
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 7,887,038 – 7,887,194 |
| Length | 156 |
| Max. P | 0.869936 |

| Location | 7,887,038 – 7,887,154 |
|---|---|
| Length | 116 |
| Sequences | 4 |
| Columns | 116 |
| Reading direction | reverse |
| Mean pairwise identity | 97.13 |
| Mean single sequence MFE | -48.30 |
| Consensus MFE | -44.54 |
| Energy contribution | -44.79 |
| Covariance contribution | 0.25 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.82 |
| Structure conservation index | 0.92 |
| SVM decision value | 0.44 |
| SVM RNA-class probability | 0.736396 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
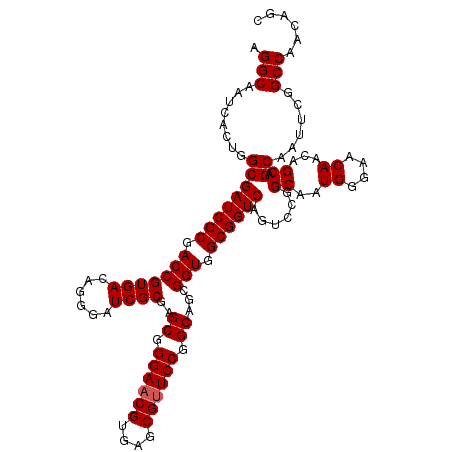
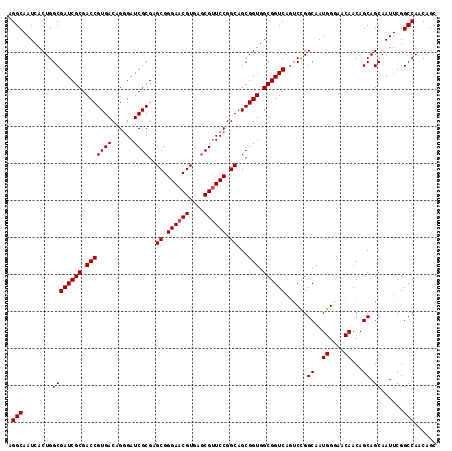
>X_DroMel_CAF1 7887038 116 - 22224390 AGGCAAUCACUGGCGAUCGCGACCGUGACAGGGAUCGCGAGCGGGAACGUGAACGUUCCGGCAGCGGUGGCGGUCAGUCUGGCAAUGGGAACAACAGCAGCAAUUCGGCCAACAGC .(((........((((((((.(((((((......))))..((.((((((....)))))).))...))).)))))).(.(((....((....)).)))).))......)))...... ( -46.54) >DroSec_CAF1 935 116 - 1 AGGCAAUCACUGGCGAUCGCGACCGUGACAGGGAUCGCGAGCGGGAACGUGAGCGAUCCGGCAGCGGUGGCGGUCAGUCCGGCAAUGGGAACAACAGCAGCAAUUCGGCCAACAGC .(((.....(((((((((((.(((((..(..(((((((..(((....)))..))))))).)..))))).)))))).).))))...((....))..............)))...... ( -51.70) >DroEre_CAF1 981 116 - 1 AGGCAAUCACUGGCGAUCGCGACCGUGACAGGGAUCGCGAGCGGGAACGUGAGCGUUCCGGCAGCGGUGGCGGUCAGUCCGGCAAUGGAAACAACAGCAGCAAUUCGGCCAACAGC .(((.....(((((((((((.(((((....((((.(((..(((....)))..)))))))....))))).)))))).).))))...((....))..............)))...... ( -49.30) >DroYak_CAF1 1003 116 - 1 AGGCAAUCACUGGCGAUCGCGACCGUGACAGAGAUCGCGAGCGGGAACGUGAGCGUUCCGGCAGCGGUGGCGGUCAUUCCGGCAAUGGCAACAACAGCAGCAAUUCGGCCAACAGC .(((........((((((((.(((((((......))))..((.((((((....)))))).))...))).))))))......((..((....))...)).))......)))...... ( -45.64) >consensus AGGCAAUCACUGGCGAUCGCGACCGUGACAGGGAUCGCGAGCGGGAACGUGAGCGUUCCGGCAGCGGUGGCGGUCAGUCCGGCAAUGGGAACAACAGCAGCAAUUCGGCCAACAGC .(((........((((((((.(((((((......))))..((.((((((....)))))).))...))).))))))......((..((....))...)).))......)))...... (-44.54 = -44.79 + 0.25)

