
| SVM decision value | 1.59 |
| SVM RNA-class probability | 0.966286 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 9154410 118 + 22224390 UUGGGUAAAACCCGGGCGUAAACACUUUAUUUUUGCGCCAAGCUCAUUUGU-UUAUAUGGCGCAAUCGGGGAAUAUAAACAAAAGGGCGAUAAUCGUACAAUAAAUACUCUACAAGUGU- .((((.....))))((((((((.........))))))))..((((.(((((-((((((..(.(....).)..))))))))))).))))................(((((.....)))))- ( -33.20) >DroSec_CAF1 7087 120 + 1 UUGGGUAAAACCCGGGCGUAAACACUUUAUUUUUGCGCCAAGCUCAUUUGUUUUAUAUGGCGCAAUCGGGGAAUAUAAACAAAAGGGCGAUAAUCGUACAAUAAAUACUCUACAAGUGUU .((((.....))))((((((((.........))))))))..((((.(((((((.((((..(.(....).)..))))))))))).))))...............((((((.....)))))) ( -30.30) >DroSim_CAF1 8166 118 + 1 UUGGGUAAAACCCGGGCGUAAACACUUUAUUUUUGCGCCAAGCUCAUUUGU-UUAUAUGGCGCAAUCGGGGAAUAUAAACAAAAGGGCGAUAAUCGCACAAUAAAUACUCUACAAGUGU- .((((.....))))((((((((.........))))))))..((((.(((((-((((((..(.(....).)..))))))))))).))))................(((((.....)))))- ( -33.20) >DroEre_CAF1 7236 117 + 1 UUGGGUAAAACCCGGGCGUAAACACUUUAUUUUUGCGCCAAGCUCAUUUGU-UUAUAUGGCGCAAUCGCGGAAUAUAAA-AAAAAGGCGGUCAACGAACAAUCAAUGCUCUACAAGUGC- .((((.....))))((((((((.........)))))))).....(((((((-((((((..(((....)))..)))))).-....(((((................))).))))))))).- ( -27.69) >DroYak_CAF1 7323 118 + 1 UUGGGUAAAACCCGGGCGUAAACAAUUUAUUUUUGCGCCAAGCUCAUUUGU-UUAUAUGGCACAAUCGGGAAAUAUAAACAAAAAGGCGGUCAACGAACACUCAAUGCUCUACAAUUGU- ..(((((..(((..((((((((.........))))))))..(((..(((((-((((((..(........)..)))))))))))..))))))....((....))..))))).........- ( -29.30) >consensus UUGGGUAAAACCCGGGCGUAAACACUUUAUUUUUGCGCCAAGCUCAUUUGU_UUAUAUGGCGCAAUCGGGGAAUAUAAACAAAAGGGCGAUAAUCGAACAAUAAAUACUCUACAAGUGU_ .((((.....))))((((((((.........))))))))..((((.(((((.((((((..(.(....).)..))))))))))).))))................................ (-25.36 = -25.96 + 0.60)



| Location | 9,154,410 – 9,154,528 |
|---|---|
| Length | 118 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 92.93 |
| Mean single sequence MFE | -27.54 |
| Consensus MFE | -23.43 |
| Energy contribution | -23.19 |
| Covariance contribution | -0.24 |
| Combinations/Pair | 1.17 |
| Mean z-score | -1.24 |
| Structure conservation index | 0.85 |
| SVM decision value | 0.03 |
| SVM RNA-class probability | 0.547206 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 9154410 118 - 22224390 -ACACUUGUAGAGUAUUUAUUGUACGAUUAUCGCCCUUUUGUUUAUAUUCCCCGAUUGCGCCAUAUAA-ACAAAUGAGCUUGGCGCAAAAAUAAAGUGUUUACGCCCGGGUUUUACCCAA -(((((((..((((((..((.(..((.....))..)....))..))))))..)..((((((((.....-...........)))))))).....))))))........(((.....))).. ( -27.29) >DroSec_CAF1 7087 120 - 1 AACACUUGUAGAGUAUUUAUUGUACGAUUAUCGCCCUUUUGUUUAUAUUCCCCGAUUGCGCCAUAUAAAACAAAUGAGCUUGGCGCAAAAAUAAAGUGUUUACGCCCGGGUUUUACCCAA ((((((((..((((((..((.(..((.....))..)....))..))))))..)..((((((((.................)))))))).....))))))).......(((.....))).. ( -28.13) >DroSim_CAF1 8166 118 - 1 -ACACUUGUAGAGUAUUUAUUGUGCGAUUAUCGCCCUUUUGUUUAUAUUCCCCGAUUGCGCCAUAUAA-ACAAAUGAGCUUGGCGCAAAAAUAAAGUGUUUACGCCCGGGUUUUACCCAA -(((((((..((((((..((.(.(((.....))).)....))..))))))..)..((((((((.....-...........)))))))).....))))))........(((.....))).. ( -30.19) >DroEre_CAF1 7236 117 - 1 -GCACUUGUAGAGCAUUGAUUGUUCGUUGACCGCCUUUUU-UUUAUAUUCCGCGAUUGCGCCAUAUAA-ACAAAUGAGCUUGGCGCAAAAAUAAAGUGUUUACGCCCGGGUUUUACCCAA -((((((...(((((.....)))))......(((......-..........))).((((((((.....-...........)))))))).....))))))........(((.....))).. ( -28.18) >DroYak_CAF1 7323 118 - 1 -ACAAUUGUAGAGCAUUGAGUGUUCGUUGACCGCCUUUUUGUUUAUAUUUCCCGAUUGUGCCAUAUAA-ACAAAUGAGCUUGGCGCAAAAAUAAAUUGUUUACGCCCGGGUUUUACCCAA -((((((...((((((...))))))......((((..(((((((((((..(........)..))))))-))))).......))))........))))))........(((.....))).. ( -23.90) >consensus _ACACUUGUAGAGUAUUUAUUGUACGAUUAUCGCCCUUUUGUUUAUAUUCCCCGAUUGCGCCAUAUAA_ACAAAUGAGCUUGGCGCAAAAAUAAAGUGUUUACGCCCGGGUUUUACCCAA .(((((((..((((((..((.(..((.....))..)....))..))))))..)..((((((((.................)))))))).....))))))........(((.....))).. (-23.43 = -23.19 + -0.24)



