
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 8,440,956 – 8,441,103 |
| Length | 147 |
| Max. P | 0.776953 |

| Location | 8,440,956 – 8,441,075 |
|---|---|
| Length | 119 |
| Sequences | 6 |
| Columns | 119 |
| Reading direction | forward |
| Mean pairwise identity | 81.96 |
| Mean single sequence MFE | -41.60 |
| Consensus MFE | -26.55 |
| Energy contribution | -26.72 |
| Covariance contribution | 0.17 |
| Combinations/Pair | 1.24 |
| Mean z-score | -2.06 |
| Structure conservation index | 0.64 |
| SVM decision value | 0.34 |
| SVM RNA-class probability | 0.697716 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
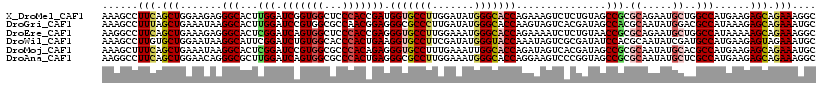
>X_DroMel_CAF1 8440956 119 + 22224390 GCCUUUCUGCUCUUCAUGGCCAGCAUUCUGCGCGGCUACAGAGACUUUCUGGUGCCCAUAUCCAAGGCACCAUCGGUGGGAGCCACCGAUCCAAGUGCCCUCUUCCAGCUGAAGGCUUU ((((((..(((.....(((((.((.....))..))))).((((((((...((((((.........))))))((((((((...))))))))..))))...))))...))).))))))... ( -44.90) >DroGri_CAF1 59418 119 + 1 GCAUUUCUGCUCUUUAUGGCGUCCAUAUUGCGUGGCUAUCGUGACUACUUGGUGCCCAUAUCAAGGGCGCCCUCCGUUGGCGCCACGGAUCCAAGUGCCUUAUUUCAGCUAAAGGCUUU (((....)))..((..(((((.(((......(((((......).))))..(((((((.......)))))))......))))))))..)).......(((((..........)))))... ( -36.60) >DroEre_CAF1 3860 119 + 1 GCCUUUCUGCUUUUUAUGGCCAGCAUUCUGCGCGGUUACAGAGAUUUUCUGGUGCCCAUUUCCAAGGCACCCUCGGUGGGAGCCACUGAUCCGAGUGCCCUCUUUCAGCUGAAGGCCUU ((((((..(((.....(((((.((.....))..))))).((((...(((.((((((.........)))))).(((((((...)))))))...)))....))))...))).))))))... ( -38.50) >DroWil_CAF1 2098 119 + 1 GCAUUUCUACUCUUCAUGGCAUCGAUAUUGCGUGGAUAUCGCGACUAUUUGGUACCCAUAUCGAAGGCACCUUCAGUGGGUGCCACAGAUCCGAAUGCCUUAUUCCAGCACAAGGCUUU ((...............(((((((...(((((.......))))).....((((((((((...((((....)))).))))))))))......)).)))))...............))... ( -35.51) >DroMoj_CAF1 91797 119 + 1 GCAUUUCUGCUCUUCAUGGCGUGCAUAUUGCGCGGCUAUCGUGACUAUCUGGUGCCAAUUUCAAAGGCACCCUCUGUGGGCGCCACGGAUCCGAGUGCCUUAUUUCAGCUGAAAGCUUU ((.((((.(((.........((((.....))))(((..(((.((......((((((.........)))))).(((((((...))))))))))))..))).......))).))))))... ( -40.00) >DroAna_CAF1 2264 119 + 1 GCCUUUCUGCUCUUCAUGGCGAGCAUAUUGCGCGGCUACCGGGACUUCCUGGUGCCCAUUUCCAAGGCGCCCUCAGUGGGCGCCACUGAUCCAAGCGCCCUGUUCCAGCUGAAGGCCUU ((((((..(((...((.(((..((.....))(((((.((((((....))))))))).........(((((((.....)))))))..........))))).))....))).))))))... ( -54.10) >consensus GCAUUUCUGCUCUUCAUGGCGAGCAUAUUGCGCGGCUACCGAGACUAUCUGGUGCCCAUAUCCAAGGCACCCUCAGUGGGCGCCACGGAUCCAAGUGCCCUAUUCCAGCUGAAGGCUUU ((.((((.(((......(((..............................((((((.........)))))).(((((((...))))))).......))).......))).))))))... (-26.55 = -26.72 + 0.17)


