| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 8,760,037 – 8,760,167 |
| Length | 130 |
| Max. P | 0.985107 |

| Location | 8,760,037 – 8,760,127 |
|---|---|
| Length | 90 |
| Sequences | 4 |
| Columns | 111 |
| Reading direction | forward |
| Mean pairwise identity | 81.11 |
| Mean single sequence MFE | -24.78 |
| Consensus MFE | -16.17 |
| Energy contribution | -16.55 |
| Covariance contribution | 0.38 |
| Combinations/Pair | 1.10 |
| Mean z-score | -2.68 |
| Structure conservation index | 0.65 |
| SVM decision value | 2.00 |
| SVM RNA-class probability | 0.985107 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
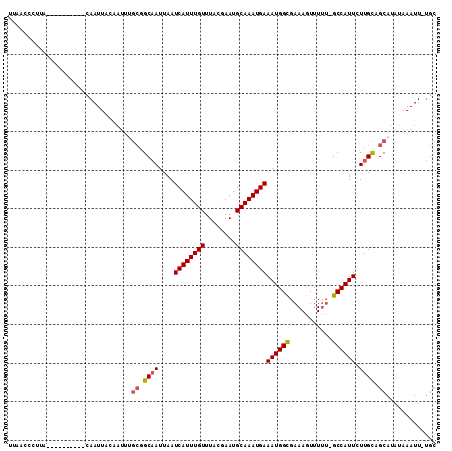
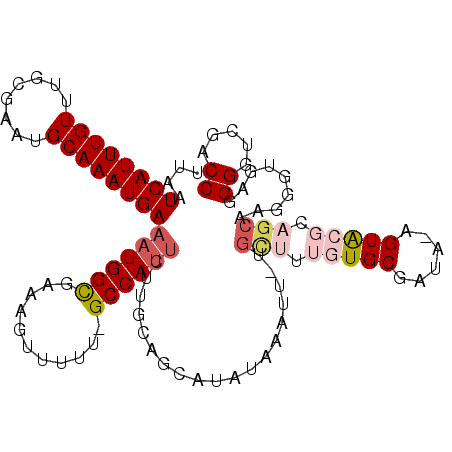
>X_DroMel_CAF1 8760037 90 + 22224390 UUAACCCUAA----------CAAUU----------GCAAUUAAUCAUUUGUUUGCGAAUGCAAAUGAAAUGGCGAAAGUUUUU-GCCAUUUUUGUAGCAUAUAAAUUGUGU .........(----------(((((----------((((.(((....))).))))..((((....(((((((((((....)))-))))))))....))))...)))))).. ( -23.50) >DroSec_CAF1 8047 99 + 1 UUAACCAUUA----------CAAUUACAAUUUGCGGCAAUUAAUCAUUUGUUUACGAAUGCAAAUGAAAUGGCGAAAGUUUUU-GCCAUUCUUGCAGCAUAUAAAUU-UGC ..........----------(((........(((.((((....((((((((........))))))))(((((((((....)))-)))))).)))).))).......)-)). ( -24.36) >DroSim_CAF1 9010 99 + 1 UUAACCCUUA----------CAAUUACAAUUUGCGGCAAUUAAUCAUUUGUUUACGAAUGCAAAUGAAAUGGCGAAAGUUUUU-GCCAUUCUUGCAGCAUAUAAAUU-UGC ..........----------(((........(((.((((....((((((((........))))))))(((((((((....)))-)))))).)))).))).......)-)). ( -24.36) >DroEre_CAF1 7212 108 + 1 UUAACCCGCGUUUGCCGUCUCAAUUACAAUUGGCGGCCAUUAAUCAUUUGUUUGUGAAUGCAAAUGAAAUGGUGAAAGUUUGGUGCCAUUCUUGCAGCAU--AAAUU-CGC .......(((...((((((............))))))......((((((((........)))))))).(((.((.(((..((....))..))).)).)))--.....-))) ( -26.90) >consensus UUAACCCUUA__________CAAUUACAAUUUGCGGCAAUUAAUCAUUUGUUUACGAAUGCAAAUGAAAUGGCGAAAGUUUUU_GCCAUUCUUGCAGCAUAUAAAUU_UGC ................................((.((((....((((((((........))))))))((((((...........)))))).)))).))............. (-16.17 = -16.55 + 0.38)


| Location | 8,760,056 – 8,760,167 |
|---|---|
| Length | 111 |
| Sequences | 5 |
