
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 8,872,026 – 8,872,207 |
| Length | 181 |
| Max. P | 0.807682 |

| Location | 8,872,026 – 8,872,146 |
|---|---|
| Length | 120 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 89.71 |
| Mean single sequence MFE | -47.06 |
| Consensus MFE | -38.68 |
| Energy contribution | -37.96 |
| Covariance contribution | -0.72 |
| Combinations/Pair | 1.20 |
| Mean z-score | -1.44 |
| Structure conservation index | 0.82 |
| SVM decision value | 0.25 |
| SVM RNA-class probability | 0.653100 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
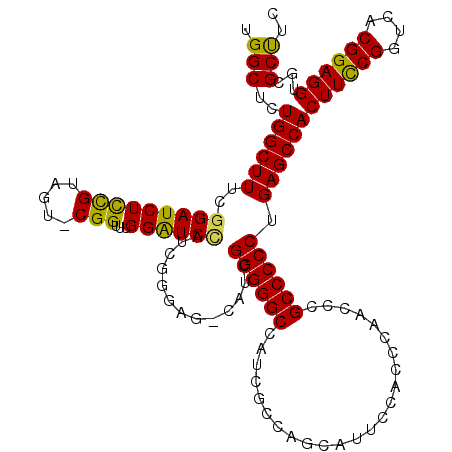
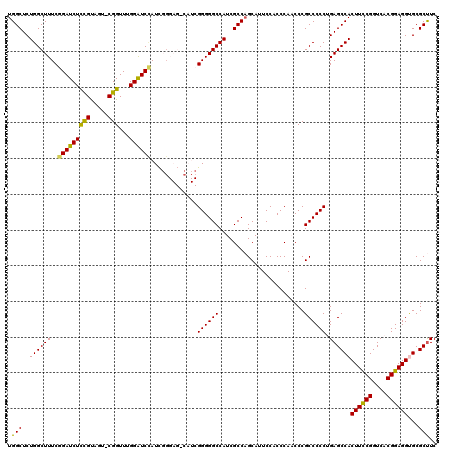
>X_DroMel_CAF1 8872026 120 + 22224390 UGGCUCUGGCUUUCGGAUCUCCGCAGUGCGGUUUGGAUCUAUCGGGAGACAUCGGGGGCCAUCGCCAGCAUUCCACCCAACCCGCCCCCUGAGCCACUUCCGGUCACGGAGGCGCGCUUC ((((..(((((..(((((((((((...))))...))))).....(....)..))..)))))..)))).......................(((((.((((((....)))))).).)))). ( -45.40) >DroSec_CAF1 2438 119 + 1 UGGCUCUGGCUUUCGGAUCUCCGUAGU-CGGUUUGGAUCCAUCGGGAGCCAUCGGGGGCCAUCGCCAGCAUUCCACCCAUCCCGCCCCCAGAGCCACUUUCGGUCACGGAGGUGCGCUUC .((((((((((..((((((((((....-)))...))))))...)..))))...((((((........................))))))))))))(((((((....)))))))....... ( -50.36) >DroSim_CAF1 2762 119 + 1 UGGCUCUGGCUUUCGGAUCUCCGUAGU-CGGUUUGGAUCCAUCGGGAGCCAUCGGGGGCCAUCGCCAGCAUUCCACCCAUCCCGCCCCCAGAGCCACUUCCGGUCACGGAGGUGCGCCUC .((((((((((..((((((((((....-)))...))))))...)..))))...((((((........................))))))))))))(((((((....)))))))....... ( -53.26) >DroEre_CAF1 2122 109 + 1 UGGCUCUGGCUUUCGGAUCUCUGUAGU-CGGUUUGGGUCAAUU----------GGGGGCCAUCGCCAGCAUUCCACCCAACCCGCCCCCUGAGCCACUUCCGGUCACGGAGGUGCGCUUC .(((...((((..(((....))).)))-)((.((((((....(----------(..(((....)))..))....)))))).)))))....((((((((((((....)))))))).)))). ( -45.30) >DroYak_CAF1 2610 111 + 1 UGGCUCUGGCUUUCGGAUCUUCGUAGU-CGGUUUGGAUCCAUCCG--------GGGGGCCAUCGCCUGCAUUCCACCCAACCCGCCCCCUGAGCCACUUCCGGUCACGGAGGCGCGCUUC .(((.((((((...(((((((((....-)))...))))))...((--------((((((........................)))))))))))))((((((....)))))).).))).. ( -40.96) >consensus UGGCUCUGGCUUUCGGAUCUCCGUAGU_CGGUUUGGAUCCAUCGGGAG_CAUCGGGGGCCAUCGCCAGCAUUCCACCCAACCCGCCCCCUGAGCCACUUCCGGUCACGGAGGUGCGCUUC .(((..((((((..(((((((((.....)))...)))))).............((((((........................)))))).))))))((((((....))))))...))).. (-38.68 = -37.96 + -0.72)

