
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 7,900,814 – 7,900,971 |
| Length | 157 |
| Max. P | 0.969745 |

| Location | 7,900,814 – 7,900,931 |
|---|---|
| Length | 117 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 93.51 |
| Mean single sequence MFE | -53.60 |
| Consensus MFE | -42.88 |
| Energy contribution | -44.62 |
| Covariance contribution | 1.75 |
| Combinations/Pair | 1.02 |
| Mean z-score | -4.28 |
| Structure conservation index | 0.80 |
| SVM decision value | 0.73 |
| SVM RNA-class probability | 0.835086 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 7900814 117 + 22224390 UCUCCUCCUUUUCCACUGAUUUGAUCCUCCUUGCGAUUCCUUCGUUAGAGGGUAUGUAGGG-G--GGGUGAAAGUGGGUUUAGGGGGGGGAACUGGUCUUAUGUGGCCGGUGACCACCUC ...((((((..((((((...((.((((((((((((...(((((....)))))..)))))))-)--)))).))))))))...))))))((..(((((((......)))))))..))..... ( -58.60) >DroSec_CAF1 1438 117 + 1 UCUCCUCCCUGUCCUCGGAUUUGAUCCUCCUUGCGAUUCCUUCGUUUGAGGGUAUGUUGGGGG--GGGUG-AAGUGGGUUUAGGGGGGGGAACUGGUCUUAUGUGGCCGGUGACCACCUC (((((((((((..(((..((((.((((((((..((((.(((((....)))))...))))))))--)))).-)))))))..)))))))))))(((((((......)))))))......... ( -54.70) >DroSim_CAF1 4553 115 + 1 UCUCCUCCCUGUCCUCUGAUUUGAUCCUCCUUGCGAUUCCUUCGUUUGAGGGUAUGUUGGG-G--GGGUG-AAGUGGGUUUAGGGG-GGGAACUGGUCUUAUGUGGCCGGUGACCACCUC (((((.(((((..(((..((((.((((((((.(((...(((((....)))))..))).)))-)--)))).-)))))))..))))))-))))(((((((......)))))))......... ( -51.70) >DroYak_CAF1 4575 118 + 1 UCUCCUCCUUGUCCUCUAAUUUGAUCCUCCUUGCGAUUCCUUCGUAUGAGGGUAUGUAGGG-GGAGGGUGAAAAUGGGUUUAGGGG-GGGAACUGGUCUUAUGUGGCCGGUGACCACCUC (((((((((...(((....(((.((((((((((((...(((((....)))))..))))..)-))))))).)))..)))...)))))-))))(((((((......)))))))......... ( -49.40) >consensus UCUCCUCCCUGUCCUCUGAUUUGAUCCUCCUUGCGAUUCCUUCGUUUGAGGGUAUGUAGGG_G__GGGUG_AAGUGGGUUUAGGGG_GGGAACUGGUCUUAUGUGGCCGGUGACCACCUC (((((((((((..(((...(((.((((.(((((((...(((((....)))))..)))))))....)))).)))..)))..)))))))))))(((((((......)))))))......... (-42.88 = -44.62 + 1.75)


