
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 8,404,190 – 8,404,334 |
| Length | 144 |
| Max. P | 0.942749 |

| Location | 8,404,190 – 8,404,295 |
|---|---|
| Length | 105 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 73.31 |
| Mean single sequence MFE | -22.38 |
| Consensus MFE | -9.96 |
| Energy contribution | -9.65 |
| Covariance contribution | -0.31 |
| Combinations/Pair | 1.09 |
| Mean z-score | -2.20 |
| Structure conservation index | 0.45 |
| SVM decision value | 0.20 |
| SVM RNA-class probability | 0.631683 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
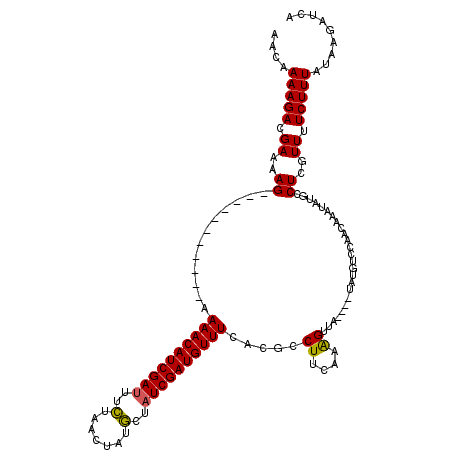
>X_DroMel_CAF1 8404190 105 - 22224390 AACAAAAGACGAAAAG-----------AAAACAUCGAUUUUUCACUAUACUAUCGAUGUUUCACGCCUUCAAAGUGA----AAUGUCCAAAAAAUAUGCCUCGUUUUCUUUAUAUGAUUA ............((((-----------(((((...((....))...........((((((((((.........))))----))))))...............)))))))))......... ( -19.30) >DroSec_CAF1 3400 88 - 1 AACAAAAGACGAAAAG-----------AAAACAUCGAUUUCUCACUAUGCUAUCGAUGUUUCAGGUCUUCAAGGU---------------------UGCCUAGUUGUCUUUAUAUGAUUA ....((((((((...(-----------(((.(((((((..(.......)..)))))))))))((((..(....).---------------------.))))..))))))))......... ( -21.70) >DroEre_CAF1 6825 116 - 1 AACAAAAGACGAAAAGUGAGCAAACCGAAAACAUCGAUUUCUAGCUAUGCAAUCGAUGUUUUGCGCCUUCAAAGUUA----UAUGUCCAACAUAUAUGCCUUGUUUUCUUUAUAAGAUCA ............((((.((((((..((((((((((((((.(.......).)))))))))))).))..........((----((((.....))))))....)))))).))))......... ( -24.00) >DroYak_CAF1 6435 120 - 1 AACAAAAGACGAAAAGUGAGGAAAACAAAAACAUCGAAUCCUAGCUGUGCUAUCGAUGUUUCGCGUCUUUAAAGUUAUGUAUAUGUCCAACAAAUAUACCUCGUUUUCUUUAAAAGAUCA ..........((....((((((((((..(((((((((....((((...))))))))))))).................((((((.........))))))...)))))))))).....)). ( -24.50) >consensus AACAAAAGACGAAAAG___________AAAACAUCGAUUUCUAACUAUGCUAUCGAUGUUUCACGCCUUCAAAGUUA____UAUGUCCAACAAAUAUGCCUCGUUUUCUUUAUAAGAUCA ....(((((.((..((............((((((((((..(.......)..)))))))))).....((....)).........................))..)).)))))......... ( -9.96 = -9.65 + -0.31)


| Location | 8,404,229 – 8,404,334 |
|---|---|
| Length | 105 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 75.47 |
| Mean single sequence MFE | -25.33 |
| Consensus MFE | -14.27 |
| Energy contribution | -14.02 |
| Covariance contribution | -0.25 |
| Combinations/Pair | 1.16 |
| Mean z-score | -2.22 |
| Structure conservation index | 0.56 |
| SVM decision value | 1.33 |
| SVM RNA-class probability | 0.942749 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
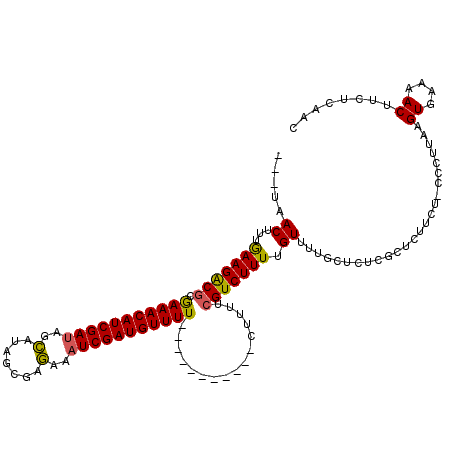
>X_DroMel_CAF1 8404229 105 + 22224390 ---UCACUUUGAAGGCGUGAAACAUCGAUAGUAUAGUGAAAAAUCGAUGUUUU-----------CUUUUCGUCUUUUGUUUUGCUCUCGCUCUUCU-CCAUGAAGUGAUAACUUCUCAAC ---((((((((((((((.(((((((((((.............)))))))))))-----------.....)))))))......((....))......-....)))))))............ ( -24.32) >DroSec_CAF1 3424 100 + 1 -----ACCUUGAAGACCUGAAACAUCGAUAGCAUAGUGAGAAAUCGAUGUUUU-----------CUUUUCGUCUUUUGUUGUGCUCUCGGUCUUCU-UCCUAAAGUGAAAAC---UCAAC -----.....(((((((.((....))((.(((((((..(((...(((......-----------....)))..)))..))))))).))))))))).-......(((....))---).... ( -26.50) >DroEre_CAF1 6864 114 + 1 ---UAACUUUGAAGGCGCAAAACAUCGAUUGCAUAGCUAGAAAUCGAUGUUUUCGGUUUGCUCACUUUUCGUCUUUUGUUUUGCUCUCGCUCUUC---CCCUGCGUGAAAACUUCUUAUU ---.......((((..((((((((.(((......(((...(((((((.....))))))))))......))).....))))))))((.(((.....---....))).))...))))..... ( -27.20) >DroYak_CAF1 6475 120 + 1 ACAUAACUUUAAAGACGCGAAACAUCGAUAGCACAGCUAGGAUUCGAUGUUUUUGUUUUCCUCACUUUUCGUCUUUUGUUUUGCUCUCGCUCUUCUGCCCUUUCGUGAAAACUUCUCACU .((.(((...(((((((.((((((((((((((...))))....))))))))))((.......)).....))))))).))).)).....((......))......((((.......)))). ( -23.30) >consensus ___UAACUUUGAAGACGCGAAACAUCGAUAGCAUAGCGAGAAAUCGAUGUUUU___________CUUUUCGUCUUUUGUUUUGCUCUCGCUCUUCU_CCCUUAAGUGAAAACUUCUCAAC .....((...(((((((.(((((((((((..(.......)..)))))))))))................))))))).)).......((((..............))))............ (-14.27 = -14.02 + -0.25)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:52:48 2006