
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 7,903,979 – 7,904,139 |
| Length | 160 |
| Max. P | 0.953961 |

| Location | 7,903,979 – 7,904,099 |
|---|---|
| Length | 120 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 96.67 |
| Mean single sequence MFE | -38.27 |
| Consensus MFE | -34.73 |
| Energy contribution | -35.07 |
| Covariance contribution | 0.33 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.33 |
| Structure conservation index | 0.91 |
| SVM decision value | 0.86 |
| SVM RNA-class probability | 0.867774 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 7903979 120 + 22224390 GCGGGUCUGAUAAAGAGUCAUGGGAUGGAAGGGGGUGGGAAAGGGGAGCUGCCUCUAGGUUAAUAAAUCGUGCUAAUAGUGGCACGAAUUUAAAGCCGGUUUACAUGGCAACAUGCAGCA (((..(((.....)))((((((........(((((..(..........)..))))).((((..((((((((((((....))))))).))))).))))......))))))....))).... ( -35.60) >DroSec_CAF1 4588 120 + 1 GCGAUUCUGAUAAAGAGGCUUCGGCUGGAAGGGGGUGGGAAAGGGGAGCUGCCUCUAGGUUAAUAAAUCGUGCUAAUAGUGGCACGAAUUUAAAGCCGGUUUACAUGGCAACAUGCAGCA ....((((.....))))(((((((((....(((((..(..........)..))))).......((((((((((((....))))))).))))).))))))....((((....)))).))). ( -38.50) >DroSim_CAF1 7708 120 + 1 GCGAGUCUGAUAAAGAGGCUUCGGCUGGAAGGGGGUGGGAAAGGGGAGCUGCCUCUAGGUUAAUAAAUCGUGCUAAUAGUGGCACGAAUUUAAAGCCGGUUUACAUGGCAACAUGCAGCA .(((((((.......)))).)))((((...(((((..(..........)..))))).((((..((((((((((((....))))))).))))).))))......((((....)))))))). ( -40.70) >consensus GCGAGUCUGAUAAAGAGGCUUCGGCUGGAAGGGGGUGGGAAAGGGGAGCUGCCUCUAGGUUAAUAAAUCGUGCUAAUAGUGGCACGAAUUUAAAGCCGGUUUACAUGGCAACAUGCAGCA ..(..(((.....)))..)....((((...(((((..(..........)..))))).((((..((((((((((((....))))))).))))).))))......((((....)))))))). (-34.73 = -35.07 + 0.33)