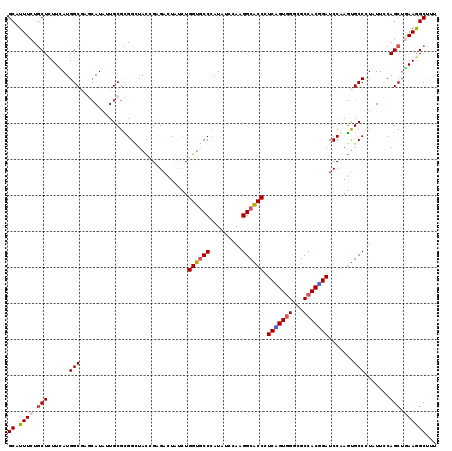
| Location | 8,440,956 – 8,441,075 |
|---|---|
| Length | 119 |
| Sequences | 6 |
| Columns | 119 |
| Reading direction | reverse |
| Mean pairwise identity | 81.96 |
| Mean single sequence MFE | -41.38 |
| Consensus MFE | -25.59 |
| Energy contribution | -26.65 |
| Covariance contribution | 1.06 |
| Combinations/Pair | 1.29 |
| Mean z-score | -1.75 |
| Structure conservation index | 0.62 |
| SVM decision value | -0.01 |
| SVM RNA-class probability | 0.526338 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 8440956 119 - 22224390 AAAGCCUUCAGCUGGAAGAGGGCACUUGGAUCGGUGGCUCCCACCGAUGGUGCCUUGGAUAUGGGCACCAGAAAGUCUCUGUAGCCGCGCAGAAUGCUGGCCAUGAAGAGCAGAAAGGC ...((((((.(((.....((((((((...((((((((...))))))))))))))))...(((((((..((((.....))))..)))((.(((....)))))))))...))).).))))) ( -47.10) >DroGri_CAF1 59418 119 - 1 AAAGCCUUUAGCUGAAAUAAGGCACUUGGAUCCGUGGCGCCAACGGAGGGCGCCCUUGAUAUGGGCACCAAGUAGUCACGAUAGCCACGCAAUAUGGACGCCAUAAAGAGCAGAAAUGC ..(((.....))).......(((((((((.((((((....).)))))....((((.......)))).)))))).(((...((.((...)).))...)))))).......(((....))) ( -37.00) >DroEre_CAF1 3860 119 - 1 AAGGCCUUCAGCUGAAAGAGGGCACUCGGAUCAGUGGCUCCCACCGAGGGUGCCUUGGAAAUGGGCACCAGAAAAUCUCUGUAACCGCGCAGAAUGCUGGCCAUAAAAAGCAGAAAGGC ...((((((.(((.....(((((((((...((.((((...)))).)))))))))))....((((...((((...((.(((((......))))))).))))))))....))).).))))) ( -39.00) >DroWil_CAF1 2098 119 - 1 AAAGCCUUGUGCUGGAAUAAGGCAUUCGGAUCUGUGGCACCCACUGAAGGUGCCUUCGAUAUGGGUACCAAAUAGUCGCGAUAUCCACGCAAUAUCGAUGCCAUGAAGAGUAGAAAUGC ...(((((((......))))))).(((...(((((((((...((((..(((((((.......)))))))...))))..((((((.......)))))).))))))..)))...))).... ( -37.50) >DroMoj_CAF1 91797 119 - 1 AAAGCUUUCAGCUGAAAUAAGGCACUCGGAUCCGUGGCGCCCACAGAGGGUGCCUUUGAAAUUGGCACCAGAUAGUCACGAUAGCCGCGCAAUAUGCACGCCAUGAAGAGCAGAAAUGC ..(((.....))).......(((...(((...((((((((((.....))))(((.........)))........))))))....))).((.....))..))).......(((....))) ( -35.00) >DroAna_CAF1 2264 119 - 1 AAGGCCUUCAGCUGGAACAGGGCGCUUGGAUCAGUGGCGCCCACUGAGGGCGCCUUGGAAAUGGGCACCAGGAAGUCCCGGUAGCCGCGCAAUAUGCUCGCCAUGAAGAGCAGAAAGGC ...((((((.((.....(((((((((....(((((((...))))))).)))))))))......((((((.((....)).))).)))))......(((((........)))))).))))) ( -52.70) >consensus AAAGCCUUCAGCUGAAAUAAGGCACUCGGAUCAGUGGCGCCCACCGAGGGUGCCUUGGAAAUGGGCACCAGAAAGUCACGAUAGCCGCGCAAUAUGCACGCCAUGAAGAGCAGAAAGGC ......(((.(((.......(((...(((.(((((((...))))))).(((((((.......)))))))...............))).((.....))..)))......))).))).... (-25.59 = -26.65 + 1.06)


