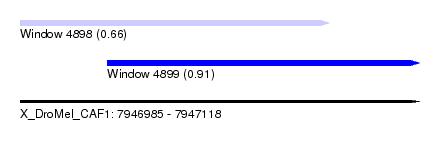

| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 7,946,985 – 7,947,118 |
| Length | 133 |
| Max. P | 0.912087 |

| Location | 7,946,985 – 7,947,088 |
|---|---|
| Length | 103 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 78.81 |
| Mean single sequence MFE | -21.23 |
| Consensus MFE | -13.89 |
| Energy contribution | -14.95 |
| Covariance contribution | 1.06 |
| Combinations/Pair | 1.06 |
| Mean z-score | -1.73 |
| Structure conservation index | 0.65 |
| SVM decision value | 0.27 |
| SVM RNA-class probability | 0.663628 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 7946985 103 + 22224390 G-AGAGAGCACUGUAUCUAA----------AAAGUAUGAGCGACAAAUUGUUGUUGCACAG------AUAUACAUAUGUGUUUACAACAACAACAAUAACAAACUAUUCCACACCCAAAA .-.(.(((((((((((((..----------.......(.(((((((....))))))).)))------))))).....)))))).)................................... ( -14.60) >DroSec_CAF1 9334 119 + 1 G-AGAGAACACUGUACGUAAAAUCAGGCAAAAAGUAUGAGCGACAAAUUGUUGUUGCGUAGCGUAGUACAGAUAUAUGUGUUUGCAACAACAACAAUAACAAACUAUUCCACACCCAAAA (-((((.....(((.(((....(((.((.....)).)))))))))..(((((((((((..(((((.........)))))...)))))))))))..........)).)))........... ( -23.10) >DroSim_CAF1 4007 120 + 1 CCAGAGAACACUGUACGUAAAAUCAGGCAAAAAGUAUGAGCGACAAAUUGUUGUUGCAUAGCGUAGCACAGAUAUAUGUGUUUGCAACAACAACAAUAACAAACUAUUCCACACCCAAAA ...(.(((...(((.(((....(((.((.....)).))))))))).((((((((((....(((.((((((......)))))))))..)))))))))).........)))).......... ( -24.70) >DroYak_CAF1 5216 98 + 1 U-AGAGAGCACCGUACGUAAA---------------------ACGAAUUGUUGUUGCAUAGCGUAGCAUAGGUAUAUGUGUUUGCAACAACAACAAAAACAAGCGAUUCCACACCCAAAA .-.(((.((..(((.......---------------------)))..(((((((((((..(((((((....)).)))))...))))))))))).........))..)))........... ( -22.50) >consensus G_AGAGAACACUGUACGUAAA_________AAAGUAUGAGCGACAAAUUGUUGUUGCAUAGCGUAGCACAGAUAUAUGUGUUUGCAACAACAACAAUAACAAACUAUUCCACACCCAAAA ...............................................(((((((((((..(((((.........)))))...)))))))))))........................... (-13.89 = -14.95 + 1.06)



| Location | 7,947,014 – 7,947,118 |
|---|---|
| Length | 104 |
| Sequences | 4 |
| Columns | 116 |
| Reading direction | forward |
| Mean pairwise identity | 88.50 |
| Mean single sequence MFE | -19.50 |
| Consensus MFE | -15.69 |
| Energy contribution | -16.75 |
| Covariance contribution | 1.06 |
| Combinations/Pair | 1.05 |
| Mean z-score | -2.12 |
| Structure conservation index | 0.80 |
| SVM decision value | 1.08 |
| SVM RNA-class probability | 0.912087 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 7947014 104 + 22224390 CGACAAAUUGUUGUUGCACAG------AUAUACAUAUGUGUUUACAACAACAACAAUAACAAACUAUUCCACACCCAAAA-AAAAGCCGCAAGCUAACA-----CACACGCGCACA ......((((((((((...((------(((((....)))))))....)))))))))).......................-....((.((.........-----.....))))... ( -14.74) >DroSec_CAF1 9373 111 + 1 CGACAAAUUGUUGUUGCGUAGCGUAGUACAGAUAUAUGUGUUUGCAACAACAACAAUAACAAACUAUUCCACACCCAAAAAAAAAGCCGCAAGCUAACA-----CACACGCGCACA .......(((((((((((..(((((.........)))))...)))))))))))................................((.((.........-----.....))))... ( -19.84) >DroSim_CAF1 4047 110 + 1 CGACAAAUUGUUGUUGCAUAGCGUAGCACAGAUAUAUGUGUUUGCAACAACAACAAUAACAAACUAUUCCACACCCAAAA-AAAAGCCGCAAGCUAACA-----CACACGCGCACA ......((((((((((....(((.((((((......)))))))))..)))))))))).......................-....((.((.........-----.....))))... ( -21.44) >DroYak_CAF1 5236 113 + 1 --ACGAAUUGUUGUUGCAUAGCGUAGCAUAGGUAUAUGUGUUUGCAACAACAACAAAAACAAGCGAUUCCACACCCAAAA-AAAAGCCGCAAGCUAACACCUCACACACGCGCACA --.....(((((((((((..(((((((....)).)))))...))))))))))).........(((...............-...((.(....))).........(....))))... ( -22.00) >consensus CGACAAAUUGUUGUUGCAUAGCGUAGCACAGAUAUAUGUGUUUGCAACAACAACAAUAACAAACUAUUCCACACCCAAAA_AAAAGCCGCAAGCUAACA_____CACACGCGCACA .......(((((((((((..(((((.........)))))...)))))))))))...............................((.(....)))..................... (-15.69 = -16.75 + 1.06)

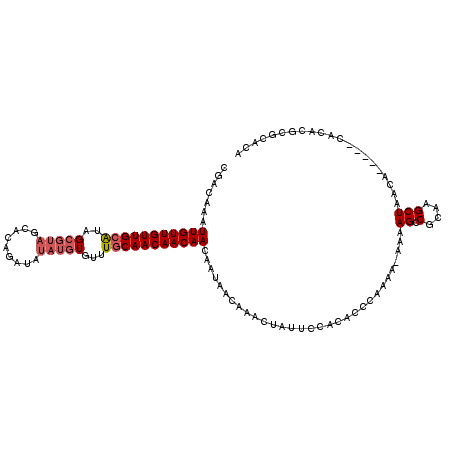

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:48:23 2006