
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 9,071,468 – 9,071,598 |
| Length | 130 |
| Max. P | 0.874997 |

| Location | 9,071,468 – 9,071,558 |
|---|---|
| Length | 90 |
| Sequences | 5 |
| Columns | 105 |
| Reading direction | forward |
| Mean pairwise identity | 78.75 |
| Mean single sequence MFE | -35.02 |
| Consensus MFE | -15.36 |
| Energy contribution | -16.20 |
| Covariance contribution | 0.84 |
| Combinations/Pair | 1.17 |
| Mean z-score | -1.91 |
| Structure conservation index | 0.44 |
| SVM decision value | -0.04 |
| SVM RNA-class probability | 0.510904 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 9071468 90 + 22224390 CUCGAAGCUCCGCAGGAAUC------CUCCUUCGGAUACGAUUACUCGCGACCC----UCGGC-----CGGAAGUAGCACUGACUUCCGGCCAUCCGUCCAGGUG .(((....((((.((((...------.)))).))))..))).(((..(.(((..----..(((-----(((((((.......))))))))))....))))..))) ( -34.80) >DroSec_CAF1 4637 90 + 1 CUCGAAGCUCCGCAGGAGUC------CUCCUUCGGAUACGAUUACUCCCGACCC----UCGGC-----CGGAAGUAGCACUGACUUCCGGCCAUCAGUACAGGUG .(((....((((.((((...------.)))).))))..)))......((.((..----..(((-----(((((((.......))))))))))....))...)).. ( -32.60) >DroSim_CAF1 5045 90 + 1 CUGGAGGCUCCACAGGAAUC------CUCCUUCGGAUACGAAUACUCGCGACCC----UCGGC-----CGGAAGUAGCACUGACUUCCGGCCAUCCGUACAGGUG ..(((((.(((...)))..)------))))..(((((.(((....)))......----..(((-----(((((((.......)))))))))))))))........ ( -35.70) >DroYak_CAF1 5109 90 + 1 CUCGAAGCUCCGCAAGAAUC------CUCCUUUGGAUACGAUUACUCGCGUCCC----UCGGC-----CGGAAGUAGCACUGACUUCCGGCCAUCUGUUCAGGUG ..........(((..((((.------.......(((..(((....)))..))).----..(((-----(((((((.......))))))))))....))))..))) ( -30.30) >DroPer_CAF1 8887 105 + 1 CUGGAGGCCCCCGAGGAGUCGUCGUCCGCCUACGGCUAUGACUACUCCAGGCCGAGCAGCAGCAGUAGUGGAAGCAGCUCGGGCUUCCGACCAUCUCCACAGCUG .(((((((((((..(((((.(((((..(((...))).))))).))))).))..((((.((.((....))....)).)))))))))))))................ ( -41.70) >consensus CUCGAAGCUCCGCAGGAAUC______CUCCUUCGGAUACGAUUACUCGCGACCC____UCGGC_____CGGAAGUAGCACUGACUUCCGGCCAUCCGUACAGGUG ........((((.((((..........)))).))))........................(((.....(((((((.......))))))))))............. (-15.36 = -16.20 + 0.84)
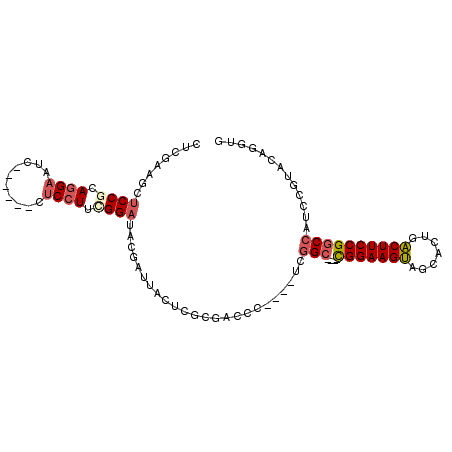