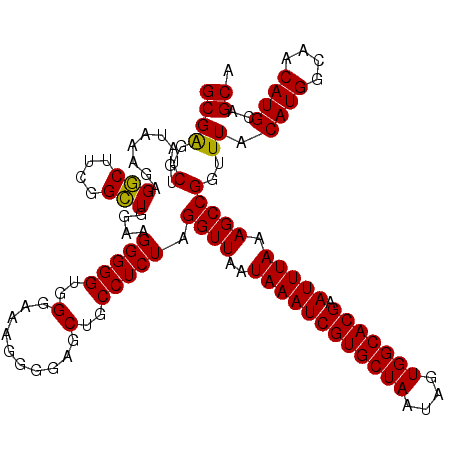
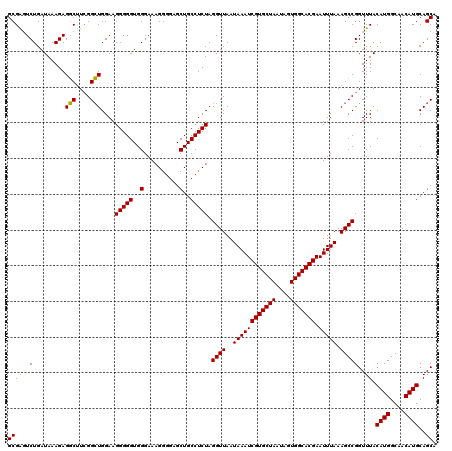
| Location | 7,904,019 – 7,904,139 |
|---|---|
| Length | 120 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 99.03 |
| Mean single sequence MFE | -40.00 |
| Consensus MFE | -39.50 |
| Energy contribution | -39.50 |
| Covariance contribution | 0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -4.21 |
| Structure conservation index | 0.99 |
| SVM decision value | 1.05 |
| SVM RNA-class probability | 0.906780 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 7904019 120 + 22224390 AAGGGGAGCUGCCUCUAGGUUAAUAAAUCGUGCUAAUAGUGGCACGAAUUUAAAGCCGGUUUACAUGGCAACAUGCAGCAAAUGCAGAAGCCGAAGAAGAAGCAGAGGCAUCAGAAGAAG .......(.(((((((.((((..((((((((((((....))))))).))))).))))(((((.((((....))))..((....))..)))))...........))))))).)........ ( -39.50) >DroSec_CAF1 4628 120 + 1 AAGGGGAGCUGCCUCUAGGUUAAUAAAUCGUGCUAAUAGUGGCACGAAUUUAAAGCCGGUUUACAUGGCAACAUGCAGCAAAUGCAGAAGCCGAAGAAGAAGCAGAGGCAUCGCAAGAAG .........(((((((.((((..((((((((((((....))))))).))))).))))(((((.((((....))))..((....))..)))))...........)))))))((....)).. ( -41.50) >DroSim_CAF1 7748 120 + 1 AAGGGGAGCUGCCUCUAGGUUAAUAAAUCGUGCUAAUAGUGGCACGAAUUUAAAGCCGGUUUACAUGGCAACAUGCAGCAAAUGCAGAAGCCGAAGAAGAAGCAGAGGCAUCACAAGAAG .......(.(((((((.((((..((((((((((((....))))))).))))).))))(((((.((((....))))..((....))..)))))...........))))))).)........ ( -39.50) >DroYak_CAF1 7848 120 + 1 AAGGGGAGCUGCCUCUAGGUUAAUAAAUCGUGCUAAUAGUGGCACGAAUUUAAAGCCGGUUUACAUGGCAACAUGCAGCAAAUGCAGAAGCCGAAGAAGAAGCAGAGGCAUCAGAAGAAG .......(.(((((((.((((..((((((((((((....))))))).))))).))))(((((.((((....))))..((....))..)))))...........))))))).)........ ( -39.50) >consensus AAGGGGAGCUGCCUCUAGGUUAAUAAAUCGUGCUAAUAGUGGCACGAAUUUAAAGCCGGUUUACAUGGCAACAUGCAGCAAAUGCAGAAGCCGAAGAAGAAGCAGAGGCAUCACAAGAAG .......(.(((((((.((((..((((((((((((....))))))).))))).))))(((((.((((....))))..((....))..)))))...........))))))).)........ (-39.50 = -39.50 + 0.00)