| Location | 8,440,995 – 8,441,103 |
|---|---|
| Length | 108 |
| Sequences | 6 |
| Columns | 110 |
| Reading direction | forward |
| Mean pairwise identity | 79.50 |
| Mean single sequence MFE | -38.18 |
| Consensus MFE | -28.01 |
| Energy contribution | -27.98 |
| Covariance contribution | -0.02 |
| Combinations/Pair | 1.26 |
| Mean z-score | -1.43 |
| Structure conservation index | 0.73 |
| SVM decision value | 0.55 |
| SVM RNA-class probability | 0.776953 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 8440995 108 + 22224390 AGAGACUUUCUGGUGCCCAUAUCCAAGGCACCAUCGGUGGGAGCCACCGAUCCAAGUGCCCUCUUCCAGCUGAAGGCUUUUUUGCGAUCGCGUGAUAAGGUAAGAA--AA ......(((((((((((.........))))))((((((((...)))))((((((((.(((.((........)).)))...)))).))))....)))......))))--). ( -34.90) >DroVir_CAF1 2104 110 + 1 CGUGACUACUUGGUGCCUAUUUCAAAGGCGCCUUCCGUGGGCGCCACGGAUCCAAGUGCCUUAUUUCAGCUGAAAGCUUUUCUGCGCUCGCGCGAUAAGGUAAGUGCAAA .((.((((((((((((((.......))))))).(((((((...)))))))...))))((((((((...((.((..((......))..))))..)))))))).)))))... ( -40.00) >DroPse_CAF1 4021 108 + 1 CGCGACUAUUUGGUGCCCAUAUCGAAGGCACCAUCGGUGGGGGCCACCGAUCCGAGUGCCUUAUUCCAGCUGAAGGCCUUUCUGCGUUCGCGCGAUAAGGUAACGA--GU ((.((.....(((((((.........)))))))(((((((...)))))))))))..(((((((((...((.(((.((......)).)))))..)))))))))....--.. ( -40.00) >DroGri_CAF1 59457 110 + 1 CGUGACUACUUGGUGCCCAUAUCAAGGGCGCCCUCCGUUGGCGCCACGGAUCCAAGUGCCUUAUUUCAGCUAAAGGCUUUUCUGCGCUCGCGCGACAAGGUAAGAGUGAA ......((((((((((((.......))))))).(((((.(....))))))......((((((.....(((.....)))....((((....))))..)))))).))))).. ( -34.40) >DroWil_CAF1 2137 110 + 1 CGCGACUAUUUGGUACCCAUAUCGAAGGCACCUUCAGUGGGUGCCACAGAUCCGAAUGCCUUAUUCCAGCACAAGGCUUUUCUGAGAUCACGUGAUAAGGUGAGAGUUGA ..(((((...((((((((((...((((....)))).))))))))))((((...(((.(((((..........))))))))))))...((((.(....).)))).))))). ( -36.70) >DroAna_CAF1 2303 104 + 1 CGGGACUUCCUGGUGCCCAUUUCCAAGGCGCCCUCAGUGGGCGCCACUGAUCCAAGCGCCCUGUUCCAGCUGAAGGCCUUUUUGCGUUCCCGCGAUAAGGUGAG------ .(((((.....(((((.....((...(((((((.....)))))))...)).....)))))..)))))........(((((.(((((....))))).)))))...------ ( -43.10) >consensus CGCGACUACUUGGUGCCCAUAUCAAAGGCACCAUCAGUGGGCGCCACCGAUCCAAGUGCCUUAUUCCAGCUGAAGGCUUUUCUGCGAUCGCGCGAUAAGGUAAGAG__AA ...........((((((.........)))))).(((((((...)))))))......(((((((((...((.....))......(((....))))))))))))........ (-28.01 = -27.98 + -0.02)

