
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 7,944,026 – 7,944,143 |
| Length | 117 |
| Max. P | 0.528494 |

| Location | 7,944,026 – 7,944,143 |
|---|---|
| Length | 117 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 80.18 |
| Mean single sequence MFE | -47.75 |
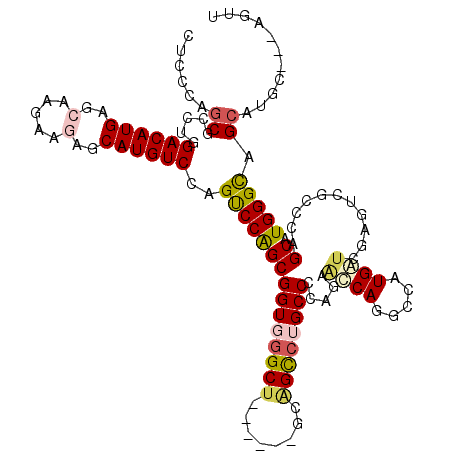
| Consensus MFE | -27.54 |
| Energy contribution | -27.98 |
| Covariance contribution | 0.45 |
| Combinations/Pair | 1.25 |
| Mean z-score | -1.71 |
| Structure conservation index | 0.58 |
| SVM decision value | -0.01 |
| SVM RNA-class probability | 0.528494 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
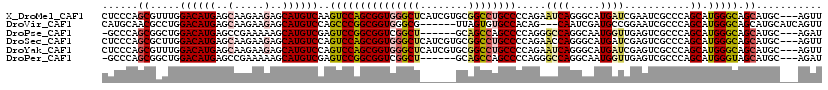
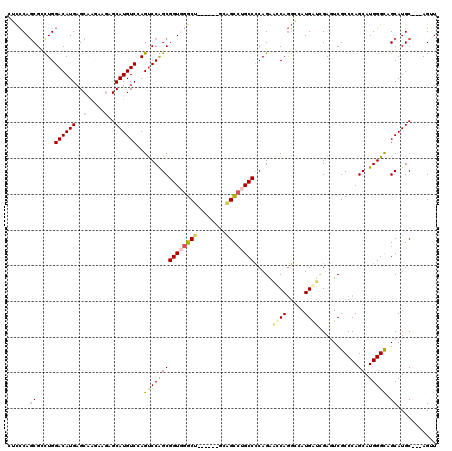
>X_DroMel_CAF1 7944026 117 - 22224390 CUCCCAGCGUUUGGACAUGAGCAAGAAGAGCAUGUCAAGUCCAGCGGUGGGCUCAUCGUGCGGCCUGCCCCAGAAUCAGGGCAUGAUCGAAUCGCCCAGCAUGGGCAGCAUGC---AGUU ..((((.((((.((((.(((((.......))...))).)))))))).)))).....((((((..((((((........))))).)..).....((((.....)))).))))).---.... ( -44.30) >DroVir_CAF1 14896 111 - 1 CAUGCAACGCCUGGACAUGAGCAAGAAGAGCAUGUCCAGCCCGGCGGUGGGCG------UUAGUGUGCCACAG---CAAUCGAUGCCGGAAUCGCCCAGCAUGGGCAGCAUGCAUCAGUU (((((...(((((((((((..(.....)..))))))).((..((((((.((((------(..((.(((....)---))))..)))))...))))))..))..)))).)))))........ ( -46.10) >DroPse_CAF1 6576 110 - 1 -GCCCAGCGGCUGGACAUGAGCCGAAAAAGCAUGUCGAGUCCGGCGGUCGGCU------GCAGCCAGCCCCAGGGCCAGGCAAUGGUUGAGUCGCCCAGCAUGGGCAGCAUGC---AGAU -(((((((((((.((((((...........)))))).)))).(((((((((((------(..(((.(((....)))..)))..))))))).)))))..)).))))).......---.... ( -50.30) >DroSec_CAF1 6398 117 - 1 CUCCCAGCGCUUGGACAUGAGCAAGAAGAGCAUGUCCAGUCCAGCGGUGGGCUCAUCGUGCGGCCUGCCCCAGAACCAGGGCAUGAUCGAGUCGCCCAGCAUGGGCAGCAUGC---AGUU ..((((.((((((((((((..(.....)..))))))))....)))).)))).....((((((..((((((........))))).)..).....((((.....)))).))))).---.... ( -49.70) >DroYak_CAF1 2213 117 - 1 CUCCCAGCGUUUGGACAUGAGCAAGAAGAGCAUGUCCAGUCCAGCGGUGGGCUCAUCGUGCGGCCUGCCCCAGAAUCAGGGCAUGAUCGAGUCGCCCAGCAUGGGCAGCAUGC---AGUU ......((((.((((((((..(.....)..))))))))(((((((((..((((((((((((..((((.........))))))))))).)))))..)).)).)))))...))))---.... ( -48.20) >DroPer_CAF1 5425 110 - 1 -GCCCAGCGGCUGGACAUGAGCCGAAAAAGCAUGUCGAGUCCGGCGGUCGGCU------GCAGCCAGCCCCAGGGCCAGGCAAUGGUUGAGUCGCCCAGCAUGGGUAGCAUGC---AGAU -(((((((((((.((((((...........)))))).)))).(((((((((((------(..(((.(((....)))..)))..))))))).)))))..)).))))).......---.... ( -47.90) >consensus CUCCCAGCGCCUGGACAUGAGCAAGAAGAGCAUGUCCAGUCCAGCGGUGGGCU______GCAGCCUGCCCCAGAACCAGGCCAUGAUCGAGUCGCCCAGCAUGGGCAGCAUGC___AGUU ......((.....((((((..(.....)..))))))..(((((((((((((((........)))))))).....((((.....))))...........)).))))).))........... (-27.54 = -27.98 + 0.45)
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:48:20 2006