
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 8,860,341 – 8,860,442 |
| Length | 101 |
| Max. P | 0.861168 |

| Location | 8,860,341 – 8,860,442 |
|---|---|
| Length | 101 |
| Sequences | 5 |
| Columns | 101 |
| Reading direction | forward |
| Mean pairwise identity | 75.94 |
| Mean single sequence MFE | -22.15 |
| Consensus MFE | -13.32 |
| Energy contribution | -14.28 |
| Covariance contribution | 0.96 |
| Combinations/Pair | 1.14 |
| Mean z-score | -1.85 |
| Structure conservation index | 0.60 |
| SVM decision value | 0.80 |
| SVM RNA-class probability | 0.854816 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 8860341 101 + 22224390 UACAUACAUGUGAGCGUAAUUUAUUUAUUGCGUGUCCAUCCAACAUUCACGCAAUACAGGCGAAAAAAAAAAUGCUAGGGUUUUUACCAUCUCUUUUGUUU .....(((...((((((........(((((((((.............)))))))))...)))................(((....)))..)))...))).. ( -16.62) >DroSec_CAF1 100312 93 + 1 U----GCAUGUGAGCGUAAUUUAUUUAUUGCGUGUCUGUCCAACAUUCACGCAAUAAAGGCGAAA----AAAUGCUAGGGUUUUUACCAUCCCCUUUGUUU .----((((.....(((......(((((((((((..((.....))..))))))))))).)))...----..)))).((((..(.....)..))))...... ( -25.90) >DroSim_CAF1 92173 93 + 1 U----GCAUGUGAGCGUAAUUUAUUUAUUGCGUGUCUGUCCAACAUUCACGCAAUAAAGGCGAAA----AAAUGCUAGGGUUUUUACCAUCUCCUUUGUUU .----((((.....(((......(((((((((((..((.....))..))))))))))).)))...----..))))...(((....)))............. ( -25.10) >DroEre_CAF1 94888 85 + 1 G----ACAUGUGAGCGUAAUUUAUUUAUUGCGUGUCUGUCCAACAUUCACGCAAUAAAGGCGAA------AAUGCUAGGGUUCUUUCCUUCUUUU------ .----.......(((((......(((((((((((..((.....))..)))))))))))......------.)))))((((.....))))......------ ( -22.42) >DroAna_CAF1 83211 77 + 1 -----UCAUGUGAGUGUAAUUUACUUAUUGCGUGCUUGUCCGACAUUCACGCAAUAAAAUCCGGA----UAAGGCCAGGCCAUGUG--------------- -----.((((((.((.(.......((((((((((..((.....))..)))))))))).....((.----.....))).))))))))--------------- ( -20.70) >consensus U____ACAUGUGAGCGUAAUUUAUUUAUUGCGUGUCUGUCCAACAUUCACGCAAUAAAGGCGAAA____AAAUGCUAGGGUUUUUACCAUCUCCUUUGUUU ...........((((........(((((((((((..((.....))..)))))))))))((((..........))))...)))).................. (-13.32 = -14.28 + 0.96)

