
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 9,000,108 – 9,000,301 |
| Length | 193 |
| Max. P | 0.991213 |

| Location | 9,000,108 – 9,000,225 |
|---|---|
| Length | 117 |
| Sequences | 4 |
| Columns | 117 |
| Reading direction | reverse |
| Mean pairwise identity | 83.87 |
| Mean single sequence MFE | -37.27 |
| Consensus MFE | -33.05 |
| Energy contribution | -33.12 |
| Covariance contribution | 0.06 |
| Combinations/Pair | 1.05 |
| Mean z-score | -2.54 |
| Structure conservation index | 0.89 |
| SVM decision value | 2.25 |
| SVM RNA-class probability | 0.991213 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 9000108 117 - 22224390 UCAGUUGGCCGCCAGCGCUUCGGCGCUGUGCAUCGGUCCAAACUUUGGCUCCCAGCGAUAUCCAUCCGUGUAUAUCCGUCAAGAGUCCACGACACAGCACAAUCGUAUCCUGGCCAA ....((((((((((((((....)))))).))...((..........(((((...(((((((.((....)).)))).)))...))))).((((..........))))..)).)))))) ( -38.90) >DroSec_CAF1 799 102 - 1 UCAGUUGGCCGCCAGCGCUUCGGCGCUGUGCAUCGGUCCAAACUUUGGCUCCCCGCCAUAUUCAUCCGUGUAUAUCCGUCAAGAGU---------------AUCGUAUCCUGGCCAG .....(((((((((((((....)))))).))..(((.........((((.....)))).......)))..................---------------..........))))). ( -34.89) >DroEre_CAF1 858 113 - 1 UCAGUUGGCCGCCAGCGCUUCGGCGCUGUGCAUCGGUCCUAAUUUUGGCUCCCCGCCAUAUCCACCCGUGUAUAUCAGUC-AGAGUCCGC-ACAUUGCC--CACAAAUCCUGGCCAA ....((((((((((((((....)))))).))...(((........((((.....)))).........((((...((....-.))....))-))...)))--..........)))))) ( -38.40) >DroYak_CAF1 839 109 - 1 UCAGUUGGCCGCCAGCGCUUCGGCGCUGUGCAUCGGUCCUAACUUUGGCUCCCCGCCACAACCACCCGUGUAUAUCCUCC-AGAGUCCAC-ACA----C--AACAUAUCCUGGCCAA ....((((((((((((((....)))))).))...((.....(((((((........(((........)))........))-)))))))..-...----.--..........)))))) ( -36.89) >consensus UCAGUUGGCCGCCAGCGCUUCGGCGCUGUGCAUCGGUCCAAACUUUGGCUCCCCGCCAUAUCCACCCGUGUAUAUCCGUC_AGAGUCCAC_ACA____C__AACAUAUCCUGGCCAA ....((((((((((((((....)))))).))..(((.........((((.....)))).......)))...........................................)))))) (-33.05 = -33.12 + 0.06)



| Location | 9,000,148 – 9,000,261 |
|---|---|
| Length | 113 |
| Sequences | 4 |
| Columns | 113 |
| Reading direction | forward |
| Mean pairwise identity | 92.48 |
| Mean single sequence MFE | -40.86 |
| Consensus MFE | -36.66 |
| Energy contribution | -37.85 |
| Covariance contribution | 1.19 |
| Combinations/Pair | 1.14 |
| Mean z-score | -1.98 |
| Structure conservation index | 0.90 |
| SVM decision value | 1.24 |