| Location | 9,154,450 – 9,154,550 |
|---|---|
| Length | 100 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 76.48 |
| Mean single sequence MFE | -20.88 |
| Consensus MFE | -10.66 |
| Energy contribution | -11.26 |
| Covariance contribution | 0.60 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.21 |
| Structure conservation index | 0.51 |
| SVM decision value | 1.37 |
| SVM RNA-class probability | 0.946788 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 9154450 100 + 22224390 AGCUCAUUUGU-UUAUAUGGCGCAAUCGGGGAAUAUAAACAAAAGGGCGAUAAUCGUACAAUAAAUACUCUACAAGUGU-UAUU---------AAAA-CCAAGUGU---UAUUAA----- .((((.(((((-((((((..(.(....).)..))))))))))).))))...............((((((.....)))))-)...---------....-........---......----- ( -20.10) >DroSec_CAF1 7127 103 + 1 AGCUCAUUUGUUUUAUAUGGCGCAAUCGGGGAAUAUAAACAAAAGGGCGAUAAUCGUACAAUAAAUACUCUACAAGUGUUUAUU---------AAUA---AAGUACCAAUAUAAA----- .((((.(((((((.((((..(.(....).)..))))))))))).)))).......((((((((((((((.....))))))))))---------....---..)))).........----- ( -21.90) >DroSim_CAF1 8206 99 + 1 AGCUCAUUUGU-UUAUAUGGCGCAAUCGGGGAAUAUAAACAAAAGGGCGAUAAUCGCACAAUAAAUACUCUACAAGUGU-UAUU---------AAUA----AGUACAAAUAUAA------ .((((.(((((-((((((..(.(....).)..))))))))))).))))..((((.((((................))))-.)))---------)...----.............------ ( -20.89) >DroEre_CAF1 7276 117 + 1 AGCUCAUUUGU-UUAUAUGGCGCAAUCGCGGAAUAUAAA-AAAAAGGCGGUCAACGAACAAUCAAUGCUCUACAAGUGC-GUUAGACCAGUAAAAUUAAAAAAAAAUAUCAUACAAAAAC .(((..(((.(-((((((..(((....)))..)))))))-)))..)))((((...((....))(((((.(.....).))-))).))))................................ ( -21.90) >DroYak_CAF1 7363 109 + 1 AGCUCAUUUGU-UUAUAUGGCACAAUCGGGAAAUAUAAACAAAAAGGCGGUCAACGAACACUCAAUGCUCUACAAUUGU-GUUU---------AAUAACAAAAUAUUAUAAUACAAAUAU .(((..(((((-((((((..(........)..)))))))))))..)))...........................((((-((((---------((((......))))).))))))).... ( -19.60) >consensus AGCUCAUUUGU_UUAUAUGGCGCAAUCGGGGAAUAUAAACAAAAGGGCGAUAAUCGAACAAUAAAUACUCUACAAGUGU_UAUU_________AAUA___AAGUACUAAUAUAAA_____ .((((.(((((.((((((..(.(....).)..))))))))))).))))........................................................................ (-10.66 = -11.26 + 0.60)

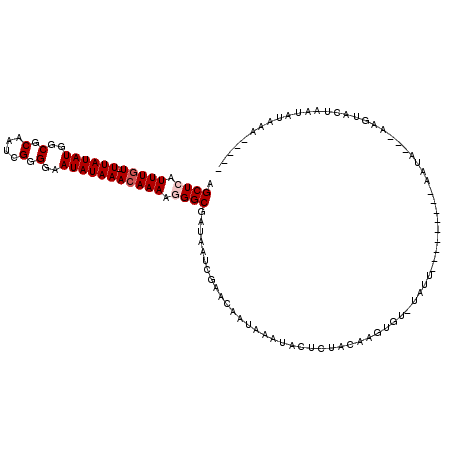
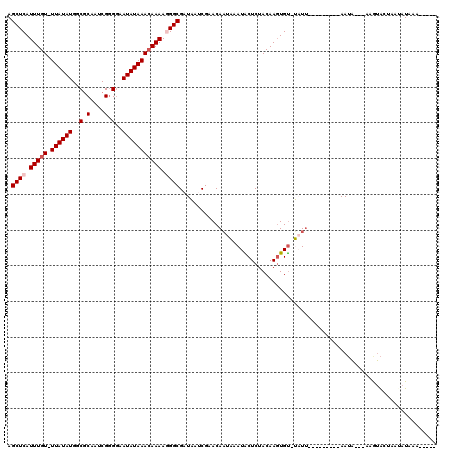
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:59:05 2006