
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 8,791,137 – 8,791,248 |
| Length | 111 |
| Max. P | 0.892074 |

| Location | 8,791,137 – 8,791,248 |
|---|---|
| Length | 111 |
| Sequences | 4 |
| Columns | 117 |
| Reading direction | reverse |
| Mean pairwise identity | 71.71 |
| Mean single sequence MFE | -34.73 |
| Consensus MFE | -19.45 |
| Energy contribution | -21.39 |
| Covariance contribution | 1.94 |
| Combinations/Pair | 1.19 |
| Mean z-score | -1.88 |
| Structure conservation index | 0.56 |
| SVM decision value | 0.97 |
| SVM RNA-class probability | 0.892074 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 8791137 111 - 22224390 ACUUUAUAUUUACCUUGAGCCCGGUCGGGA--GUGCCUGGCAACUUUUGGCAACACGGCUGGGGAAUUUGCGUGCCUUGAAAGCGCCACGA-AAAUACCCCAAUUGGU---UAGAGG ............((((.((((.(((((...--.((((.(....)....))))...)))))((((.((((.(((((((....)).).)))).-)))).))))....)))---).)))) ( -37.00) >DroSec_CAF1 39392 91 - 1 ACUUUUUAUUUACCUUGAGACCGGGUGAGA--GUGCCUGGCAACUUUUGGCAACUCGGCUGGGGAA-----------------------GA-AAUAGCCCCAGUUGGGUGGUAGAGG ........((((((....(.((((((....--..)))))))...........((((((((((((.(-----------------------..-..)..)))))))))))))))))).. ( -35.20) >DroSim_CAF1 32366 88 - 1 ACUUUUUAUUUACCUUGAGACCGGGUGAGA--GUGCCUGGCAACUUUUGGCAACUCGGCUGGGGAA-----------------------GA-AAUAGCCCCAGUUGGG---UAGAGG .((((.......((..(((.((((((....--..))))))...)))..))..((((((((((((.(-----------------------..-..)..)))))))))))---))))). ( -33.40) >DroEre_CAF1 35044 110 - 1 ACUUUUUAUUUACCUUAA-GCCGGGAGAGAGUGUGCCUGGCAAU-UUUGGCAACCCAGCUGUGAAGUUUGCAUGCCUUGGAAGCCCCACGAAAGUAGC-CCGGUUGG----UAGAGG ........((((((...(-((((((......((((...(((..(-(..((((..(.((((....)))).)..))))..))..))).)))).......)-))))))))----)))).. ( -33.32) >consensus ACUUUUUAUUUACCUUGAGACCGGGUGAGA__GUGCCUGGCAACUUUUGGCAACUCGGCUGGGGAA_______________________GA_AAUAGCCCCAGUUGGG___UAGAGG .((((.......((..(((.(((((((......)))))))...)))..))...(((((((((((.................................)))))))))))....)))). (-19.45 = -21.39 + 1.94)

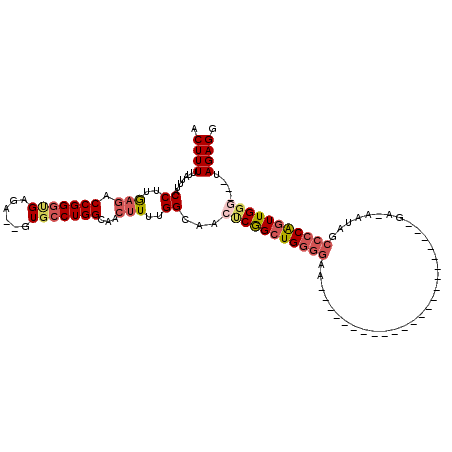

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:55:51 2006