| SVM RNA-class probability | 0.934578 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 9000148 113 + 22224390 GGAUAUACACGGAUGGAUAUCGCUGGGAGCCAAAGUUUGGACCGAUGCACAGCGCCGAAGCGCUGGCGGCCAACUGACGAGGAACUUUGGUUCGUGGUCAUAACGAUAACGAG .........((.((.(.(((.((((.((((((((((((((.(((.....((((((....)))))).)))))..((....)))))))))))))).)))).))).).))..)).. ( -40.00) >DroSec_CAF1 824 113 + 1 GGAUAUACACGGAUGAAUAUGGCGGGGAGCCAAAGUUUGGACCGAUGCACAGCGCCGAAGCGCUGGCGGCCAACUGACGAGGAACUUUGGUUCGUCGUCACAACGAUAACGAG .........((........((((((.((((((((((((((.(((.....((((((....)))))).)))))..((....)))))))))))))).)))))).........)).. ( -43.33) >DroEre_CAF1 894 113 + 1 UGAUAUACACGGGUGGAUAUGGCGGGGAGCCAAAAUUAGGACCGAUGCACAGCGCCGAAGCGCUGGCGGCCAACUGACGAGGAACUUUGGUUUGGGGUCACAACGAUAACAAG .........((..((....((((.....)))).......((((..(((.((((((....))))))))).(((((..(.(.....).)..).)))))))).)).))........ ( -32.60) >DroYak_CAF1 871 113 + 1 GGAUAUACACGGGUGGUUGUGGCGGGGAGCCAAAGUUAGGACCGAUGCACAGCGCCGAAGCGCUGGCGGCCAACUGACAAGGAACUUUGGUUCGUGGUUACAACGAUAACAAG ............((.((((((((.(.(((((((((((.((.(((.....((((((....)))))).)))))..((....)).))))))))))).).)))))))).))...... ( -47.50) >consensus GGAUAUACACGGAUGGAUAUGGCGGGGAGCCAAAGUUAGGACCGAUGCACAGCGCCGAAGCGCUGGCGGCCAACUGACGAGGAACUUUGGUUCGUGGUCACAACGAUAACAAG ............((.(.((((((((.(((((((((((.((.(((.....((((((....)))))).)))))..((....)).))))))))))).)))))))).).))...... (-36.66 = -37.85 + 1.19)
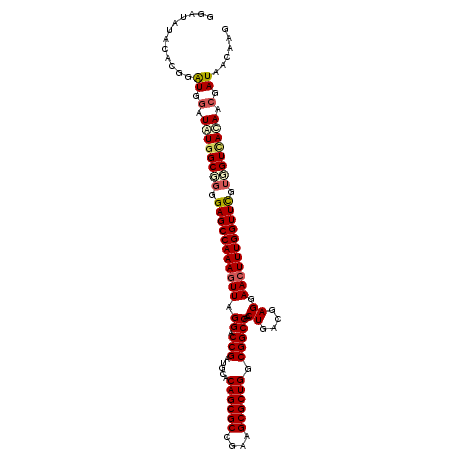


| Location | 9,000,148 – 9,000,261 |
|---|---|
| Length | 113 |
| Sequences | 4 |
| Columns | 113 |
| Reading direction | reverse |
| Mean pairwise identity | 92.48 |
| Mean single sequence MFE | -34.62 |
| Consensus MFE | -29.74 |
| Energy contribution | -29.43 |
| Covariance contribution | -0.31 |
| Combinations/Pair | 1.11 |
| Mean z-score | -1.88 |
| Structure conservation index | 0.86 |
| SVM decision value | 0.83 |
| SVM RNA-class probability | 0.861178 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 9000148 113 - 22224390 CUCGUUAUCGUUAUGACCACGAACCAAAGUUCCUCGUCAGUUGGCCGCCAGCGCUUCGGCGCUGUGCAUCGGUCCAAACUUUGGCUCCCAGCGAUAUCCAUCCGUGUAUAUCC .....(((((((.((((...((((....))))...))))(..((((((((((((....)))))).))...(((....)))..))))..))))))))................. ( -36.20) >DroSec_CAF1 824 113 - 1 CUCGUUAUCGUUGUGACGACGAACCAAAGUUCCUCGUCAGUUGGCCGCCAGCGCUUCGGCGCUGUGCAUCGGUCCAAACUUUGGCUCCCCGCCAUAUUCAUCCGUGUAUAUCC .............((((((.((((....)))).)))))).((((((((((((((....)))))).))...)).))))....((((.....))))................... ( -38.70) >DroEre_CAF1 894 113 - 1 CUUGUUAUCGUUGUGACCCCAAACCAAAGUUCCUCGUCAGUUGGCCGCCAGCGCUUCGGCGCUGUGCAUCGGUCCUAAUUUUGGCUCCCCGCCAUAUCCACCCGUGUAUAUCA .........((((.((((((((......(.....).....))))..((((((((....)))))).))...)))).))))..((((.....))))................... ( -30.20) >DroYak_CAF1 871 113 - 1 CUUGUUAUCGUUGUAACCACGAACCAAAGUUCCUUGUCAGUUGGCCGCCAGCGCUUCGGCGCUGUGCAUCGGUCCUAACUUUGGCUCCCCGCCACAACCACCCGUGUAUAUCC .........(((((......((.((((((((((..(((....))).((((((((....)))))).))...))....)))))))).))......)))))............... ( -33.40) >consensus CUCGUUAUCGUUGUGACCACGAACCAAAGUUCCUCGUCAGUUGGCCGCCAGCGCUUCGGCGCUGUGCAUCGGUCCAAACUUUGGCUCCCCGCCAUAUCCACCCGUGUAUAUCC ...(((((....)))))...((.((((((((((..(((....))).((((((((....)))))).))...))....)))))))).)).......................... (-29.74 = -29.43 + -0.31)