| Location | 9,071,468 – 9,071,558 |
|---|---|
| Length | 90 |
| Sequences | 5 |
| Columns | 105 |
| Reading direction | reverse |
| Mean pairwise identity | 78.75 |
| Mean single sequence MFE | -40.32 |
| Consensus MFE | -20.80 |
| Energy contribution | -21.12 |
| Covariance contribution | 0.32 |
| Combinations/Pair | 1.30 |
| Mean z-score | -2.18 |
| Structure conservation index | 0.52 |
| SVM decision value | 0.89 |
| SVM RNA-class probability | 0.874997 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 9071468 90 - 22224390 CACCUGGACGGAUGGCCGGAAGUCAGUGCUACUUCCG-----GCCGA----GGGUCGCGAGUAAUCGUAUCCGAAGGAG------GAUUCCUGCGGAGCUUCGAG ..((.....))..((((((((((.......)))))))-----)))((----((...((((....)))).((((.(((..------....))).)))).))))... ( -38.30) >DroSec_CAF1 4637 90 - 1 CACCUGUACUGAUGGCCGGAAGUCAGUGCUACUUCCG-----GCCGA----GGGUCGGGAGUAAUCGUAUCCGAAGGAG------GACUCCUGCGGAGCUUCGAG ..((((.(((..(((((((((((.......)))))))-----)))).----.))))))).....(((..((((.((((.------...)))).))))....))). ( -41.90) >DroSim_CAF1 5045 90 - 1 CACCUGUACGGAUGGCCGGAAGUCAGUGCUACUUCCG-----GCCGA----GGGUCGCGAGUAUUCGUAUCCGAAGGAG------GAUUCCUGUGGAGCCUCCAG ........(((((((((((((((.......)))))))-----)))..----(((((....).))))..)))))..((((------(.((((...))))))))).. ( -41.00) >DroYak_CAF1 5109 90 - 1 CACCUGAACAGAUGGCCGGAAGUCAGUGCUACUUCCG-----GCCGA----GGGACGCGAGUAAUCGUAUCCAAAGGAG------GAUUCUUGCGGAGCUUCGAG .............((((((((((.......)))))))-----)))((----((..((((((.((((...(((...))).------))))))))))...))))... ( -36.30) >DroPer_CAF1 8887 105 - 1 CAGCUGUGGAGAUGGUCGGAAGCCCGAGCUGCUUCCACUACUGCUGCUGCUCGGCCUGGAGUAGUCAUAGCCGUAGGCGGACGACGACUCCUCGGGGGCCUCCAG ((((.((((((..((((((....)))).))..))))))....))))(((...((((((((((.(((....(((....)))..))).)))))....)))))..))) ( -44.10) >consensus CACCUGUACAGAUGGCCGGAAGUCAGUGCUACUUCCG_____GCCGA____GGGUCGCGAGUAAUCGUAUCCGAAGGAG______GAUUCCUGCGGAGCUUCGAG ............(((((((((((.......))))))).....))))......(((.((((....)))).((((.(((((........))))).)))))))..... (-20.80 = -21.12 + 0.32)



| Location | 9,071,502 – 9,071,598 |
|---|---|
| Length | 96 |
| Sequences | 5 |
| Columns | 105 |
| Reading direction | forward |
| Mean pairwise identity | 80.02 |
| Mean single sequence MFE | -41.08 |
| Consensus MFE | -22.32 |
| Energy contribution | -22.04 |
| Covariance contribution | -0.28 |
| Combinations/Pair | 1.37 |
| Mean z-score | -1.97 |
| Structure conservation index | 0.54 |
| SVM decision value | 0.28 |
| SVM RNA-class probability | 0.665609 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 9071502 96 + 22224390 AUUACUCGCGACCC----UCGGC-----CGGAAGUAGCACUGACUUCCGGCCAUCCGUCCAGGUGCCAUCCAGUGACCCGUACGGCGAGGAGGAACCGGCGGACA ..............----..(((-----(((((((.......)))))))))).((((.((.(((.((.(((..((.((.....)))).))))).))))))))).. ( -38.80) >DroSec_CAF1 4671 96 + 1 AUUACUCCCGACCC----UCGGC-----CGGAAGUAGCACUGACUUCCGGCCAUCAGUACAGGUGUCAUCCAGUGACCCGUACGGCGAGGAGGAACCGGCGGACA ....((((((.((.----..(((-----(((((((.......))))))))))....((((.((.(((((...))))))))))))))).))))............. ( -41.80) >DroSim_CAF1 5079 96 + 1 AAUACUCGCGACCC----UCGGC-----CGGAAGUAGCACUGACUUCCGGCCAUCCGUACAGGUGCCAUCCAGUGACCCGUACGGCGAGGAGGAACCGGCGGACA .....((((...((----(((((-----(((((((.......))))))))))..((((((.(((((......)).))).)))))).)))).(....).))))... ( -42.90) >DroYak_CAF1 5143 96 + 1 AUUACUCGCGUCCC----UCGGC-----CGGAAGUAGCACUGACUUCCGGCCAUCUGUUCAGGUGCCAUCCAGUGACCCGUACGGCGAGGAGGAACCGGCGGACA .........((((.----(((((-----(((((((.......)))))))))).........(((.((.(((..((.((.....)))).))))).))))).)))). ( -38.50) >DroPer_CAF1 8927 105 + 1 ACUACUCCAGGCCGAGCAGCAGCAGUAGUGGAAGCAGCUCGGGCUUCCGACCAUCUCCACAGCUGCCCAGCGGCAGCGAAGGCUACGAGGAUGAGGUGGCGGACA ........((.((((((.((.((....))....)).)))))).))((((.(((((((....(((((...)))))(((....)))........))))))))))).. ( -43.40) >consensus AUUACUCGCGACCC____UCGGC_____CGGAAGUAGCACUGACUUCCGGCCAUCCGUACAGGUGCCAUCCAGUGACCCGUACGGCGAGGAGGAACCGGCGGACA ....................(((.....(((((((.......)))))))))).(((((...(((.((.(((..((.((.....)))).))))).))).))))).. (-22.32 = -22.04 + -0.28)


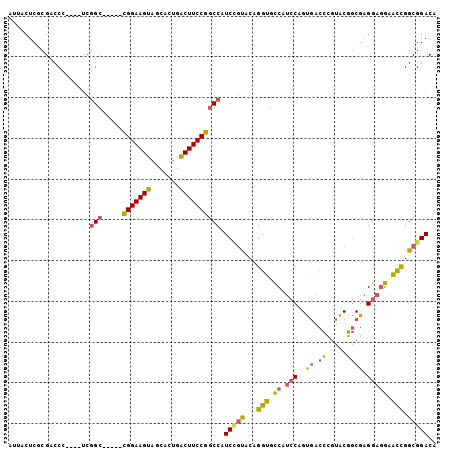
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:58:16 2006