| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 8,282,807 – 8,282,923 |
| Length | 116 |
| Max. P | 0.991156 |

| Location | 8,282,807 – 8,282,923 |
|---|---|
| Length | 116 |
| Sequences | 4 |
| Columns | 118 |
| Reading direction | forward |
| Mean pairwise identity | 92.88 |
| Mean single sequence MFE | -27.81 |
| Consensus MFE | -26.19 |
| Energy contribution | -26.00 |
| Covariance contribution | -0.19 |
| Combinations/Pair | 1.04 |
| Mean z-score | -1.53 |
| Structure conservation index | 0.94 |
| SVM decision value | 0.57 |
| SVM RNA-class probability | 0.783702 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
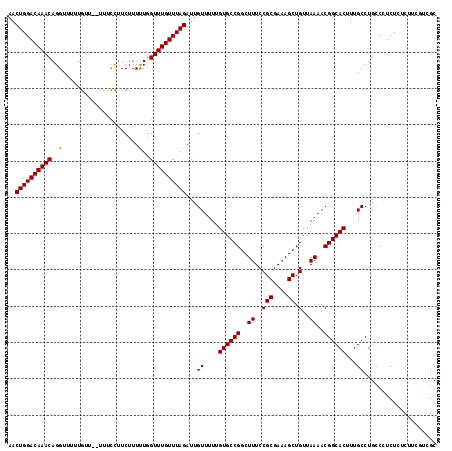
>X_DroMel_CAF1 8282807 116 + 22224390 AACUGGACAAACAGGUUUUUGUU--UUUCCUUCUUUUUAGUUUGUUUAGAUUGUUUUUGUGCCGGCUUUCCGCGAAAGCUGUUAAAACGGCACUUUGCCUGUCCUCUCUCUCCUUCGC ..(((((((((((((........--...)))........))))))))))...((....((((((..((..(((....)).)..))..))))))...)).................... ( -27.10) >DroSec_CAF1 4197 116 + 1 AACUGGACAAACAGGUUUUUGUU--UUUCCUUCUUUUUGGUUUGUUUAGAUUGUUUUUGUGCCGGCUUUCCGCGAAAGCUGUUAAAACGGCACUUUGCCUGCCCCCUGUCUUCGUAGC ..(((((((((((((........--...)))........))))))))))...((....((((((..((..(((....)).)..))..))))))...))((((...........)))). ( -27.70) >DroSim_CAF1 4160 116 + 1 AACUGGACAAACAGGUUUUUGUU--UUUCCUUCUUUUUGGUUUGUUUAGAUUGUUUUUGUGCCGGCUUUCCGCGAAAGCUGUUAAAACGGCACUUUGCCUGCCCUCUCUCUUCGUGGC ..(((((((((((((........--...)))........)))))))))).........((((((..((..(((....)).)..))..))))))...(((................))) ( -28.69) >DroEre_CAF1 7320 115 + 1 AACUGGACAAACAGGUUUUUGUUUUUUUUUGUCUUUCUGGUUUGUUUAGAUUGUUUUUGUGCCGGCUUUCCGCGAAAGCUGUUAAAACGGCACUUUGCCUGCCCUCUCUC---UUCGC ..(((((((((((((....................))).))))))))))...((....((((((..((..(((....)).)..))..))))))...))............---..... ( -27.75) >consensus AACUGGACAAACAGGUUUUUGUU__UUUCCUUCUUUUUGGUUUGUUUAGAUUGUUUUUGUGCCGGCUUUCCGCGAAAGCUGUUAAAACGGCACUUUGCCUGCCCUCUCUCUUCGUCGC ..((((((((((..(.....................)..))))))))))...((....((((((..((..(((....)).)..))..))))))...)).................... (-26.19 = -26.00 + -0.19)


