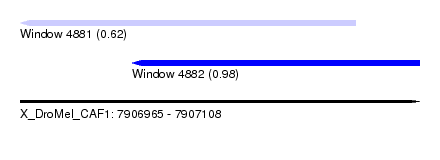
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 7,906,965 – 7,907,108 |
| Length | 143 |
| Max. P | 0.975924 |

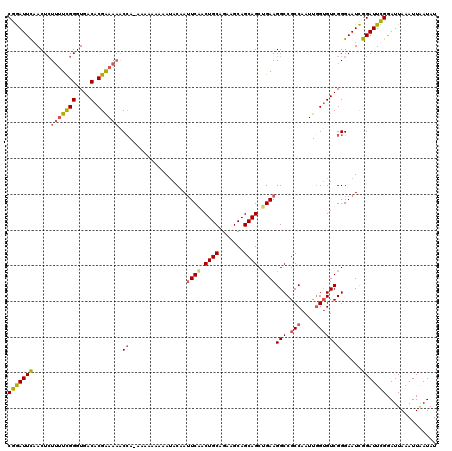
| Location | 7,906,965 – 7,907,085 |
|---|---|
| Length | 120 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 91.10 |
| Mean single sequence MFE | -28.83 |
| Consensus MFE | -21.65 |
| Energy contribution | -22.53 |
| Covariance contribution | 0.88 |
| Combinations/Pair | 1.20 |
| Mean z-score | -1.79 |
| Structure conservation index | 0.75 |
| SVM decision value | 0.18 |
| SVM RNA-class probability | 0.622445 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 7906965 120 - 22224390 CGAAUUUAACUCUUAUCGGGUGACACGAAAAACCAAAAAAAAAAAUACAAUUCGACUGCAGAAGCAGCAGCUGAAGGCCGCCAAUUGGUGUCGGGAAUUGGAUUUGGAUUAAAUUAAUAU ..(((((((..(..(((.(((..(.(((...(((((..............((((.((((.......)))).))))((...))..))))).))))..))).)))..)..)))))))..... ( -28.30) >DroSec_CAF1 7598 119 - 1 CGGAUUCAACUAUUUUCGGGUGACACGAAACGCCA-AAAAAAAAAUACAAUUCAACUGCAGAAGCAGCAGCUGAAGGCCGCCAAUUGGUGUCGGGAAUCGGAUUCGGAUUAAAUUAAUAU (((((((......((((((....).)))))..((.-..............((((.((((.......)))).))))(((.(((....)))))))).....))))))).............. ( -29.70) >DroSim_CAF1 10732 119 - 1 CGGAUUCAACUCUUUUCGGGUGACACGAAAAGCAA-AAAAAAAAAUACAAUUCAACUGCAGAAGCAGCAGCUGAAGGCCGCCAAUUGGUGUCGGGAAUCGGAUUCGGAUUAAAUUAAUAU (((((((....((((((((....).)))))))...-..............((((.((((.......)))).))))(((.(((....)))))).......))))))).............. ( -30.40) >DroYak_CAF1 10870 119 - 1 CGGAUUCAACUCUAUUUGGGUGACACGGAAAACCAAAAAAAAAAAAACAA-UCGACUGCAGAAGCAGCAGCAGAAGGCCGCCAAUUAGAGUCGGGAAUCGGAUUUGGAUUAAAUUGAUAU (.(((((.((((((....((((....((....))................-((..((((.......))))..))....))))...))))))...))))).)................... ( -26.90) >consensus CGGAUUCAACUCUUUUCGGGUGACACGAAAAACCA_AAAAAAAAAUACAAUUCAACUGCAGAAGCAGCAGCUGAAGGCCGCCAAUUGGUGUCGGGAAUCGGAUUCGGAUUAAAUUAAUAU (((((((.....(((((((....).)))))).((................((((.((((.......)))).))))(((.(((....)))))))).....))))))).............. (-21.65 = -22.53 + 0.88)


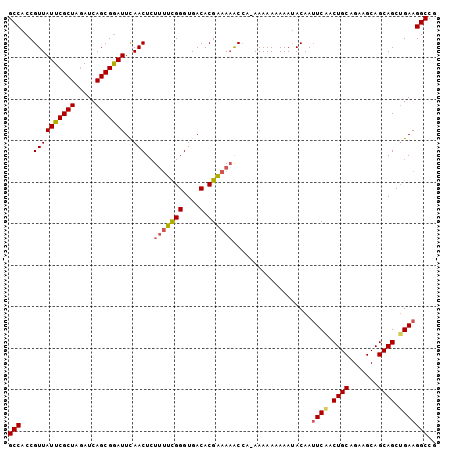
| Location | 7,907,005 – 7,907,108 |
|---|---|
| Length | 103 |
| Sequences | 4 |
| Columns | 103 |
| Reading direction | reverse |
| Mean pairwise identity | 91.73 |
| Mean single sequence MFE | -26.50 |
| Consensus MFE | -23.39 |
| Energy contribution | -23.95 |
| Covariance contribution | 0.56 |
| Combinations/Pair | 1.14 |
| Mean z-score | -2.37 |
| Structure conservation index | 0.88 |
| SVM decision value | 1.76 |
| SVM RNA-class probability | 0.975924 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 7907005 103 - 22224390 GCCACCGUUAUUCGCUAGAUCAGCGAAUUUAACUCUUAUCGGGUGACACGAAAAACCAAAAAAAAAAAUACAAUUCGACUGCAGAAGCAGCAGCUGAAGGCCG (((...((((((((((.....))))))..)))).....((((....).)))......................((((.((((.......)))).))))))).. ( -24.50) >DroSec_CAF1 7638 102 - 1 GCCACCGUUAUUCGCUAGAUCAGCGGAUUCAACUAUUUUCGGGUGACACGAAACGCCA-AAAAAAAAAUACAAUUCAACUGCAGAAGCAGCAGCUGAAGGCCG (((.(((((..((....))..)))))..........((((((....).))))).....-..............((((.((((.......)))).))))))).. ( -26.30) >DroSim_CAF1 10772 102 - 1 GCCACCGUUAUUCGCUAGAUCAGCGGAUUCAACUCUUUUCGGGUGACACGAAAAGCAA-AAAAAAAAAUACAAUUCAACUGCAGAAGCAGCAGCUGAAGGCCG (((.(((((..((....))..)))))........((((((((....).)))))))...-..............((((.((((.......)))).))))))).. ( -29.60) >DroYak_CAF1 10910 102 - 1 GCCACCGUUAUUCGCUAAAUCAGCGGAUUCAACUCUAUUUGGGUGACACGGAAAACCAAAAAAAAAAAAACAA-UCGACUGCAGAAGCAGCAGCAGAAGGCCG (((.((((.(((((((.....)))))))(((.(((.....)))))).))))......................-((..((((.......))))..)).))).. ( -25.60) >consensus GCCACCGUUAUUCGCUAGAUCAGCGGAUUCAACUCUUUUCGGGUGACACGAAAAACCA_AAAAAAAAAUACAAUUCAACUGCAGAAGCAGCAGCUGAAGGCCG (((...((((((((((.....)))))))..)))..(((((((....).))))))...................((((.((((.......)))).))))))).. (-23.39 = -23.95 + 0.56)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:48:08 2006