
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 1,055,031 – 1,055,122 |
| Length | 91 |
| Max. P | 0.998597 |

| Location | 1,055,031 – 1,055,122 |
|---|---|
| Length | 91 |
| Sequences | 5 |
| Columns | 93 |
| Reading direction | forward |
| Mean pairwise identity | 91.41 |
| Mean single sequence MFE | -28.92 |
| Consensus MFE | -25.76 |
| Energy contribution | -27.24 |
| Covariance contribution | 1.48 |
| Combinations/Pair | 1.06 |
| Mean z-score | -2.27 |
| Structure conservation index | 0.89 |
| SVM decision value | 3.15 |
| SVM RNA-class probability | 0.998597 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
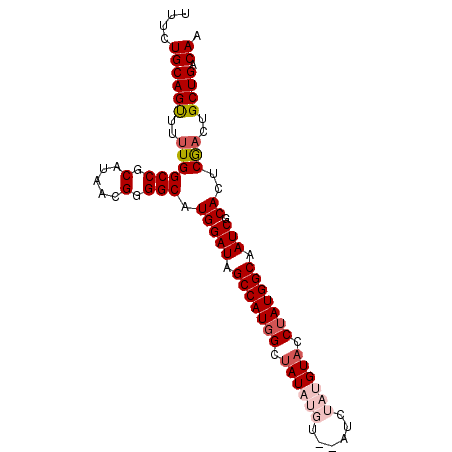
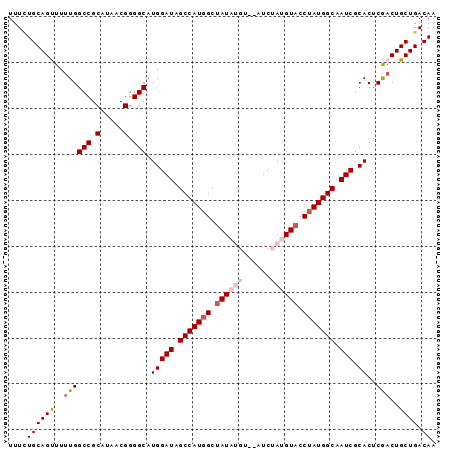
>X_DroMel_CAF1 1055031 91 + 22224390 UUUCUGCAGUUUUUGGCCGCAUAACGGGGCAUGGAUAGCCAUGGCUAUAUGU--AUCUAUGUACCUAUGGCAAUCGCACUCGACUGCUGACAA ..((.((((((....(((.(.....).))).(((((.(((((((.((((((.--...)))))).))))))).))).))...)))))).))... ( -31.30) >DroSec_CAF1 3975 93 + 1 UUUGUGCAGUUUUUGGCCGCAUAACGGGGCAUGGAUAGCCAUGGCUAUAUGUAUAUCUAUGUACCUAUGGCAAUCGCACUCGACUGCUGACAA .((((.((((..((((((.(.....).))).(((((.(((((((.((((((......)))))).))))))).))).))..)))..)))))))) ( -31.00) >DroSim_CAF1 4910 93 + 1 UUUCUGCAGUUUUUGGCCGCAUAACGGGGCAUGGAUAGCCAUGGCUAUAUGUAUAUCUAUGUACCUAUGGCAAUCGCACUCGACUGCUGACAA ..((.((((((....(((.(.....).))).(((((.(((((((.((((((......)))))).))))))).))).))...)))))).))... ( -31.50) >DroEre_CAF1 3998 87 + 1 UUUUUGCAGCUUUUGGCCGCACAACGGGGCAUGGAUAGCCAUUGCAAU------AUCUACGUACCUAUGGCAAUCGCACUCGACUGCUGACAA ...(((((((..((((((.(.....).))).(((((.((((((((...------......)))...))))).))).))..)))..)))).))) ( -23.10) >DroYak_CAF1 6536 87 + 1 UUUCUGCAGUUUUUGGCCGCAUUGCGGGGCAUGGAUAGCCAUGGCUAU------AUCUAUGUACCUAUGGCAAUCGCACUCACCUGCUGACAA ....((((((...(((..(((((((((((((((((((((....)))).------.))))))).)))...))))).))..)))...)))).)). ( -27.70) >consensus UUUCUGCAGUUUUUGGCCGCAUAACGGGGCAUGGAUAGCCAUGGCUAUAUGU__AUCUAUGUACCUAUGGCAAUCGCACUCGACUGCUGACAA ....((((((..((((((.(.....).))).(((((.(((((((.((((((......)))))).))))))).))).))..)))..)))).)). (-25.76 = -27.24 + 1.48)