| Columns | 112 |
| Reading direction | forward |
| Mean pairwise identity | 80.44 |
| Mean single sequence MFE | -29.90 |
| Consensus MFE | -15.64 |
| Energy contribution | -17.20 |
| Covariance contribution | 1.56 |
| Combinations/Pair | 1.12 |
| Mean z-score | -2.43 |
| Structure conservation index | 0.52 |
| SVM decision value | 1.08 |
| SVM RNA-class probability | 0.912072 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
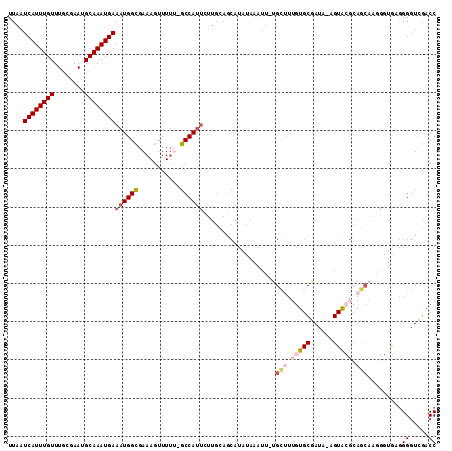
>X_DroMel_CAF1 8760056 111 + 22224390 UUAAUCAUUUGUUUGCGAAUGCAAAUGAAAUGGCGAAAGUUUUU-GCCAUUUUUGUAGCAUAUAAAUUGUGUUUUGUGCGAUAAAGUGCGCAGCAAGGGUGUGGGGUUGACC ((((((....((((((....))))))(((((((((((....)))-)))))))).....(((((...((((....((((((......))))))))))..))))).)))))).. ( -31.00) >DroSec_CAF1 8076 109 + 1 UUAAUCAUUUGUUUACGAAUGCAAAUGAAAUGGCGAAAGUUUUU-GCCAUUCUUGCAGCAUAUAAAUU-UGCUUUGUGCGAUA-AGUACGCAGCAAGGGUGGGGGGUCGACC ....((((((((........))))))))...(((....)))(((-.((((((((((((((........-))))..((((....-.))))...)))))))))).)))...... ( -34.10) >DroSim_CAF1 9039 105 + 1 UUAAUCAUUUGUUUACGAAUGCAAAUGAAAUGGCGAAAGUUUUU-GCCAUUCUUGCAGCAUAUAAAUU-UGCUUUGUGCGAUA-AGUACGCA----GGGUGAGGGGUCGACC ....((((((((........))))))))(((((((((....)))-))))))(((((((((........-))))..((((....-.)))))))----))....((......)) ( -31.30) >DroEre_CAF1 7251 100 + 1 UUAAUCAUUUGUUUGUGAAUGCAAAUGAAAUGGUGAAAGUUUGGUGCCAUUCUUGCAGCAU--AAAUU-CGCCUUGUGCGAUA-AGUA--CAGCAA------GGGGUCGACC ....((((((((........)))))))).(((.((.(((..((....))..))).)).)))--....(-(((((..(((....-....--..))).------.))).))).. ( -26.60) >DroYak_CAF1 6559 86 + 1 UUAAUCAUUUGUUUGUGAAUGCAAAUGAAAUGGCGAAAGUUUUG-GCCA-----------------UU-UGCCAUGUGCGAUA-AGUA--CAGCAA----AAAGGGUCGACC ....((((((((........))))))))...(((....)))(((-(((.-----------------((-(((..(((((....-.)))--))))))----)...)))))).. ( -26.50) >consensus UUAAUCAUUUGUUUGCGAAUGCAAAUGAAAUGGCGAAAGUUUUU_GCCAUUCUUGCAGCAUAUAAAUU_UGCUUUGUGCGAUA_AGUACGCAGCAAGGGUGAGGGGUCGACC ....((((((((........))))))))((((((...........))))))...................(((.(((((......))))).))).........((.....)) (-15.64 = -17.20 + 1.56)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:55:45 2006