| Location | 9,000,188 – 9,000,301 |
|---|---|
| Length | 113 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 75.27 |
| Mean single sequence MFE | -39.68 |
| Consensus MFE | -24.98 |
| Energy contribution | -22.98 |
| Covariance contribution | -1.99 |
| Combinations/Pair | 1.36 |
| Mean z-score | -1.76 |
| Structure conservation index | 0.63 |
| SVM decision value | 1.18 |
| SVM RNA-class probability | 0.926887 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 9000188 113 - 22224390 AAAAGCAGGCAGCUGCCUCAAAAAAAUUUCACCUUCAACCCUCGUUAUCGUUAUGACCACGAACCAAAG----UUCCUCG---UCAGUUGGCCGCCAGCGCUUCGGCGCUGUGCAUCGGU ....(.((((....)))))...........(((.(((((...((....))...((((...((((....)----)))...)---))))))))..((((((((....)))))).))...))) ( -33.70) >DroPse_CAF1 5813 109 - 1 AGCAGCUAGCAGCUGGCC---AAAGAU------UUCAACCCUCGCUGGGGUUUC--CUCUGGGCCACAGCGGACUCAUCGGCCACAGUUGGCUGCCAGUGCUCCGGCGCUGAGCAUCGCC .(((((((((.(.(((((---......------...((((((....))))))((--(.(((.....))).)))......)))))).)))))))))((((((....))))))......... ( -47.10) >DroSec_CAF1 864 113 - 1 AAAAGCAGGCAGCUGCCUCAAAAAAAUUUCACCUUCAACCCUCGUUAUCGUUGUGACGACGAACCAAAG----UUCCUCG---UCAGUUGGCCGCCAGCGCUUCGGCGCUGUGCAUCGGU ....(.((((....)))))...........(((.(((((...((....))...((((((.((((....)----))).)))---))))))))..((((((((....)))))).))...))) ( -39.30) >DroEre_CAF1 934 113 - 1 AAAAGCAGGCAGCUGUCUCGACGAAAUUUCACGUUCAACCCUUGUUAUCGUUGUGACCCCAAACCAAAG----UUCCUCG---UCAGUUGGCCGCCAGCGCUUCGGCGCUGUGCAUCGGU .......((((((((...(((.(((...(((((...(((....))).....))))).............----))).)))---.))))).)))((((((((....)))))).))...... ( -33.49) >DroYak_CAF1 911 113 - 1 AAAAGCAGGCAGCUGUCGCGACGAAAUUUCACGUUCAACCCUUGUUAUCGUUGUAACCACGAACCAAAG----UUCCUUG---UCAGUUGGCCGCCAGCGCUUCGGCGCUGUGCAUCGGU .......((((((((.(((((((((((................))).)))))))......((((....)----)))...)---.))))).)))((((((((....)))))).))...... ( -37.39) >DroPer_CAF1 5800 109 - 1 AGCGGCUAGCAGCUGGCC---AAAGAU------UUCAACCCUCGCUGGGGUUUC--CUCUGGGCCACAGCGGACUCAUCGGCCACAGUUGGCUGCCAGUGCUCCGGCGCUGAGCAUCGCC .(((((((((.(.(((((---......------...((((((....))))))((--(.(((.....))).)))......)))))).))))))(((((((((....)))))).))).))). ( -47.10) >consensus AAAAGCAGGCAGCUGCCUC_AAAAAAUUUCAC_UUCAACCCUCGUUAUCGUUGUGACCACGAACCAAAG____UUCCUCG___UCAGUUGGCCGCCAGCGCUUCGGCGCUGUGCAUCGGU ....(((((((((((............................(((((....)))))...(((..........)))........))))).)))..((((((....)))))))))...... (-24.98 = -22.98 + -1.99)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:57:59 2006