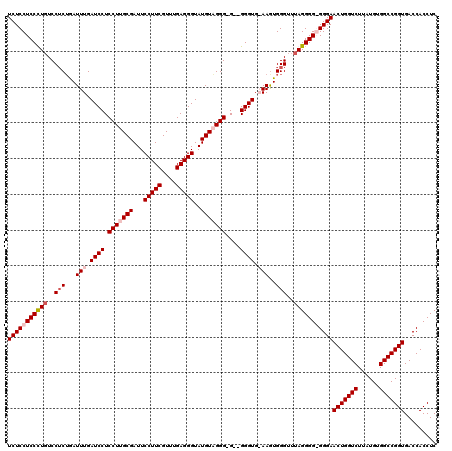
| Location | 7,900,854 – 7,900,971 |
|---|---|
| Length | 117 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 94.08 |
| Mean single sequence MFE | -52.63 |
| Consensus MFE | -45.05 |
| Energy contribution | -45.98 |
| Covariance contribution | 0.94 |
| Combinations/Pair | 1.05 |
| Mean z-score | -3.16 |
| Structure conservation index | 0.86 |
| SVM decision value | 1.65 |
| SVM RNA-class probability | 0.969745 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 7900854 117 + 22224390 UUCGUUAGAGGGUAUGUAGGG-G--GGGUGAAAGUGGGUUUAGGGGGGGGAACUGGUCUUAUGUGGCCGGUGACCACCUCCUCCUCCUGCGACUGAUUACGCCUUCUCCUGCUGUUCUUG .......(((..((.((((((-(--(((((..(((.((((((((((((((((((((((......))))))).......))))))))))).)))).))).))))))).)))))))..))). ( -54.11) >DroSec_CAF1 1478 117 + 1 UUCGUUUGAGGGUAUGUUGGGGG--GGGUG-AAGUGGGUUUAGGGGGGGGAACUGGUCUUAUGUGGCCGGUGACCACCUCCUCCUCCUGCGACUGACUACGCCUUCUCCUGCUGUUCCUG ........((((...((.(((((--(((((-.(((.((((((((((((((((((((((......))))))).......))))))))))).)))).))).)))))))))).))...)))). ( -58.31) >DroSim_CAF1 4593 115 + 1 UUCGUUUGAGGGUAUGUUGGG-G--GGGUG-AAGUGGGUUUAGGGG-GGGAACUGGUCUUAUGUGGCCGGUGACCACCUCCUCCUCCUGCGACUGACUACGCCUUCUCCUGCUGUUCCUG ........((((...((.(((-(--(((((-.(((.((((((((((-(((((((((((......))))))).......))).))))))).)))).))).))))))).)).))...)))). ( -50.11) >DroYak_CAF1 4615 118 + 1 UUCGUAUGAGGGUAUGUAGGG-GGAGGGUGAAAAUGGGUUUAGGGG-GGGAACUGGUCUUAUGUGGCCGGUGACCACCUCCUCCUCCUGCGACUGACAACGCCUACUCCUGCUGUUCCUG ........((((...((((((-(..(((((....(.((((((((((-(((((((((((......))))))).......))).))))))).)))).)...))))).)))))))...)))). ( -48.01) >consensus UUCGUUUGAGGGUAUGUAGGG_G__GGGUG_AAGUGGGUUUAGGGG_GGGAACUGGUCUUAUGUGGCCGGUGACCACCUCCUCCUCCUGCGACUGACUACGCCUUCUCCUGCUGUUCCUG ........((((...((((((....(((((...((.((((((((((((((((((((((......))))))).......))))))))))).)))).))..)))))..))))))...)))). (-45.05 = -45.98 + 0.94)

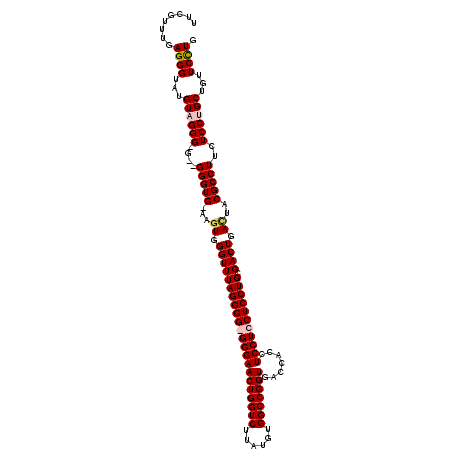
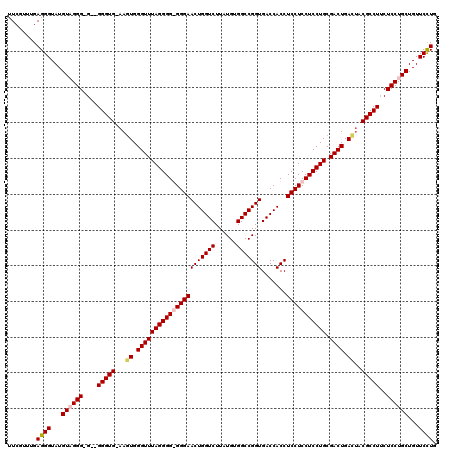
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:47:44 2006