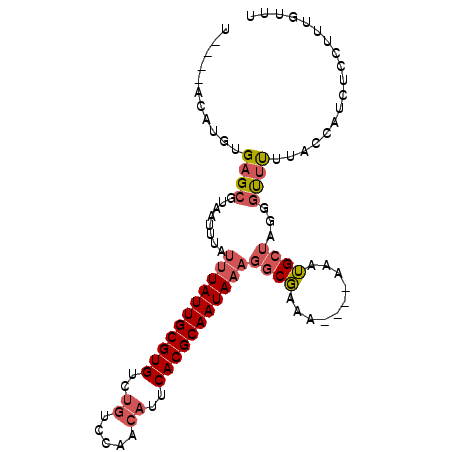

| Location | 8,860,341 – 8,860,442 |
|---|---|
| Length | 101 |
| Sequences | 5 |
| Columns | 101 |
| Reading direction | reverse |
| Mean pairwise identity | 75.94 |
| Mean single sequence MFE | -20.03 |
| Consensus MFE | -14.12 |
| Energy contribution | -13.84 |
| Covariance contribution | -0.28 |
| Combinations/Pair | 1.23 |
| Mean z-score | -1.60 |
| Structure conservation index | 0.70 |
| SVM decision value | 0.83 |
| SVM RNA-class probability | 0.861168 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
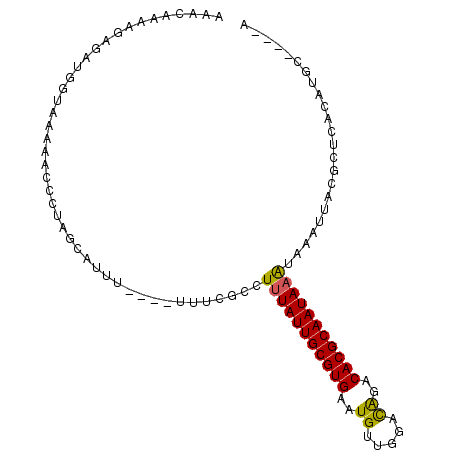
>X_DroMel_CAF1 8860341 101 - 22224390 AAACAAAAGAGAUGGUAAAAACCCUAGCAUUUUUUUUUUCGCCUGUAUUGCGUGAAUGUUGGAUGGACACGCAAUAAAUAAAUUACGCUCACAUGUAUGUA ....((((((((((.((.......)).))))))))))........(((((((((.............)))))))))......................... ( -16.82) >DroSec_CAF1 100312 93 - 1 AAACAAAGGGGAUGGUAAAAACCCUAGCAUUU----UUUCGCCUUUAUUGCGUGAAUGUUGGACAGACACGCAAUAAAUAAAUUACGCUCACAUGC----A ......((((...........)))).((....----....)).(((((((((((..((.....))..)))))))))))........((......))----. ( -22.30) >DroSim_CAF1 92173 93 - 1 AAACAAAGGAGAUGGUAAAAACCCUAGCAUUU----UUUCGCCUUUAUUGCGUGAAUGUUGGACAGACACGCAAUAAAUAAAUUACGCUCACAUGC----A ......((((((((.((.......)).)))))----)))....(((((((((((..((.....))..)))))))))))........((......))----. ( -21.30) >DroEre_CAF1 94888 85 - 1 ------AAAAGAAGGAAAGAACCCUAGCAUU------UUCGCCUUUAUUGCGUGAAUGUUGGACAGACACGCAAUAAAUAAAUUACGCUCACAUGU----C ------......(((.......)))(((...------......(((((((((((..((.....))..)))))))))))........))).......----. ( -17.33) >DroAna_CAF1 83211 77 - 1 ---------------CACAUGGCCUGGCCUUA----UCCGGAUUUUAUUGCGUGAAUGUCGGACAAGCACGCAAUAAGUAAAUUACACUCACAUGA----- ---------------..((((..((((.....----.))))..(((((((((((..((.....))..))))))))))).............)))).----- ( -22.40) >consensus AAACAAAAGAGAUGGUAAAAACCCUAGCAUUU____UUUCGCCUUUAUUGCGUGAAUGUUGGACAGACACGCAAUAAAUAAAUUACGCUCACAUGC____A ...........................................(((((((((((..((.....))..)))))))))))....................... (-14.12 = -13.84 + -0.28)

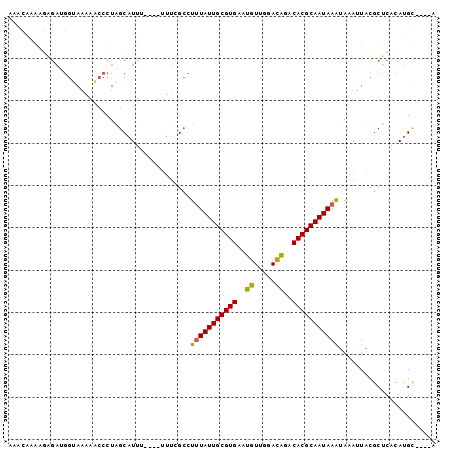
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:56:13 2006