| Location | 8,282,807 – 8,282,923 |
|---|---|
| Length | 116 |
| Sequences | 4 |
| Columns | 118 |
| Reading direction | reverse |
| Mean pairwise identity | 92.88 |
| Mean single sequence MFE | -27.55 |
| Consensus MFE | -26.43 |
| Energy contribution | -26.25 |
| Covariance contribution | -0.19 |
| Combinations/Pair | 1.04 |
| Mean z-score | -2.36 |
| Structure conservation index | 0.96 |
| SVM decision value | 2.25 |
| SVM RNA-class probability | 0.991156 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
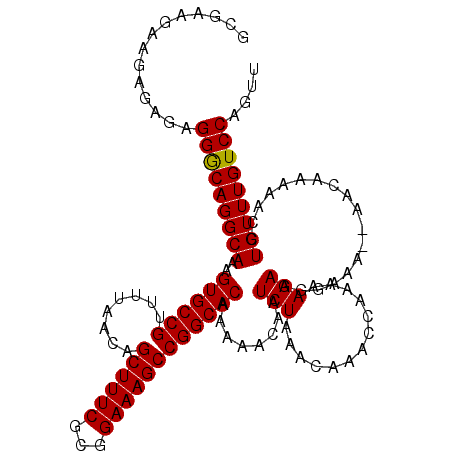
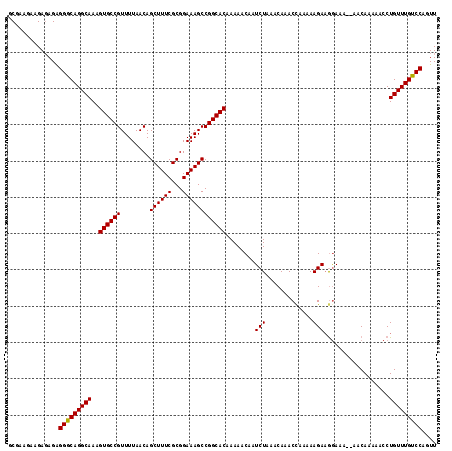
>X_DroMel_CAF1 8282807 116 - 22224390 GCGAAGGAGAGAGAGGACAGGCAAAGUGCCGUUUUAACAGCUUUCGCGGAAAGCCGGCACAAAAACAAUCUAAACAAACUAAAAAGAAGGAAA--AACAAAAACCUGUUUGUCCAGUU ..............((((((((...((((((........((((((...))))))))))))...........................(((...--........))))))))))).... ( -27.60) >DroSec_CAF1 4197 116 - 1 GCUACGAAGACAGGGGGCAGGCAAAGUGCCGUUUUAACAGCUUUCGCGGAAAGCCGGCACAAAAACAAUCUAAACAAACCAAAAAGAAGGAAA--AACAAAAACCUGUUUGUCCAGUU ..............(((((((((..((((((........((((((...))))))))))))..................((........))...--..........))))))))).... ( -28.00) >DroSim_CAF1 4160 116 - 1 GCCACGAAGAGAGAGGGCAGGCAAAGUGCCGUUUUAACAGCUUUCGCGGAAAGCCGGCACAAAAACAAUCUAAACAAACCAAAAAGAAGGAAA--AACAAAAACCUGUUUGUCCAGUU ..............(((((((((..((((((........((((((...))))))))))))..................((........))...--..........))))))))).... ( -28.20) >DroEre_CAF1 7320 115 - 1 GCGAA---GAGAGAGGGCAGGCAAAGUGCCGUUUUAACAGCUUUCGCGGAAAGCCGGCACAAAAACAAUCUAAACAAACCAGAAAGACAAAAAAAAACAAAAACCUGUUUGUCCAGUU .....---......(((((((((..((((((........((((((...))))))))))))........(((.........)))......................))))))))).... ( -26.40) >consensus GCGAAGAAGAGAGAGGGCAGGCAAAGUGCCGUUUUAACAGCUUUCGCGGAAAGCCGGCACAAAAACAAUCUAAACAAACCAAAAAGAAGGAAA__AACAAAAACCUGUUUGUCCAGUU ..............(((((((((..((((((........((((((...))))))))))))........(((.............)))..................))))))))).... (-26.43 = -26.25 + -0.19)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:51:06 2006