
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 8,934,871 – 8,935,029 |
| Length | 158 |
| Max. P | 0.941432 |

| Location | 8,934,871 – 8,934,989 |
|---|---|
| Length | 118 |
| Sequences | 5 |
| Columns | 118 |
| Reading direction | reverse |
| Mean pairwise identity | 92.19 |
| Mean single sequence MFE | -31.46 |
| Consensus MFE | -24.84 |
| Energy contribution | -25.44 |
| Covariance contribution | 0.60 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.15 |
| Structure conservation index | 0.79 |
| SVM decision value | 1.32 |
| SVM RNA-class probability | 0.941432 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 8934871 118 - 22224390 CACGUGCAAGCCGCCUGUCAAAAUGGCUUGCGCUUCGUUCACACACACACCCGGCAAAUGAAAACGCCGAAUUGGCCACAAUUAUCAAUGAAAAGUGGGCGAAUUCACGCAACGCCAA ...((((((((((..........))))))))))(((((((((.........((((..........))))(((((....)))))...........)))))))))............... ( -35.50) >DroSec_CAF1 5146 118 - 1 CACGUGCAAGCCGCCUGUCAAAAUGGCUUGCGCUUCGUUCACACACACACCCGGCAAAUGAAAACGCCGAAUUGGCCACAAUUAUCAAUGAAAAUACGGCGAAUUCACGCAACGCCAA ...((((((((((..........))))))))))...................(((...((((..((((((((((....)))))....((....)).)))))..))))......))).. ( -32.80) >DroSim_CAF1 5379 118 - 1 CACGUGCAAGCCGCCUGUCAAAAUGGCUUGCGCUUCGUUCACACACACACCCGGCAAAUGAAAACGCCGAAUUGGCCACAAUUAUCAAUGAAAAUAUGGCGAAUUCACGCAACGCCAA ...((((((((((..........))))))))))((((((............((((..........))))(((((....)))))...))))))....(((((...........))))). ( -32.50) >DroEre_CAF1 4171 116 - 1 CACGUGCAAGCCGCCUGUCAAAAUGGCUUGCGCUUCUAUCACA--CACACCCGGCAAAUGAAAACGCCGAAUUGGCGGAAAUUAUCAAUGAAAACUGUCCAAAGUCACGUAACGCCAC ...((((((((((..........))))))))))..........--......((((..........))))...(((((((.....)).(((...(((......)))..)))..))))). ( -29.10) >DroYak_CAF1 4250 116 - 1 CACGUGCAAGCCGCCUGUCAAAAUGGCUUGCGCUUCGUUCACA--CACACCCGGCAAAUGAAAACGCCGAAUUGGCCGGAAUUAUCAAUGAAAACUGCCCGAAAUCACGCAACGCCAC ...((((((((((..........))))))))))..........--.......(((...(((.((..(((.......)))..)).)))........(((..(....)..)))..))).. ( -27.40) >consensus CACGUGCAAGCCGCCUGUCAAAAUGGCUUGCGCUUCGUUCACACACACACCCGGCAAAUGAAAACGCCGAAUUGGCCACAAUUAUCAAUGAAAACUGGGCGAAUUCACGCAACGCCAA ...((((((((((..........))))))))))...................(((..........(((.....)))......................(((......)))...))).. (-24.84 = -25.44 + 0.60)