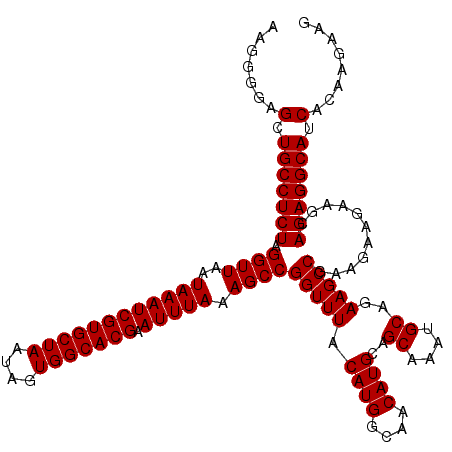

| Location | 7,904,019 – 7,904,139 |
|---|---|
| Length | 120 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 99.03 |
| Mean single sequence MFE | -31.77 |
| Consensus MFE | -31.60 |
| Energy contribution | -31.60 |
| Covariance contribution | -0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.97 |
| Structure conservation index | 0.99 |
| SVM decision value | 1.44 |
| SVM RNA-class probability | 0.953961 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 7904019 120 - 22224390 CUUCUUCUGAUGCCUCUGCUUCUUCUUCGGCUUCUGCAUUUGCUGCAUGUUGCCAUGUAAACCGGCUUUAAAUUCGUGCCACUAUUAGCACGAUUUAUUAACCUAGAGGCAGCUCCCCUU ..........((((((((..........(((..(((((.....)))).)..))).........((...(((((.(((((........))))))))))....))))))))))......... ( -31.60) >DroSec_CAF1 4628 120 - 1 CUUCUUGCGAUGCCUCUGCUUCUUCUUCGGCUUCUGCAUUUGCUGCAUGUUGCCAUGUAAACCGGCUUUAAAUUCGUGCCACUAUUAGCACGAUUUAUUAACCUAGAGGCAGCUCCCCUU ......((..((((((((..........(((..(((((.....)))).)..))).........((...(((((.(((((........))))))))))....))))))))))))....... ( -32.30) >DroSim_CAF1 7748 120 - 1 CUUCUUGUGAUGCCUCUGCUUCUUCUUCGGCUUCUGCAUUUGCUGCAUGUUGCCAUGUAAACCGGCUUUAAAUUCGUGCCACUAUUAGCACGAUUUAUUAACCUAGAGGCAGCUCCCCUU ..........((((((((..........(((..(((((.....)))).)..))).........((...(((((.(((((........))))))))))....))))))))))......... ( -31.60) >DroYak_CAF1 7848 120 - 1 CUUCUUCUGAUGCCUCUGCUUCUUCUUCGGCUUCUGCAUUUGCUGCAUGUUGCCAUGUAAACCGGCUUUAAAUUCGUGCCACUAUUAGCACGAUUUAUUAACCUAGAGGCAGCUCCCCUU ..........((((((((..........(((..(((((.....)))).)..))).........((...(((((.(((((........))))))))))....))))))))))......... ( -31.60) >consensus CUUCUUCUGAUGCCUCUGCUUCUUCUUCGGCUUCUGCAUUUGCUGCAUGUUGCCAUGUAAACCGGCUUUAAAUUCGUGCCACUAUUAGCACGAUUUAUUAACCUAGAGGCAGCUCCCCUU ..........((((((((..........(((..(((((.....)))).)..))).........((...(((((.(((((........))))))))))....))))))))))......... (-31.60 = -31.60 + -0.00)

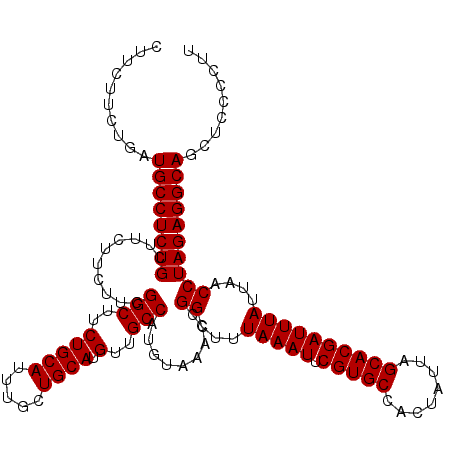
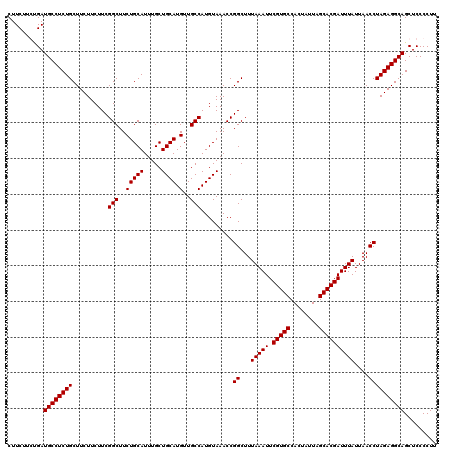
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:47:58 2006