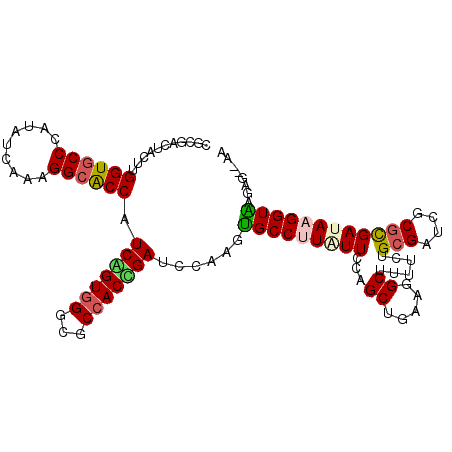

| Location | 8,440,995 – 8,441,103 |
|---|---|
| Length | 108 |
| Sequences | 6 |
| Columns | 110 |
| Reading direction | reverse |
| Mean pairwise identity | 79.50 |
| Mean single sequence MFE | -37.18 |
| Consensus MFE | -25.99 |
| Energy contribution | -25.60 |
| Covariance contribution | -0.39 |
| Combinations/Pair | 1.30 |
| Mean z-score | -1.34 |
| Structure conservation index | 0.70 |
| SVM decision value | 0.31 |
| SVM RNA-class probability | 0.682342 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 8440995 108 - 22224390 UU--UUCUUACCUUAUCACGCGAUCGCAAAAAAGCCUUCAGCUGGAAGAGGGCACUUGGAUCGGUGGCUCCCACCGAUGGUGCCUUGGAUAUGGGCACCAGAAAGUCUCU ((--((((..((.......((....))......((((((........))))))....))((((((((...))))))))(((((((.......)))))))))))))..... ( -39.50) >DroVir_CAF1 2104 110 - 1 UUUGCACUUACCUUAUCGCGCGAGCGCAGAAAAGCUUUCAGCUGAAAUAAGGCACUUGGAUCCGUGGCGCCCACGGAAGGCGCCUUUGAAAUAGGCACCAAGUAGUCACG ..((.(((..(((((((((....)))......(((.....)))...)))))).((((((.(((((((...)))))))....((((.......)))).))))))))))).. ( -37.20) >DroPse_CAF1 4021 108 - 1 AC--UCGUUACCUUAUCGCGCGAACGCAGAAAGGCCUUCAGCUGGAAUAAGGCACUCGGAUCGGUGGCCCCCACCGAUGGUGCCUUCGAUAUGGGCACCAAAUAGUCGCG ..--..............(((((.((.((....(((((..........))))).)))).((((((((...))))))))(((((((.......)))))))......))))) ( -36.90) >DroGri_CAF1 59457 110 - 1 UUCACUCUUACCUUGUCGCGCGAGCGCAGAAAAGCCUUUAGCUGAAAUAAGGCACUUGGAUCCGUGGCGCCAACGGAGGGCGCCCUUGAUAUGGGCACCAAGUAGUCACG ............(((.(((....))))))....(((((..........)))))((((((.((((((....).)))))....((((.......)))).))))))....... ( -36.20) >DroWil_CAF1 2137 110 - 1 UCAACUCUCACCUUAUCACGUGAUCUCAGAAAAGCCUUGUGCUGGAAUAAGGCAUUCGGAUCUGUGGCACCCACUGAAGGUGCCUUCGAUAUGGGUACCAAAUAGUCGCG .........(((((((((((.(((((..(((..(((((((......))))))).)))))))))))((((((.......))))))...)))).)))).............. ( -33.00) >DroAna_CAF1 2303 104 - 1 ------CUCACCUUAUCGCGGGAACGCAAAAAGGCCUUCAGCUGGAACAGGGCGCUUGGAUCAGUGGCGCCCACUGAGGGCGCCUUGGAAAUGGGCACCAGGAAGUCCCG ------...........(((....))).....((.((((..((((..(((((((((....(((((((...))))))).)))))))))(.......).)))))))).)).. ( -40.30) >consensus UU__CUCUUACCUUAUCGCGCGAACGCAGAAAAGCCUUCAGCUGGAAUAAGGCACUUGGAUCCGUGGCGCCCACCGAAGGCGCCUUCGAUAUGGGCACCAAAUAGUCACG ..........((.....(((....)))......(((((..........)))))....)).(((((((...))))))).((.((((.......)))).))........... (-25.99 = -25.60 + -0.39)


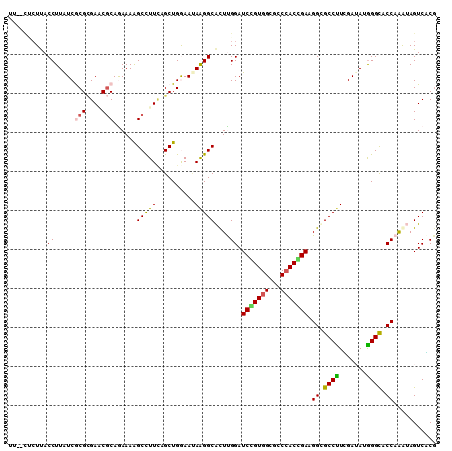
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:53:09 2006