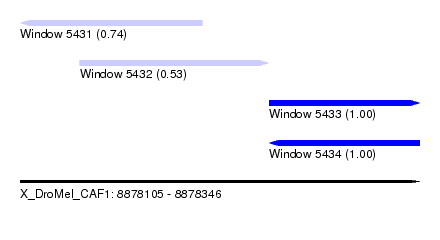
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 8,878,105 – 8,878,346 |
| Length | 241 |
| Max. P | 0.999749 |

| Location | 8,878,105 – 8,878,215 |
|---|---|
| Length | 110 |
| Sequences | 5 |
| Columns | 116 |
| Reading direction | reverse |
| Mean pairwise identity | 79.41 |
| Mean single sequence MFE | -30.03 |
| Consensus MFE | -12.04 |
| Energy contribution | -13.20 |
| Covariance contribution | 1.16 |
| Combinations/Pair | 1.06 |
| Mean z-score | -2.18 |
| Structure conservation index | 0.40 |
| SVM decision value | 0.45 |
| SVM RNA-class probability | 0.740930 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
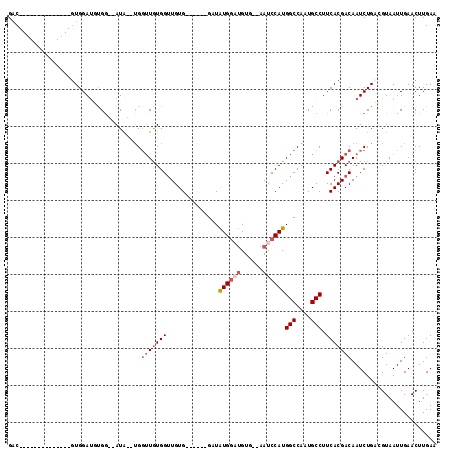
>X_DroMel_CAF1 8878105 110 - 22224390 GACAUGUGGAUGUGGAUGUGGAUGUGA--AUA--UGGUUGUGGUUGUGUAUAUCGAUAUGGUUGUG--AAUCCAUGGCCAAUGCCUUCACGACAAUCUGCCGUAAUUGAACUUGAA .((((.((.((....)).)).))))..--.((--((((.(..((((((..........((((..((--....))..)))).......))))))...).))))))............ ( -24.63) >DroSec_CAF1 8448 90 - 1 GAC--------------GUGGAUUCGG--AUA--UGGUUGUGGUUGUG------GCUAUGGAUGUG--AAUCCAUGGCCAAUGCCUUCACGACAAUCUGAAGUAAUUGAACUUGAA ...--------------.....(((((--((.--..(((((((...((------(((((((((...--.)))))))))))......))))))).)))))))............... ( -33.50) >DroSim_CAF1 8754 90 - 1 GAC--------------GUGGAUUCGG--AUA--UGGUUGUGGUUGUG------GCUAUGGAUGUG--AAUCCAUGGCCAAUGCCUUCACGACAAUCUGAAGUAAUUGAACUUGAA ...--------------.....(((((--((.--..(((((((...((------(((((((((...--.)))))))))))......))))))).)))))))............... ( -33.50) >DroEre_CAF1 7914 94 - 1 GAC--------------GUGGAAGUGGGGAUACAAGCAUAUGGAUGUG------GAUGUGUUUGUG--GAUCCAUGGCCAAUGCCUUCACGACAAUCUGCCGUAAUUGAACUUGAA ..(--------------(((((.(((((..(((((((((((.......------.)))))))))))--..)))))(((....))))))))).((((........))))........ ( -28.80) >DroYak_CAF1 8788 94 - 1 GUC--------------UUGGAUGUGGCCAUU--UGGUUGUGGUUGUG------GAUGUGGACGUGGAGAUCCAUGGCCAAUGCCUUCACGACAAUCUGGCGUAAUUGAACUUGAA ...--------------((.(((...((((..--..(((((((....(------(...(((.(((((....))))).)))...)).)))))))....))))...))).))...... ( -29.70) >consensus GAC______________GUGGAUGUGG__AUA__UGGUUGUGGUUGUG______GAUAUGGAUGUG__AAUCCAUGGCCAAUGCCUUCACGACAAUCUGACGUAAUUGAACUUGAA ....................................(((((((..............((((((......))))))(((....))).)))))))....................... (-12.04 = -13.20 + 1.16)


| Location | 8,878,141 – 8,878,255 |
|---|---|
| Length | 114 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 80.38 |
| Mean single sequence MFE | -16.75 |
| Consensus MFE | -10.72 |
| Energy contribution | -11.16 |
| Covariance contribution | 0.44 |
| Combinations/Pair | 1.08 |
| Mean z-score | -1.63 |
| Structure conservation index | 0.64 |
| SVM decision value | -0.00 |