| Location | 8,872,106 – 8,872,207 |
|---|---|
| Length | 101 |
| Sequences | 4 |
| Columns | 103 |
| Reading direction | forward |
| Mean pairwise identity | 92.48 |
| Mean single sequence MFE | -34.17 |
| Consensus MFE | -29.58 |
| Energy contribution | -30.32 |
| Covariance contribution | 0.75 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.75 |
| Structure conservation index | 0.87 |
| SVM decision value | 0.64 |
| SVM RNA-class probability | 0.807682 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
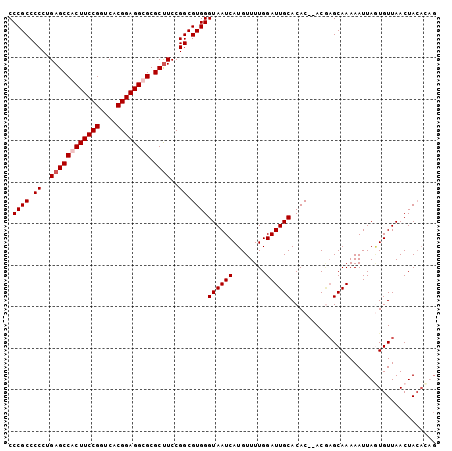
>X_DroMel_CAF1 8872106 101 + 22224390 CCCGCCCCCUGAGCCACUUCCGGUCACGGAGGCGCGCUUCCGGCGUGGGUAAUCAUGUUUUGGAUUGCACAC--ACGAGCAAAAAUUAAUGUUAACUACACAG .((((.((..(((((.((((((....)))))).).))))..)).))))((((((........))))))....--...((((........)))).......... ( -29.90) >DroSim_CAF1 2841 101 + 1 CCCGCCCCCAGAGCCACUUCCGGUCACGGAGGUGCGCCUCCGGCGUGGGUAAUCAUGUUUUGGAUUGCACAC--ACGAGCAAAAAUUAGUGUUAACUACACAG ...((..((.((((((((((((....)))))))).).))).))((((.((((((........))))))...)--))).))........((((.....)))).. ( -35.70) >DroEre_CAF1 2191 103 + 1 CCCGCCCCCUGAGCCACUUCCGGUCACGGAGGUGCGCUUCCGGCGUGGGUAAUCAUGUUUUGGAUUGCACACACACACGCAAAAAUUAGUGUUAACUAAUGUU .((((.((..((((((((((((....)))))))).))))..)).))))((((((........))))))................((((((....))))))... ( -37.20) >DroYak_CAF1 2681 101 + 1 CCCGCCCCCUGAGCCACUUCCGGUCACGGAGGCGCGCUUCCGGCGUGGGUAAUCAUGUUUUGGAUUGCACAC--ACUAGCAAAAAUUACUGUUAACUACAGUG .((((.((..(((((.((((((....)))))).).))))..)).))))((((((........))))))....--............((((((.....)))))) ( -33.90) >consensus CCCGCCCCCUGAGCCACUUCCGGUCACGGAGGCGCGCUUCCGGCGUGGGUAAUCAUGUUUUGGAUUGCACAC__ACGAGCAAAAAUUAGUGUUAACUACACAG .((((.((..((((((((((((....)))))))).))))..)).))))((((((........))))))................................... (-29.58 = -30.32 + 0.75)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:56:18 2006