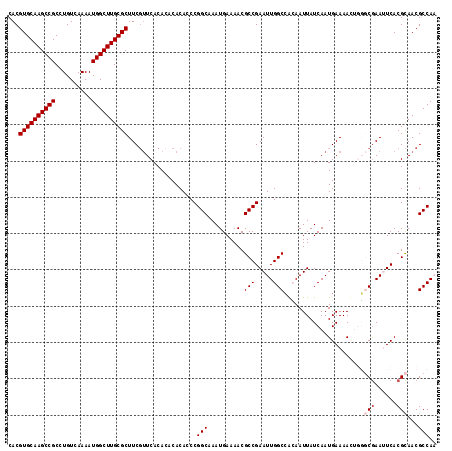
| Location | 8,934,909 – 8,935,029 |
|---|---|
| Length | 120 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 94.07 |
| Mean single sequence MFE | -38.72 |
| Consensus MFE | -32.92 |
| Energy contribution | -33.32 |
| Covariance contribution | 0.40 |
| Combinations/Pair | 1.06 |
| Mean z-score | -1.34 |
| Structure conservation index | 0.85 |
| SVM decision value | 0.27 |
| SVM RNA-class probability | 0.662602 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 8934909 120 - 22224390 AAGGUUCUGGAAGUGGCAGGCGAAAUCGGAGUCCAUGGCUCACGUGCAAGCCGCCUGUCAAAAUGGCUUGCGCUUCGUUCACACACACACCCGGCAAAUGAAAACGCCGAAUUGGCCACA ............(((((.((((((....((((.....))))..((((((((((..........))))))))))))))))............((((..........)))).....))))). ( -40.30) >DroSec_CAF1 5184 120 - 1 AAGGUUCUGGAAGUGGCAGGCGAAAUCGGAGUCCAUGGCUCACGUGCAAGCCGCCUGUCAAAAUGGCUUGCGCUUCGUUCACACACACACCCGGCAAAUGAAAACGCCGAAUUGGCCACA ............(((((.((((((....((((.....))))..((((((((((..........))))))))))))))))............((((..........)))).....))))). ( -40.30) >DroSim_CAF1 5417 120 - 1 AAGGUUCUGGAAGUGGCAGGCGAAAUCGGAGUCCAUGGCUCACGUGCAAGCCGCCUGUCAAAAUGGCUUGCGCUUCGUUCACACACACACCCGGCAAAUGAAAACGCCGAAUUGGCCACA ............(((((.((((((....((((.....))))..((((((((((..........))))))))))))))))............((((..........)))).....))))). ( -40.30) >DroEre_CAF1 4209 117 - 1 AAUGUUCUGGAAGUGA-AACCGAAAUCGGAGUCCAUGGCUCACGUGCAAGCCGCCUGUCAAAAUGGCUUGCGCUUCUAUCACA--CACACCCGGCAAAUGAAAACGCCGAAUUGGCGGAA ..(((..(((((((((-..(((....))).((.....))))))((((((((((..........)))))))))))))))..)))--...................((((.....))))... ( -34.50) >DroYak_CAF1 4288 117 - 1 AAUGUUCUCGAAGUGG-AAGCGAAAUCGGUGUCCAUGGCUCACGUGCAAGCCGCCUGUCAAAAUGGCUUGCGCUUCGUUCACA--CACACCCGGCAAAUGAAAACGCCGAAUUGGCCGGA ............(((.-.((((((....(((.(....)..)))((((((((((..........))))))))))))))))..))--)....(((((.(((...........))).))))). ( -38.20) >consensus AAGGUUCUGGAAGUGGCAGGCGAAAUCGGAGUCCAUGGCUCACGUGCAAGCCGCCUGUCAAAAUGGCUUGCGCUUCGUUCACACACACACCCGGCAAAUGAAAACGCCGAAUUGGCCACA ..((((......((((...(((((....((((.....))))..((((((((((..........))))))))))))))))))).........((((..........))))....))))... (-32.92 = -33.32 + 0.40)


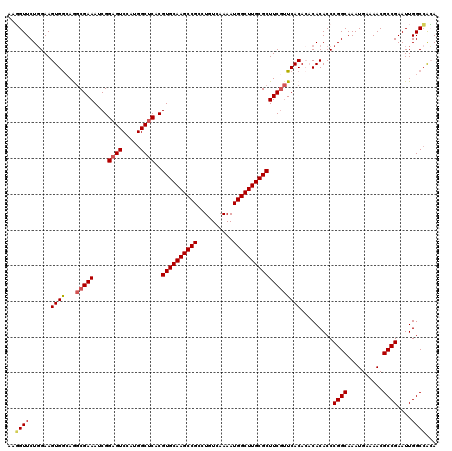
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:56:57 2006