| SVM RNA-class probability | 0.532327 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 8878141 114 + 22224390 UGGCCAUGGAUU--CACAACCAUAUCGAUAUACACAACCACAACCA--UAU--UCACAUCCACAUCCACAUCCACAUGUCCAUGUUGGUCAUCAGCAACUUUCACUCACCGGCUAAAUGA (((((((((...--.....)))).......................--...--......((((((..((((....))))..))).)))......................)))))..... ( -14.80) >DroSec_CAF1 8484 94 + 1 UGGCCAUGGAUU--CACAUCCAUAGC------CACAACCACAACCA--UAU--CCGAAUCCAC--------------GUCCAUGUUGGUCAUCAGCAACUUUCACUCACCGGCAAAAUGA ((((.((((((.--...)))))).))------))............--...--.........(--------------((...(((((((..................)))))))..))). ( -17.87) >DroSim_CAF1 8790 94 + 1 UGGCCAUGGAUU--CACAUCCAUAGC------CACAACCACAACCA--UAU--CCGAAUCCAC--------------GUCCAUGUUGGUCAUCAGCAACUUUCACUCACCGGCAAGCUGA ((((.((((((.--...)))))).))------))............--...--..........--------------(.((.....)).).(((((...................))))) ( -18.51) >DroEre_CAF1 7950 98 + 1 UGGCCAUGGAUC--CACAAACACAUC------CACAUCCAUAUGCUUGUAUCCCCACUUCCAC--------------GUCCAUGUUGGUCAUCAGCAACUUUCACUCACCGGCAAAAUGA ..(((((((((.--............------...)))))).((((.(....((.((......--------------......)).))....))))).............)))....... ( -13.99) >DroYak_CAF1 8824 98 + 1 UGGCCAUGGAUCUCCACGUCCACAUC------CACAACCACAACCA--AAUGGCCACAUCCAA--------------GACCAUGUUGGUCAUCAGCAACUUUCACUCACCGGCAAAAUGA ..(((.(((((......)))))....------..............--.((((((((((....--------------....))).)))))))..................)))....... ( -18.60) >consensus UGGCCAUGGAUU__CACAUCCAUAUC______CACAACCACAACCA__UAU__CCACAUCCAC______________GUCCAUGUUGGUCAUCAGCAACUUUCACUCACCGGCAAAAUGA ..(((.(((((......)))))............................................................(((((.....))))).............)))....... (-10.72 = -11.16 + 0.44)

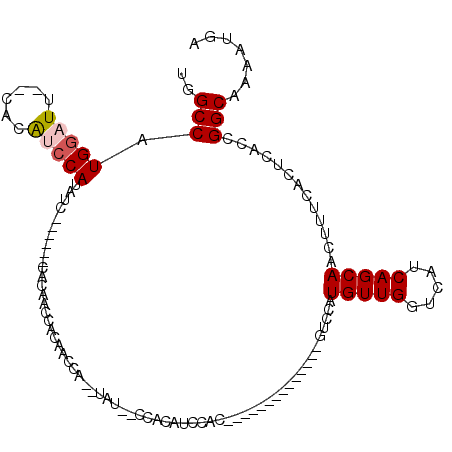
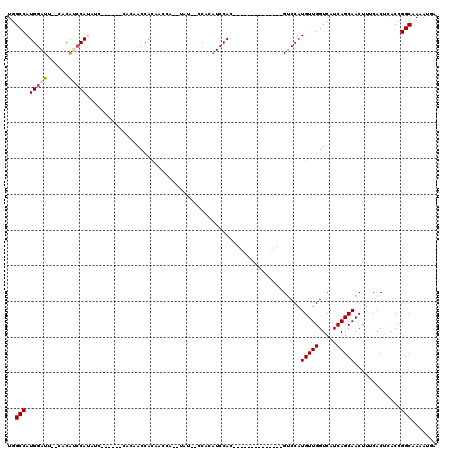
| Location | 8,878,255 – 8,878,346 |
|---|---|
| Length | 91 |
| Sequences | 5 |
| Columns | 95 |
| Reading direction | forward |
| Mean pairwise identity | 92.70 |
| Mean single sequence MFE | -27.86 |
| Consensus MFE | -23.62 |
| Energy contribution | -24.42 |
| Covariance contribution | 0.80 |
| Combinations/Pair | 1.00 |
| Mean z-score | -3.81 |
| Structure conservation index | 0.85 |
| SVM decision value | 4.00 |
| SVM RNA-class probability | 0.999749 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 8878255 91 + 22224390 CGAUGACGAUGUGUGCCUCCUGCACAUUUUGUCAUCGUUUUUCAUGUGGGAAAAUCUCUCGCCUCUGCAAAAGAUCCAA----AGAAAAAAAGAC (((((((((.((((((.....)))))).)))))))))((((((..((((((.....)))))).(((.....))).....----.))))))..... ( -30.50) >DroSec_CAF1 8578 94 + 1 CGAUGACGAUGUGUGCCUCCUGCACAUUUUGUCAUCGUUUUUCAUGUGGGAAAAUCUCUCGCCUCUGCAAAAGAUCCAAA-AAAGAAAAAAAGAC (((((((((.((((((.....)))))).)))))))))((((((..((((((.....)))))).(((.....)))......-...))))))..... ( -30.50) >DroSim_CAF1 8884 95 + 1 CGAUGACGAUGUGUGCCUCCUGCACAUUUUGUCAUCGUUUUUCAUGUGGGAAAAUCUCUCGCCUCUGCAAAAGAUCCAAAAAAAGAAAAAAAUAC (((((((((.