
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 8,352,223 – 8,352,321 |
| Length | 98 |
| Max. P | 0.874107 |

| Location | 8,352,223 – 8,352,321 |
|---|---|
| Length | 98 |
| Sequences | 4 |
| Columns | 106 |
| Reading direction | forward |
| Mean pairwise identity | 76.28 |
| Mean single sequence MFE | -32.75 |
| Consensus MFE | -18.42 |
| Energy contribution | -18.18 |
| Covariance contribution | -0.25 |
| Combinations/Pair | 1.35 |
| Mean z-score | -2.12 |
| Structure conservation index | 0.56 |
| SVM decision value | 0.88 |
| SVM RNA-class probability | 0.874107 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 8352223 98 + 22224390 -UCGAU---CGAUUGUGUU----UUGGAUUGCCGGCGAAUUUUUGCGCCGGCGGCUGAUCCACUAAUCAAGUUGCAUUUAUAUGCCCCACCGAUCCGUCGGCUAUA -..(((---((((((((..----(..(.(((((((((........))))))))))..)..))).)))).....((((....))))......))))........... ( -34.20) >DroPse_CAF1 14261 97 + 1 ---GAUUGCCGAUUGUGUCGUUUGUCGUUUGUCGUCG----UCUACGGCGGCAUCUGAUCCACUAAUCAAGGUGCAUUUAUAUGCCCAGCGGAUCCGAUGGCGA-- ---..(((((.((((..((((..((((..((((((((----....))))))))..))))..)).......((.((((....))))))....))..)))))))))-- ( -34.00) >DroSec_CAF1 17673 99 + 1 AUCGAU---CGAUUGUGUU----UUGGAUUGCCGGCGAAUUUUUGCGCCGACGGCUGAUCCACUAAUCAAGGUGCAUUUAUAUGCCCCACCGAUCCGCCGGCCAUA ...(((---((((((((..----(..(.(((.(((((........))))).))))..)..))).))))..((.((((....)))).))...))))........... ( -31.00) >DroYak_CAF1 15794 86 + 1 ---GAU---CGAUUGUGUU----UUGGAUUGCCGGCGG----------UGGCGGCUGAUCCACUAAUCAAGGUGCAUUUAUAUGCCCCACCGAUCCGGCGACUUUA ---(((---(.(.......----.).))))((((((((----------(((.(((.(((.((((......)))).))).....)))))))))..)))))....... ( -31.80) >consensus ___GAU___CGAUUGUGUU____UUGGAUUGCCGGCGA___UUUGCGCCGGCGGCUGAUCCACUAAUCAAGGUGCAUUUAUAUGCCCCACCGAUCCGACGGCUAUA ........................(((...(((((((........)))))))(((.((((..((.....))..((((....))))......)))).)))..))).. (-18.42 = -18.18 + -0.25)


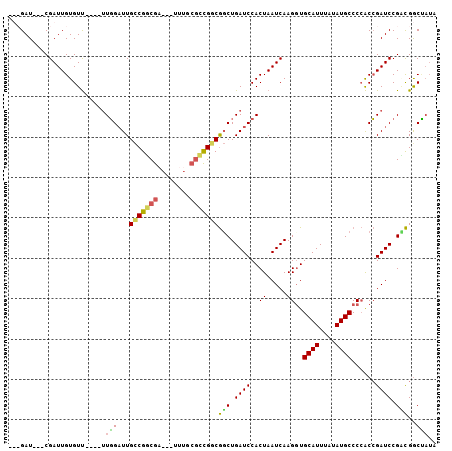
| Location | 8,352,223 – 8,352,321 |
|---|---|
| Length | 98 |
| Sequences | 4 |
| Columns | 106 |
| Reading direction | reverse |
| Mean pairwise identity | 76.28 |
| Mean single sequence MFE | -29.44 |
| Consensus MFE | -18.45 |
| Energy contribution | -20.95 |
| Covariance contribution | 2.50 |
| Combinations/Pair | 1.19 |
| Mean z-score | -1.86 |
| Structure conservation index | 0.63 |
| SVM decision value | 0.69 |
| SVM RNA-class probability | 0.824240 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 8352223 98 - 22224390 UAUAGCCGACGGAUCGGUGGGGCAUAUAAAUGCAACUUGAUUAGUGGAUCAGCCGCCGGCGCAAAAAUUCGCCGGCAAUCCAA----AACACAAUCG---AUCGA- ...........(((((((.((((((....))))...((((((....))))))))(((((((........))))))).......----......))))---)))..- ( -31.60) >DroPse_CAF1 14261 97 - 1 --UCGCCAUCGGAUCCGCUGGGCAUAUAAAUGCACCUUGAUUAGUGGAUCAGAUGCCGCCGUAGA----CGACGACAAACGACAAACGACACAAUCGGCAAUC--- --..(.((((.((((((((((((((....)))).))......)))))))).)))).)((((....----...((............)).......))))....--- ( -27.14) >DroSec_CAF1 17673 99 - 1 UAUGGCCGGCGGAUCGGUGGGGCAUAUAAAUGCACCUUGAUUAGUGGAUCAGCCGUCGGCGCAAAAAUUCGCCGGCAAUCCAA----AACACAAUCG---AUCGAU ....(((((((((((((((((((((....)))).)))(((((....)))))))))...........)))))))))).......----..........---...... ( -31.00) >DroYak_CAF1 15794 86 - 1 UAAAGUCGCCGGAUCGGUGGGGCAUAUAAAUGCACCUUGAUUAGUGGAUCAGCCGCCA----------CCGCCGGCAAUCCAA----AACACAAUCG---AUC--- .......(((((..(((((((((((....)))).)).......((((.....))))))----------)))))))).......----..........---...--- ( -28.00) >consensus UAUAGCCGACGGAUCGGUGGGGCAUAUAAAUGCACCUUGAUUAGUGGAUCAGCCGCCGGCGCAAA___UCGCCGGCAAUCCAA____AACACAAUCG___AUC___ ....((((((((((((((((.((((....))))...((((((....))))))))))))........)))))))))).............................. (-18.45 = -20.95 + 2.50)


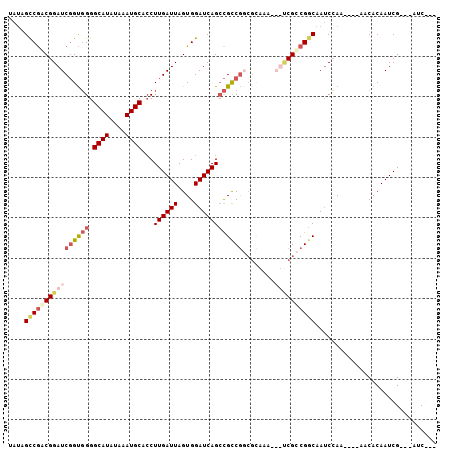
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:51:47 2006