
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 8,667,774 – 8,667,869 |
| Length | 95 |
| Max. P | 0.965121 |

| Location | 8,667,774 – 8,667,869 |
|---|---|
| Length | 95 |
| Sequences | 5 |
| Columns | 95 |
| Reading direction | forward |
| Mean pairwise identity | 91.74 |
| Mean single sequence MFE | -29.94 |
| Consensus MFE | -24.86 |
| Energy contribution | -25.02 |
| Covariance contribution | 0.16 |
| Combinations/Pair | 1.04 |
| Mean z-score | -1.43 |
| Structure conservation index | 0.83 |
| SVM decision value | 0.36 |
| SVM RNA-class probability | 0.704118 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
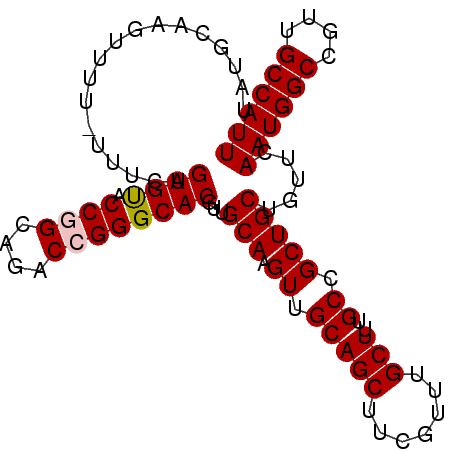
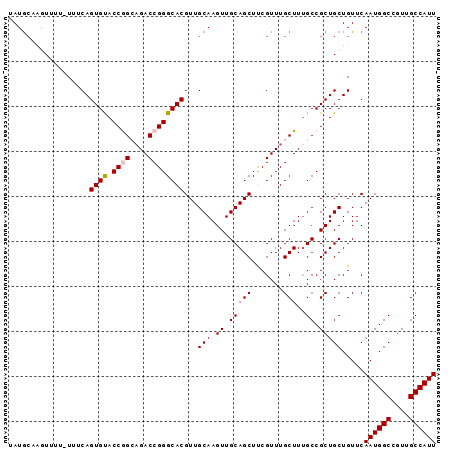
>X_DroMel_CAF1 8667774 95 + 22224390 UAUGCUAGUUUUUUUUCAGUGUACCGGCAGACCGGGCACGUUGCAAGUUGCAGCUUCGUUUGCUUUGCCGCUGCUGUUCAAUGGCCGUUGCCAUU ................(((((...(((((((.(((((..(((((.....)))))...))))).))))))).)))))...((((((....)))))) ( -28.90) >DroSec_CAF1 104446 94 + 1 UAUGCCAGUUUC-UUUCAGUGUACCAGCAGACCGGGCACGUUGCAAGUUGCAGCUUCGUUUGCUUUGCCGCUGCUGUUCAAUGGCCGUUGCCAUU ...((((.....-...(((((....(((((((..(((....(((.....))))))..)))))))....)))))........)))).......... ( -26.49) >DroSim_CAF1 68833 94 + 1 UAUGCCAGUUUC-UUUCAGUGUACCAGCAGACCGGGCACGUUGCAAGUUGCAGCUUCGUUUGCUUUGCCGCUGCUGUUCAAUGGCCGUUGCCAUU ...((((.....-...(((((....(((((((..(((....(((.....))))))..)))))))....)))))........)))).......... ( -26.49) >DroEre_CAF1 116202 94 + 1 UAUAGAUAUUUU-UUCCAGUGCACCGGGCGACCGGGCACGUUGCAAGUUGCAGCUCCGUUUGCUUUGCCGCUGCUGUUCAAUGGCCAUUGCCAUU .((((.......-.....((((.((((....))))))))((.(((((..((((......))))))))).))..))))..((((((....)))))) ( -30.70) >DroYak_CAF1 107891 94 + 1 UAUGCAUAUUUU-UUUCAGUGCACCGGGAGACCGGGCACGUUGCAAGUUGCAGCUCCGUUUGCUUGGCCGCUGCUGUUCAAUGGCCGUUGCCAUU ...(((......-.....((((.((((....))))))))(((((.....)))))......))).((((.((.(((((...))))).)).)))).. ( -37.10) >consensus UAUGCAAGUUUU_UUUCAGUGUACCGGCAGACCGGGCACGUUGCAAGUUGCAGCUUCGUUUGCUUUGCCGCUGCUGUUCAAUGGCCGUUGCCAUU ..................((((.((((....))))))))...(((.((.(((((.......)))..)).))))).....((((((....)))))) (-24.86 = -25.02 + 0.16)

