
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 8,257,692 – 8,257,792 |
| Length | 100 |
| Max. P | 0.986186 |

| Location | 8,257,692 – 8,257,792 |
|---|---|
| Length | 100 |
| Sequences | 3 |
| Columns | 101 |
| Reading direction | forward |
| Mean pairwise identity | 92.05 |
| Mean single sequence MFE | -29.77 |
| Consensus MFE | -27.33 |
| Energy contribution | -26.23 |
| Covariance contribution | -1.10 |
| Combinations/Pair | 1.21 |
| Mean z-score | -3.67 |
| Structure conservation index | 0.92 |
| SVM decision value | 2.03 |
| SVM RNA-class probability | 0.986186 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 8257692 100 + 22224390 U-UUCAGCAUGCUUACUCACGCACACACUUGUGCGCUUUAAACGCUUACGGUUCUGUGUGUGCGUGUAUGUAAAUGUCUAAUACUUGUGUCAAUUUUCCCU .-....((((..((((.((((((((((((((((.((.......)).)))))....)))))))))))...))))))))........................ ( -26.70) >DroSec_CAF1 7972 101 + 1 UUUUCAGCAUGCUCACUCACGCACACACUUGUACGCUUUAAACGCUUGCGGUUCUGUGUGUGCGUGUAUGUAAAUGUCUAAUACUUAUGUCAAUUUUCCCU ......((((...((..(((((((((((.....(((...........))).....)))))))))))..))...))))........................ ( -29.40) >DroYak_CAF1 12605 98 + 1 U-UUCAGCAUGCUCACUCACGCACACAUUUGUACGCUUUAAACGCGUGCGGUU--GUGUGUGUGUGUAUGUAAAUGUCUAAUACUUGUGUCAAUUUUUCCA .-....((((((.....((((((((((.((((((((.......)))))))).)--))))))))).)))))).............................. ( -33.20) >consensus U_UUCAGCAUGCUCACUCACGCACACACUUGUACGCUUUAAACGCUUGCGGUUCUGUGUGUGCGUGUAUGUAAAUGUCUAAUACUUGUGUCAAUUUUCCCU ......((((...((..((((((((((((((((.((.......)).)))))....)))))))))))..))...))))........................ (-27.33 = -26.23 + -1.10)


| Location | 8,257,692 – 8,257,792 |
|---|---|
| Length | 100 |
| Sequences | 3 |
| Columns | 101 |
| Reading direction | reverse |
| Mean pairwise identity | 92.05 |
| Mean single sequence MFE | -26.23 |
| Consensus MFE | -23.32 |
| Energy contribution | -24.10 |
| Covariance contribution | 0.78 |
| Combinations/Pair | 1.04 |
| Mean z-score | -2.61 |
| Structure conservation index | 0.89 |
| SVM decision value | 1.23 |
| SVM RNA-class probability | 0.934175 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 8257692 100 - 22224390 AGGGAAAAUUGACACAAGUAUUAGACAUUUACAUACACGCACACACAGAACCGUAAGCGUUUAAAGCGCACAAGUGUGUGCGUGAGUAAGCAUGCUGAA-A ....................((((.(((((((...(((((((((((.(...)....((((.....))))....))))))))))).))))..))))))).-. ( -27.40) >DroSec_CAF1 7972 101 - 1 AGGGAAAAUUGACAUAAGUAUUAGACAUUUACAUACACGCACACACAGAACCGCAAGCGUUUAAAGCGUACAAGUGUGUGCGUGAGUGAGCAUGCUGAAAA ....................((((.(((...(((.(((((((((((.(((((....).))))...(....)..))))))))))).)))...)))))))... ( -29.20) >DroYak_CAF1 12605 98 - 1 UGGAAAAAUUGACACAAGUAUUAGACAUUUACAUACACACACACAC--AACCGCACGCGUUUAAAGCGUACAAAUGUGUGCGUGAGUGAGCAUGCUGAA-A ....................((((.(((...(((.(((.((((((.--....(.((((.......)))).)...)))))).))).)))...))))))).-. ( -22.10) >consensus AGGGAAAAUUGACACAAGUAUUAGACAUUUACAUACACGCACACACAGAACCGCAAGCGUUUAAAGCGUACAAGUGUGUGCGUGAGUGAGCAUGCUGAA_A ....................((((.(((.(.(((.(((((((((((.....(((...........))).....))))))))))).))).).)))))))... (-23.32 = -24.10 + 0.78)
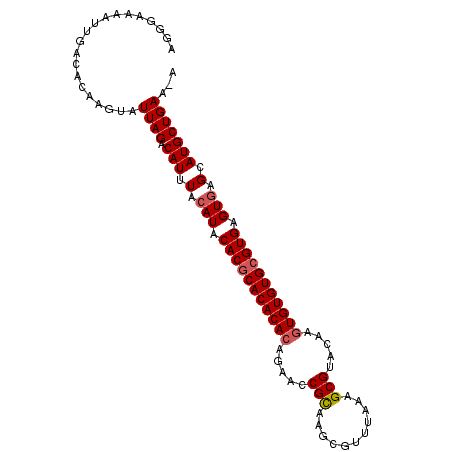



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:50:44 2006