
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 8,719,875 – 8,719,995 |
| Length | 120 |
| Max. P | 0.988363 |

| Location | 8,719,875 – 8,719,995 |
|---|---|
| Length | 120 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 90.83 |
| Mean single sequence MFE | -45.40 |
| Consensus MFE | -38.27 |
| Energy contribution | -38.63 |
| Covariance contribution | 0.36 |
| Combinations/Pair | 1.19 |
| Mean z-score | -3.00 |
| Structure conservation index | 0.84 |
| SVM decision value | 1.90 |
| SVM RNA-class probability | 0.981719 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
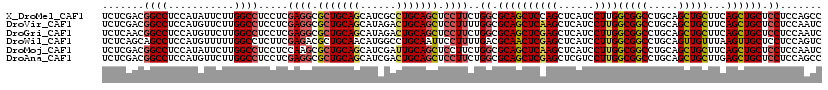
>X_DroMel_CAF1 8719875 120 + 22224390 UCUCGACGGCCUCCAUAUUCUUGGCCUCCUCGAGGCGCUGCAGCAUCGCCUGCAGCUCCUUCUGGCGCAGCUCCAGCUCAUCCUUGGCGGCCUGCAGCUGCUUCAGCUGCUCCUCCAGCC ((((((.((((...........))))...)))))).(((((((......))))))).......((((.(((....)))).....(((.((...(((((((...))))))).)).)))))) ( -47.70) >DroVir_CAF1 1353 120 + 1 UCUCGACGGCCUCCAUGUUCUUGGCCUCCUCGAGGCGCUGCAGCAUAGACUGCAGCUCCUUUUGGCGCAGCUCAAGCUCAUCCUUGGCGGCCUGCAGCUGCUUCAGCUGCUCCUCCAAUC .......((((.(((((..(((((((.((..((((.(((((((......))))))).))))..)).)..)).))))..))....))).)))).(((((((...))))))).......... ( -49.70) >DroGri_CAF1 4414 120 + 1 UCUCAACGGCCUCCAUGUUCUUGGCCUCCUCGAGGCGCUGCAGCAUAGACUGCAGCUCCUUCUGGCGCAGCUCGAGCUCAUCCUUGGCGGCCUGCAGCUGCUUCAGCUGCUCCUCCAAUC .......((((.(((((..(((((((.((..((((.(((((((......))))))).))))..)).)..)).))))..))....))).)))).(((((((...))))))).......... ( -49.40) >DroWil_CAF1 815 120 + 1 UCUCAGCAGCCUCCAUGUUUUUGGCCUCUUCGAGACGCUGCAACAUGGCCUGCAAUUCCUUUUGACGCAACUCGAGCUCAUCCUUGGCGGCCUGCAGUUGUUUAAGUUGCUCCUCCAGUC .....((((...(((((((...(((.((.....)).)))..))))))).)))).............((((((..(((.((....))((.....))....)))..)))))).......... ( -29.70) >DroMoj_CAF1 1342 120 + 1 UCUCGACGGCCUCCAUAUUCUUGGCCUCCUCCAAGCGCUGCAGCAUCGAUUGCAGCUCCUUCUGGCGCAGCUCAAGCUCAUCCUUGGCGGCCUGCAGCUGCUUCAGCUGCUCCUCCAAUC .......((((.(((....(((((......))))).(((((.(((.....))).(((......)))))))).............))).)))).(((((((...))))))).......... ( -42.10) >DroAna_CAF1 651 120 + 1 UCUCGACGGCCUCCAUGUUCUUGGCCUCCUCGAGGCGCUGCAGCAUCGACUGCAGCUCCUUCUGGCGCAGCUCGAGCUCGUCCUUGGCGGCCUGCAGCUGCUUGAGCUGCUCCUCCAGCC .(((((.((((...........))))...)))))(((((((((......))))))).......((.(((((((((((.(((.....)))((.....)).))))))))))).))....)). ( -53.80) >consensus UCUCGACGGCCUCCAUGUUCUUGGCCUCCUCGAGGCGCUGCAGCAUCGACUGCAGCUCCUUCUGGCGCAGCUCGAGCUCAUCCUUGGCGGCCUGCAGCUGCUUCAGCUGCUCCUCCAAUC .......((((...........)))).....((((.(((((((......))))))).))))..((.((((((((((......))))(((((.....)))))...)))))).))....... (-38.27 = -38.63 + 0.36)