| Location | 1,055,031 – 1,055,122 |
|---|---|
| Length | 91 |
| Sequences | 5 |
| Columns | 93 |
| Reading direction | reverse |
| Mean pairwise identity | 91.41 |
| Mean single sequence MFE | -24.52 |
| Consensus MFE | -21.65 |
| Energy contribution | -21.77 |
| Covariance contribution | 0.12 |
| Combinations/Pair | 1.09 |
| Mean z-score | -1.23 |
| Structure conservation index | 0.88 |
| SVM decision value | 0.14 |
| SVM RNA-class probability | 0.603100 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
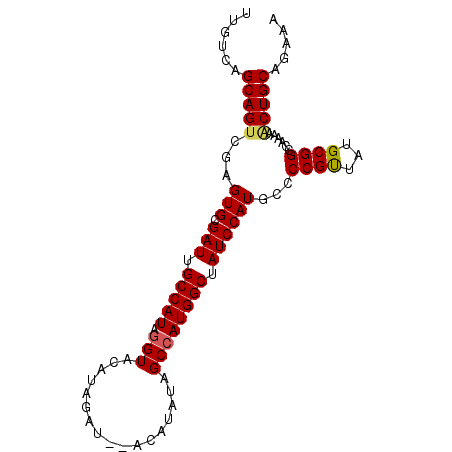
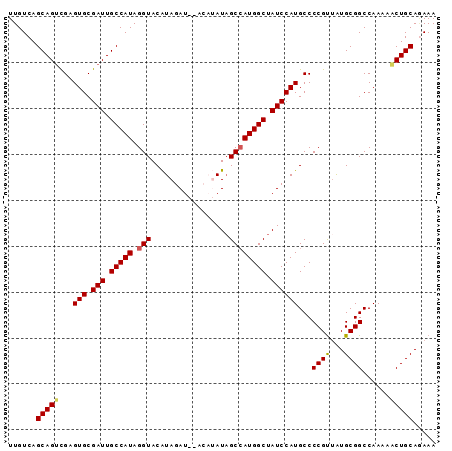
>X_DroMel_CAF1 1055031 91 - 22224390 UUGUCAGCAGUCGAGUGCGAUUGCCAUAGGUACAUAGAU--ACAUAUAGCCAUGGCUAUCCAUGCCCCGUUAUGCGGCCAAAAACUGCAGAAA ......(((((...(((.(((.(((((.(((..(((...--...))).)))))))).))))))...(((.....)))......)))))..... ( -24.30) >DroSec_CAF1 3975 93 - 1 UUGUCAGCAGUCGAGUGCGAUUGCCAUAGGUACAUAGAUAUACAUAUAGCCAUGGCUAUCCAUGCCCCGUUAUGCGGCCAAAAACUGCACAAA ((((..(((((...(((.(((.(((((.(((..(((........))).)))))))).))))))...(((.....)))......))))))))). ( -24.40) >DroSim_CAF1 4910 93 - 1 UUGUCAGCAGUCGAGUGCGAUUGCCAUAGGUACAUAGAUAUACAUAUAGCCAUGGCUAUCCAUGCCCCGUUAUGCGGCCAAAAACUGCAGAAA ......(((((...(((.(((.(((((.(((..(((........))).)))))))).))))))...(((.....)))......)))))..... ( -24.10) >DroEre_CAF1 3998 87 - 1 UUGUCAGCAGUCGAGUGCGAUUGCCAUAGGUACGUAGAU------AUUGCAAUGGCUAUCCAUGCCCCGUUGUGCGGCCAAAAGCUGCAAAAA ......(((((((....)))))))....((..(((.(((------(..((....)))))).)))..))....((((((.....)))))).... ( -24.90) >DroYak_CAF1 6536 87 - 1 UUGUCAGCAGGUGAGUGCGAUUGCCAUAGGUACAUAGAU------AUAGCCAUGGCUAUCCAUGCCCCGCAAUGCGGCCAAAAACUGCAGAAA ...((.((.((.(.(((.(((.(((((.(((........------...)))))))).)))))).))).))..(((((.......))))))).. ( -24.90) >consensus UUGUCAGCAGUCGAGUGCGAUUGCCAUAGGUACAUAGAU__ACAUAUAGCCAUGGCUAUCCAUGCCCCGUUAUGCGGCCAAAAACUGCAGAAA ......(((((...(((.(((.(((((.(((.................)))))))).))))))...((((...))))......)))))..... (-21.65 = -21.77 + 0.12)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 09:42:03 2006