((((((.....)))))).)))))))))((((((..((((((.....)))))).(((.....)))..........))))))..... ( -30.50) >DroEre_CAF1 8048 84 + 1 CGAUGACGAUG--UGCCUCCUGCACAUUUUGUCAUCGUUUUUCAUGUGGGAAAAUCUCUCGCCUCUGCAAAAGAUCCAAA---------AAAGAC .((((((((((--(((.....)))))))..))))))((((((...((((((.....)))))).(((.....)))......---------)))))) ( -23.90) >DroYak_CAF1 8922 86 + 1 CGAUGACGAUG--UGCCUCCUGCACAUUUUGUCAUCGUUUUUCAUGUGGGAAAAUCUCUCGCCUCUGCAAAAGAUCCAAA-------AAAAAGAC .((((((((((--(((.....)))))))..))))))((((((...((((((.....)))))).(((.....)))......-------..)))))) ( -23.90) >consensus CGAUGACGAUGUGUGCCUCCUGCACAUUUUGUCAUCGUUUUUCAUGUGGGAAAAUCUCUCGCCUCUGCAAAAGAUCCAAA___AGAAAAAAAGAC (((((((((.((((((.....)))))).)))))))))........((((((.....)))))).(((.....)))..................... (-23.62 = -24.42 + 0.80)



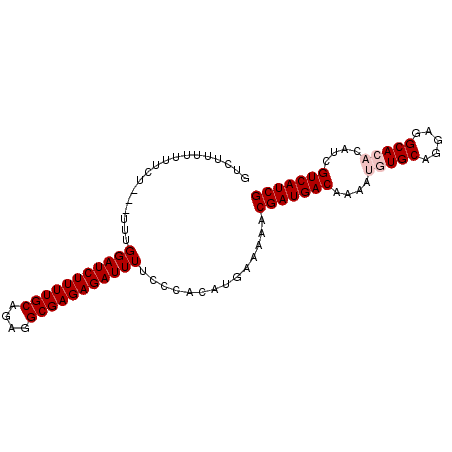
| Location | 8,878,255 – 8,878,346 |
|---|---|
| Length | 91 |
| Sequences | 5 |
| Columns | 95 |
| Reading direction | reverse |
| Mean pairwise identity | 92.70 |
| Mean single sequence MFE | -29.50 |
| Consensus MFE | -24.10 |
| Energy contribution | -24.90 |
| Covariance contribution | 0.80 |
| Combinations/Pair | 1.00 |
| Mean z-score | -3.22 |
| Structure conservation index | 0.82 |
| SVM decision value | 3.11 |
| SVM RNA-class probability | 0.998477 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 8878255 91 - 22224390 GUCUUUUUUUCU----UUGGAUCUUUUGCAGAGGCGAGAGAUUUUCCCACAUGAAAAACGAUGACAAAAUGUGCAGGAGGCACACAUCGUCAUCG .....((((((.----.((((((((((((....)))))))))....)))...))))))(((((((....(((((.....)))))....))))))) ( -30.30) >DroSec_CAF1 8578 94 - 1 GUCUUUUUUUCUUU-UUUGGAUCUUUUGCAGAGGCGAGAGAUUUUCCCACAUGAAAAACGAUGACAAAAUGUGCAGGAGGCACACAUCGUCAUCG .....((((((...-...(((((((((((....)))))))))))........))))))(((((((....(((((.....)))))....))))))) ( -29.94) >DroSim_CAF1 8884 95 - 1 GUAUUUUUUUCUUUUUUUGGAUCUUUUGCAGAGGCGAGAGAUUUUCCCACAUGAAAAACGAUGACAAAAUGUGCAGGAGGCACACAUCGUCAUCG .....((((((.......(((((((((((....)))))))))))........))))))(((((((....(((((.....)))))....))))))) ( -30.36) >DroEre_CAF1 8048 84 - 1 GUCUUU---------UUUGGAUCUUUUGCAGAGGCGAGAGAUUUUCCCACAUGAAAAACGAUGACAAAAUGUGCAGGAGGCA--CAUCGUCAUCG ((.(((---------(.((((((((((((....)))))))))....)))...)))).))((((((...((((((.....)))--))).)))))). ( -28.10) >DroYak_CAF1 8922 86 - 1 GUCUUUUU-------UUUGGAUCUUUUGCAGAGGCGAGAGAUUUUCCCACAUGAAAAACGAUGACAAAAUGUGCAGGAGGCA--CAUCGUCAUCG ((.((((.-------..((((((((((((....)))))))))....)))...)))).))((((((...((((((.....)))--))).)))))). ( -28.80) >consensus GUCUUUUUUUCU___UUUGGAUCUUUUGCAGAGGCGAGAGAUUUUCCCACAUGAAAAACGAUGACAAAAUGUGCAGGAGGCACACAUCGUCAUCG ..................(((((((((((....)))))))))))..............(((((((....(((((.....)))))....))))))) (-24.10 = -24.90 + 0.80)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:56:23 2006