| Location | 8,719,875 – 8,719,995 |
|---|---|
| Length | 120 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 90.83 |
| Mean single sequence MFE | -49.15 |
| Consensus MFE | -43.75 |
| Energy contribution | -43.20 |
| Covariance contribution | -0.55 |
| Combinations/Pair | 1.28 |
| Mean z-score | -2.36 |
| Structure conservation index | 0.89 |
| SVM decision value | 2.12 |
| SVM RNA-class probability | 0.988363 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
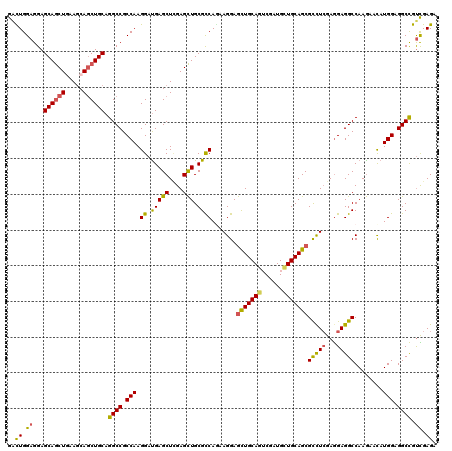
>X_DroMel_CAF1 8719875 120 - 22224390 GGCUGGAGGAGCAGCUGAAGCAGCUGCAGGCCGCCAAGGAUGAGCUGGAGCUGCGCCAGAAGGAGCUGCAGGCGAUGCUGCAGCGCCUCGAGGAGGCCAAGAAUAUGGAGGCCGUCGAGA ((((......(((((((...))))))).((((.((..((.(((((....))).)))).((.((.(((((((......))))))).))))..)).))))...........))))....... ( -53.10) >DroVir_CAF1 1353 120 - 1 GAUUGGAGGAGCAGCUGAAGCAGCUGCAGGCCGCCAAGGAUGAGCUUGAGCUGCGCCAAAAGGAGCUGCAGUCUAUGCUGCAGCGCCUCGAGGAGGCCAAGAACAUGGAGGCCGUCGAGA (((.......(((((((...))))))).((((.(((....((..((((.(((...((...(((.((((((((....)))))))).)))...)).)))))))..))))).))))))).... ( -54.10) >DroGri_CAF1 4414 120 - 1 GAUUGGAGGAGCAGCUGAAGCAGCUGCAGGCCGCCAAGGAUGAGCUCGAGCUGCGCCAGAAGGAGCUGCAGUCUAUGCUGCAGCGCCUCGAGGAGGCCAAGAACAUGGAGGCCGUUGAGA ..........(((((((...))))))).((((.(((.((.(((((....))).)))).......((((((((....))))))))(((((...)))))........))).))))....... ( -50.50) >DroWil_CAF1 815 120 - 1 GACUGGAGGAGCAACUUAAACAACUGCAGGCCGCCAAGGAUGAGCUCGAGUUGCGUCAAAAGGAAUUGCAGGCCAUGUUGCAGCGUCUCGAAGAGGCCAAAAACAUGGAGGCUGCUGAGA ......((.(((...........((((((....((...(((((((....))).))))....))..)))))).(((((((...(.(((((...))))))...)))))))..))).)).... ( -35.30) >DroMoj_CAF1 1342 120 - 1 GAUUGGAGGAGCAGCUGAAGCAGCUGCAGGCCGCCAAGGAUGAGCUUGAGCUGCGCCAGAAGGAGCUGCAAUCGAUGCUGCAGCGCUUGGAGGAGGCCAAGAAUAUGGAGGCCGUCGAGA (((.......(((((((...))))))).((((.(((...((...((((.(((...((...(((.((((((........)))))).)))...)).))))))).)).))).))))))).... ( -47.90) >DroAna_CAF1 651 120 - 1 GGCUGGAGGAGCAGCUCAAGCAGCUGCAGGCCGCCAAGGACGAGCUCGAGCUGCGCCAGAAGGAGCUGCAGUCGAUGCUGCAGCGCCUCGAGGAGGCCAAGAACAUGGAGGCCGUCGAGA ((((......((((((.....)))))).((((.((..((.(((((....))).)))).((.((.((((((((....)))))))).))))..)).))))...........))))....... ( -54.00) >consensus GACUGGAGGAGCAGCUGAAGCAGCUGCAGGCCGCCAAGGAUGAGCUCGAGCUGCGCCAGAAGGAGCUGCAGUCGAUGCUGCAGCGCCUCGAGGAGGCCAAGAACAUGGAGGCCGUCGAGA ..((.((...((((((.....)))))).((((.(((.((.(((((....))).)))).......(((((((......)))))))(((((...)))))........))).)))).)).)). (-43.75 = -43.20 + -0.55)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:55:31 2006