
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 8,996,334 – 8,996,454 |
| Length | 120 |
| Max. P | 0.951398 |

| Location | 8,996,334 – 8,996,454 |
|---|---|
| Length | 120 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 95.28 |
| Mean single sequence MFE | -42.92 |
| Consensus MFE | -35.90 |
| Energy contribution | -37.40 |
| Covariance contribution | 1.50 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.67 |
| Structure conservation index | 0.84 |
| SVM decision value | -0.04 |
| SVM RNA-class probability | 0.514737 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 8996334 120 + 22224390 UCUGCCGCCGCCACCUCCGCACACGGGUUAUGCUAACUACGGCGGACCACCACAUGGCCCGCCGCCUGGACCACCCGGUGGACCAGCACGACCCUAUUACCAGCCCCAGUACGGCGGACA (((((((..((.......))....((((..((((.....((((((.(((.....))).))))))...((.((((...)))).))))))..)))).................))))))).. ( -47.10) >DroSec_CAF1 8106 120 + 1 UCUGCCGCCGCCACCUCCGCACACGGGUUAUGCCAACUACGGCGGACCACCACAUGGCCCGCCGCCUGGACCACCCGGUGGACCAGCACGACCCUAUUACCAGCCCCAGUACGGCGGACA (((((((..((.......))....((((..(((......((((((.(((.....))).))))))..(((.((((...)))).))))))..)))).................))))))).. ( -46.30) >DroEre_CAF1 8254 120 + 1 UCUACCGCCGCCACCUCCGCACACGGGUUAUGCCAACUACGGCGGACCACCACAUGGCCCGCCACCUGGACCGCCCGGAGGACCAGCACGACCAUAUUACCAGCCCCAGUACGGAGGACA .............((((((.((..(((((...........(((((.(((.....))).)))))..((((.((.......)).))))...............)))))..)).))))))... ( -40.60) >DroYak_CAF1 9860 120 + 1 UCUGCCGCCGCCACCUCCGCACACGGGUUAUGCCAACUACGGCGGACCACCACAUGGGCCACCGCCUGGACCACCCGGAGGACCAGCAAGACCGUAUUACCAGCCCCAGUACGGAGGACA .............((((((.((..(((((..(((......)))((.((.((....((((....))))((.....)))).)).)).................)))))..)).))))))... ( -37.70) >consensus UCUGCCGCCGCCACCUCCGCACACGGGUUAUGCCAACUACGGCGGACCACCACAUGGCCCGCCGCCUGGACCACCCGGAGGACCAGCACGACCCUAUUACCAGCCCCAGUACGGAGGACA .............((((((.((..(((((..........((((((.(((.....))).)))))).((((.((((...)))).))))...............)))))..)).))))))... (-35.90 = -37.40 + 1.50)

| Location | 8,996,334 – 8,996,454 |
|---|---|
| Length | 120 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 95.28 |
| Mean single sequence MFE | -60.88 |
| Consensus MFE | -55.14 |
| Energy contribution | -55.83 |
| Covariance contribution | 0.69 |
| Combinations/Pair | 1.08 |
| Mean z-score | -2.31 |
| Structure conservation index | 0.91 |
| SVM decision value | 1.41 |
| SVM RNA-class probability | 0.951398 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
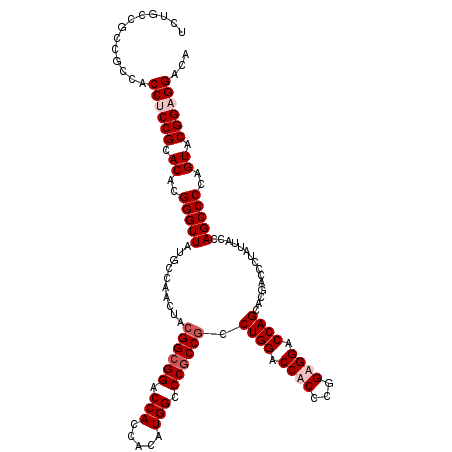
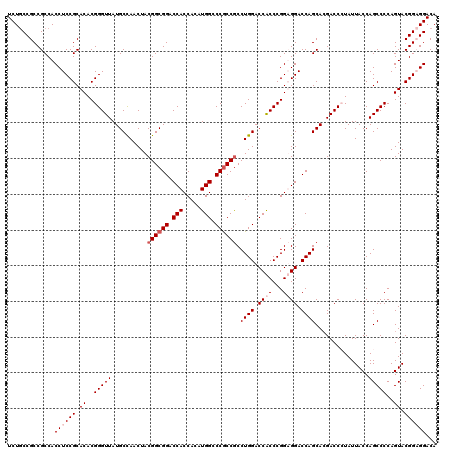
>X_DroMel_CAF1 8996334 120 - 22224390 UGUCCGCCGUACUGGGGCUGGUAAUAGGGUCGUGCUGGUCCACCGGGUGGUCCAGGCGGCGGGCCAUGUGGUGGUCCGCCGUAGUUAGCAUAACCCGUGUGCGGAGGUGGCGGCGGCAGA ((.((((((((((....(((.((...((((..((((((.((((...)))).))..((((((((((((...))))))))))))....))))..))))...))))).))).))))))))).. ( -64.10) >DroSec_CAF1 8106 120 - 1 UGUCCGCCGUACUGGGGCUGGUAAUAGGGUCGUGCUGGUCCACCGGGUGGUCCAGGCGGCGGGCCAUGUGGUGGUCCGCCGUAGUUGGCAUAACCCGUGUGCGGAGGUGGCGGCGGCAGA ((.((((((((((....(((.((...((((..((((((.((((...)))).))..((((((((((((...))))))))))))....))))..))))...))))).))).))))))))).. ( -64.10) >DroEre_CAF1 8254 120 - 1 UGUCCUCCGUACUGGGGCUGGUAAUAUGGUCGUGCUGGUCCUCCGGGCGGUCCAGGUGGCGGGCCAUGUGGUGGUCCGCCGUAGUUGGCAUAACCCGUGUGCGGAGGUGGCGGCGGUAGA .(((((((((((((((..((.((((.(((((((.((((....)))))))).)))...((((((((((...))))))))))...)))).))...)))).)))))))))..))......... ( -57.00) >DroYak_CAF1 9860 120 - 1 UGUCCUCCGUACUGGGGCUGGUAAUACGGUCUUGCUGGUCCUCCGGGUGGUCCAGGCGGUGGCCCAUGUGGUGGUCCGCCGUAGUUGGCAUAACCCGUGUGCGGAGGUGGCGGCGGCAGA ...(((((((((.(((((..((((.......))))..))))).(((((.(.(((.(((((((.((((...)))).)))))))...))))...))))).)))))))))..((....))... ( -58.30) >consensus UGUCCGCCGUACUGGGGCUGGUAAUAGGGUCGUGCUGGUCCACCGGGUGGUCCAGGCGGCGGGCCAUGUGGUGGUCCGCCGUAGUUGGCAUAACCCGUGUGCGGAGGUGGCGGCGGCAGA ...(((((((((..((((..(((.........)))..))))..(((((.(((...((((((((((((...))))))))))))....)))...))))).)))))))))..((....))... (-55.14 = -55.83 + 0.69)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:57:52 2006