| Location | 7,887,078 – 7,887,194 |
|---|---|
| Length | 116 |
| Sequences | 4 |
| Columns | 116 |
| Reading direction | reverse |
| Mean pairwise identity | 96.41 |
| Mean single sequence MFE | -56.85 |
| Consensus MFE | -51.47 |
| Energy contribution | -51.10 |
| Covariance contribution | -0.37 |
| Combinations/Pair | 1.08 |
| Mean z-score | -2.07 |
| Structure conservation index | 0.91 |
| SVM decision value | 0.87 |
| SVM RNA-class probability | 0.869936 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
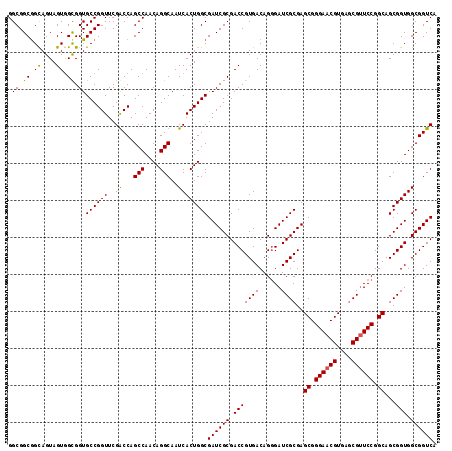
>X_DroMel_CAF1 7887078 116 - 22224390 GGCGGCAGCAGUAGUGGCGGUGCCGGUUCGACCAGCCAACAGGCAAUCACUGGCGAUCGCGACCGUGACAGGGAUCGCGAGCGGGAACGUGAACGUUCCGGCAGCGGUGGCGGUCA ..(((((.(.((....))).)))))....((((.(((....)))..((((((.(..((((((((.......)).))))))((.((((((....)))))).)).))))))).)))). ( -54.80) >DroSec_CAF1 975 116 - 1 GGCGGCGGCAGUAGUGGUGGUGCCGGUUCGACCAGCCAACAGGCAAUCACUGGCGAUCGCGACCGUGACAGGGAUCGCGAGCGGGAACGUGAGCGAUCCGGCAGCGGUGGCGGUCA .(((((((((.((....)).))))((.....)).)))..(((.......)))))((((((.(((((..(..(((((((..(((....)))..))))))).)..))))).)))))). ( -60.20) >DroEre_CAF1 1021 116 - 1 GGCGGCGGCAGUAGUAGCGGUGCCGGUUCGGCCAGCCAACAGGCAAUCACUGGCGAUCGCGACCGUGACAGGGAUCGCGAGCGGGAACGUGAGCGUUCCGGCAGCGGUGGCGGUCA .((.((.((....)).)).))((((((...(((........)))....))))))((((((.(((((....((((.(((..(((....)))..)))))))....))))).)))))). ( -56.80) >DroYak_CAF1 1043 116 - 1 GGCGGCGGUAGUAGUGGCGGUGCCGGUUCGGCCAGCCAACAGGCAAUCACUGGCGAUCGCGACCGUGACAGAGAUCGCGAGCGGGAACGUGAGCGUUCCGGCAGCGGUGGCGGUCA (.(((.(((.((..((((.(.((((...))))).))))....)).))).))).)((((((.(((((((......))))..((.((((((....)))))).))...))).)))))). ( -55.60) >consensus GGCGGCGGCAGUAGUGGCGGUGCCGGUUCGACCAGCCAACAGGCAAUCACUGGCGAUCGCGACCGUGACAGGGAUCGCGAGCGGGAACGUGAGCGUUCCGGCAGCGGUGGCGGUCA .((.((.((....)).)).))((((((..((...(((....)))..))))))))((((((.(((((((......))))..((.((((((....)))))).))...))).)))))). (-51.47 = -51.10 + -0.37)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:47:17 2006