
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 8,282,631 – 8,282,734 |
| Length | 103 |
| Max. P | 0.996073 |

| Location | 8,282,631 – 8,282,734 |
|---|---|
| Length | 103 |
| Sequences | 3 |
| Columns | 104 |
| Reading direction | forward |
| Mean pairwise identity | 79.22 |
| Mean single sequence MFE | -25.83 |
| Consensus MFE | -18.49 |
| Energy contribution | -19.93 |
| Covariance contribution | 1.45 |
| Combinations/Pair | 1.05 |
| Mean z-score | -1.41 |
| Structure conservation index | 0.72 |
| SVM decision value | 0.01 |
| SVM RNA-class probability | 0.539452 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
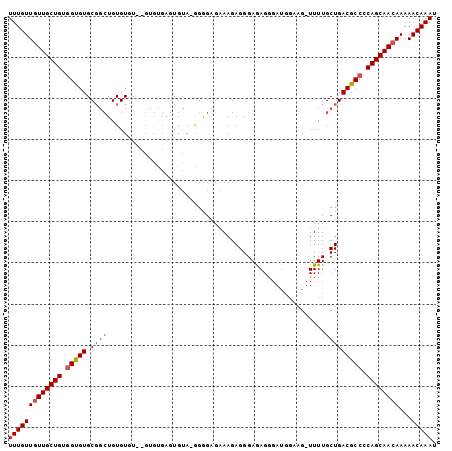
>X_DroMel_CAF1 8282631 103 + 22224390 UUUGUUGUUGCUGUGGUGUGCGGCUGUGUGUGUGUGUGAGUGUA-GGAGCGAAAGAGGGAAAGGGAUGGUAGUUUUUGCUGACGCCCCAGCAACAAAAACAAAU (((((((((((((.(((((.(((((((.(.((..(....)..))-.).)))..........((((((....))))))))))))))).))))))))...))))). ( -29.20) >DroSec_CAF1 4030 94 + 1 UUUGUUGUUGCUGUGGUGUGCGGCUGUGU----GUGUGAGUGUA-GGGGAGAAAGAGGGAGAGG----GAAG-UUUUGCUGACGCCCCAGCAACAAAAACAAAU (((((((((((((.(((((....((.((.----.(....)..))-.)).......((.((((..----....-)))).)).))))).))))))))...))))). ( -25.80) >DroEre_CAF1 7156 92 + 1 UUUGUUGUUGCUGUAGCGUGCCUGUGUGUGU---------UGAAGAGGGAGGAAGAGGGAGAGAGAUGGAAG-UUUUGCUGACGCC-CAGCAA-AAAAACAAAU ((((((.((((((..((((.(((.(.(.(..---------...).).).))).......((..((((....)-)))..)).)))).-))))))-...)))))). ( -22.50) >consensus UUUGUUGUUGCUGUGGUGUGCGGCUGUGUGU__GUGUGAGUGUA_GGGGAGAAAGAGGGAGAGGGAUGGAAG_UUUUGCUGACGCCCCAGCAACAAAAACAAAU (((((((((((((.(((((.((((.....................................................))))))))).))))))))...))))). (-18.49 = -19.93 + 1.45)


| Location | 8,282,631 – 8,282,734 |
|---|---|
| Length | 103 |
| Sequences | 3 |
| Columns | 104 |
| Reading direction | reverse |
| Mean pairwise identity | 79.22 |
| Mean single sequence MFE | -16.45 |
| Consensus MFE | -12.86 |
| Energy contribution | -12.97 |
| Covariance contribution | 0.11 |
| Combinations/Pair | 1.05 |
| Mean z-score | -2.64 |
| Structure conservation index | 0.78 |
| SVM decision value | 2.65 |
| SVM RNA-class probability | 0.996073 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
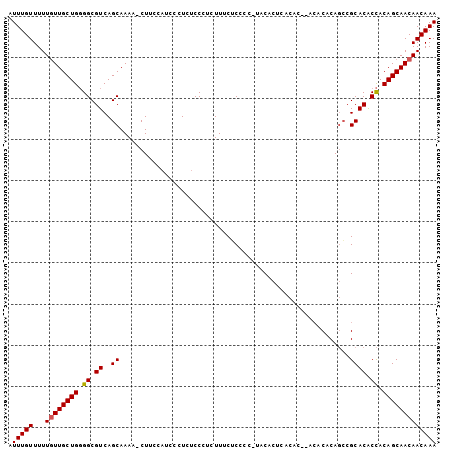
>X_DroMel_CAF1 8282631 103 - 22224390 AUUUGUUUUUGUUGCUGGGGCGUCAGCAAAAACUACCAUCCCUUUCCCUCUUUCGCUCC-UACACUCACACACACACACAGCCGCACACCACAGCAACAACAAA .(((((...((((((((((((....))...........................((..(-(..................))..))...)).))))))))))))) ( -17.77) >DroSec_CAF1 4030 94 - 1 AUUUGUUUUUGUUGCUGGGGCGUCAGCAAAA-CUUC----CCUCUCCCUCUUUCUCCCC-UACACUCACAC----ACACAGCCGCACACCACAGCAACAACAAA .(((((...((((((((.((.(((.((....-....----...................-...........----.....)).).)).)).))))))))))))) ( -17.13) >DroEre_CAF1 7156 92 - 1 AUUUGUUUUU-UUGCUG-GGCGUCAGCAAAA-CUUCCAUCUCUCUCCCUCUUCCUCCCUCUUCA---------ACACACACAGGCACGCUACAGCAACAACAAA .((((((...-((((((-(((((..((....-................................---------..........))))))).)))))).)))))) ( -14.45) >consensus AUUUGUUUUUGUUGCUGGGGCGUCAGCAAAA_CUUCCAUCCCUCUCCCUCUUUCUCCCC_UACACUCACAC__ACACACAGCCGCACACCACAGCAACAACAAA .(((((...((((((((.((.((..((........................................................)))).)).))))))))))))) (-12.86 = -12.97 + 0.11)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:51:05 2006