| Location | 8,667,774 – 8,667,869 |
|---|---|
| Length | 95 |
| Sequences | 5 |
| Columns | 95 |
| Reading direction | reverse |
| Mean pairwise identity | 91.74 |
| Mean single sequence MFE | -27.46 |
| Consensus MFE | -24.06 |
| Energy contribution | -23.58 |
| Covariance contribution | -0.48 |
| Combinations/Pair | 1.10 |
| Mean z-score | -1.71 |
| Structure conservation index | 0.88 |
| SVM decision value | 1.58 |
| SVM RNA-class probability | 0.965121 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
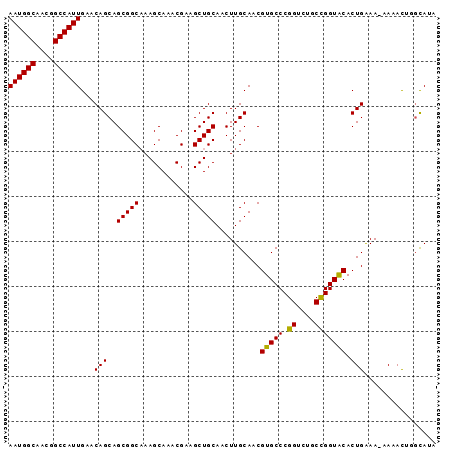
>X_DroMel_CAF1 8667774 95 - 22224390 AAUGGCAACGGCCAUUGAACAGCAGCGGCAAAGCAAACGAAGCUGCAACUUGCAACGUGCCCGGUCUGCCGGUACACUGAAAAAAAACUAGCAUA ((((((....)))))).....((.(((((............)))))..........(((((.((....)))))))...............))... ( -25.90) >DroSec_CAF1 104446 94 - 1 AAUGGCAACGGCCAUUGAACAGCAGCGGCAAAGCAAACGAAGCUGCAACUUGCAACGUGCCCGGUCUGCUGGUACACUGAAA-GAAACUGGCAUA ((((((....)))))).....((.(((((............)))))..(((.((..(((((.((....)))))))..)).))-)......))... ( -24.90) >DroSim_CAF1 68833 94 - 1 AAUGGCAACGGCCAUUGAACAGCAGCGGCAAAGCAAACGAAGCUGCAACUUGCAACGUGCCCGGUCUGCUGGUACACUGAAA-GAAACUGGCAUA ((((((....)))))).....((.(((((............)))))..(((.((..(((((.((....)))))))..)).))-)......))... ( -24.90) >DroEre_CAF1 116202 94 - 1 AAUGGCAAUGGCCAUUGAACAGCAGCGGCAAAGCAAACGGAGCUGCAACUUGCAACGUGCCCGGUCGCCCGGUGCACUGGAA-AAAAUAUCUAUA ((((((....)))))).....((((((((............)))))....)))...((((((((....)))).))))((((.-......)))).. ( -28.40) >DroYak_CAF1 107891 94 - 1 AAUGGCAACGGCCAUUGAACAGCAGCGGCCAAGCAAACGGAGCUGCAACUUGCAACGUGCCCGGUCUCCCGGUGCACUGAAA-AAAAUAUGCAUA ((((((....)))))).....(((((..((........)).)))))....((((..((((((((....)))).))))(....-).....)))).. ( -33.20) >consensus AAUGGCAACGGCCAUUGAACAGCAGCGGCAAAGCAAACGAAGCUGCAACUUGCAACGUGCCCGGUCUGCCGGUACACUGAAA_AAAACUGGCAUA ((((((....))))))...(((..(((((............)))))..........(((((.((....))))))).)))................ (-24.06 = -23.58 + -0.48)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:55:11 2006