
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 7,702,833 – 7,702,979 |
| Length | 146 |
| Max. P | 0.974068 |

| Location | 7,702,833 – 7,702,939 |
|---|---|
| Length | 106 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 84.26 |
| Mean single sequence MFE | -25.57 |
| Consensus MFE | -16.72 |
| Energy contribution | -17.41 |
| Covariance contribution | 0.69 |
| Combinations/Pair | 1.12 |
| Mean z-score | -1.60 |
| Structure conservation index | 0.65 |
| SVM decision value | 0.01 |
| SVM RNA-class probability | 0.536988 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 7702833 106 - 22224390 GACGCACUCAAGUGUCUAAAA----------ACGAGUAUGUCAUAAGCUCACAUUUGUACAGCUGAGGUCAAAAUGGUAGCAAUUAU----UUGCAAUUAUUUGAAAACAAUAUCAGUAG (((((......))))).....----------..((((.........))))...........(((((.((..(((((((.((((....----)))).)))))))....))....))))).. ( -22.20) >DroSec_CAF1 812 115 - 1 GACGCACUCAAGUGUCUAAAAAUUCCAAAGAACGAGUGUGUCAUAAGCUCACAGUUGCACAGCUGAGGUCAAAAUGGUAGCAAUUGU----UUGCAAUUAUUUGAAAACAAUAUCAG-AA (((((((((.....(((...........)))..)))))))))....((..((((((((.((..((....))...))...))))))))----..)).....(((((........))))-). ( -29.10) >DroSim_CAF1 8982 115 - 1 GACGCACUCAAGUGUCUAAAAAUUCCAAAGAACGAGUGUGUCAUAAGCUCACAGUUGCACAGCUGAGGUCAAAAUGGUAGCAAUUGU----UUGCAAUUAUUUGAAAACAAUAUUAG-AA (((((((((.....(((...........)))..)))))))))....((..((((((((.((..((....))...))...))))))))----..))......................-.. ( -27.40) >DroYak_CAF1 1626 119 - 1 GACGCACUCAAGUGUUUAAAAAUUCCACAGAACGAGUGUGUCAUAAGCUCGCAGUCGCACAGUUGCGGUCAAAAAAGUAGCAUACAUGGCAUUUUAUUUAUUUGAAAUCAAUAUCGG-AA (((((((((..(((...........))).....)))))))))......((((((.(.....)))))))(((((.(((((((.......))....))))).)))))............-.. ( -23.60) >consensus GACGCACUCAAGUGUCUAAAAAUUCCAAAGAACGAGUGUGUCAUAAGCUCACAGUUGCACAGCUGAGGUCAAAAUGGUAGCAAUUAU____UUGCAAUUAUUUGAAAACAAUAUCAG_AA (((((((((........................)))))))))....((........)).....((...(((((.((((.((((........)))).)))))))))...)).......... (-16.72 = -17.41 + 0.69)

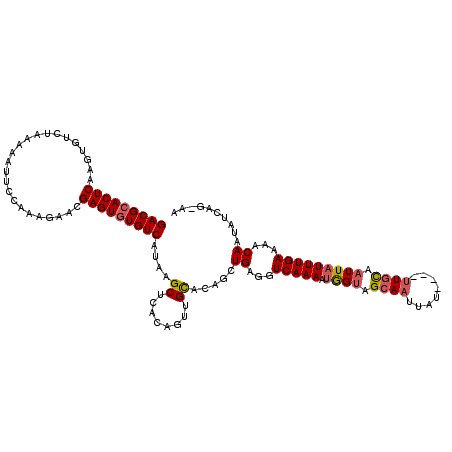

| Location | 7,702,869 – 7,702,979 |
|---|---|
| Length | 110 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 86.94 |
| Mean single sequence MFE | -38.25 |
| Consensus MFE | -30.27 |
| Energy contribution | -30.90 |
| Covariance contribution | 0.63 |
| Combinations/Pair | 1.17 |
| Mean z-score | -2.63 |
| Structure conservation index | 0.79 |
| SVM decision value | 1.72 |
| SVM RNA-class probability | 0.974068 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 7702869 110 + 22224390 CUACCAUUUUGACCUCAGCUGUACAAAUGUGAGCUUAUGACAUACUCGU----------UUUUAGACACUUGAGUGCGUCAGGAGUUCAACGGGUGUGAUUCUAUGUGGCAAAGUGUAUU ....(((((((.((.((....((((..(((((((((.((((.((((((.----------...........)))))).)))).)))))).)))..))))......)).))))))))).... ( -34.70) >DroSec_CAF1 847 120 + 1 CUACCAUUUUGACCUCAGCUGUGCAACUGUGAGCUUAUGACACACUCGUUCUUUGGAAUUUUUAGACACUUGAGUGCGUCAGGAGUUCAACGGGUGUGAUGCUAUGUGGCAAAGUGUAUU ....(((((((.((.((((...(((.((((((((((.((((.((((((...((((((...))))))....)))))).)))).)))))).)))).)))...))..)).))))))))).... ( -39.40) >DroSim_CAF1 9017 120 + 1 CUACCAUUUUGACCUCAGCUGUGCAACUGUGAGCUUAUGACACACUCGUUCUUUGGAAUUUUUAGACACUUGAGUGCGUCAGGAGUUCAACGGGUGUGAUGCUAUGUGGCAAAGUGUAUU ....(((((((.((.((((...(((.((((((((((.((((.((((((...((((((...))))))....)))))).)))).)))))).)))).)))...))..)).))))))))).... ( -39.40) >DroYak_CAF1 1665 120 + 1 CUACUUUUUUGACCGCAACUGUGCGACUGCGAGCUUAUGACACACUCGUUCUGUGGAAUUUUUAAACACUUGAGUGCGUCAGAAGUUCAUGGGGUGCCAUAGUUUGUGGAAAUGUGUAUU ............(((((((((((((.((..((((((.((((.((((((....(((...........))).)))))).)))).))))))..))..)).))))).))))))........... ( -39.50) >consensus CUACCAUUUUGACCUCAGCUGUGCAACUGUGAGCUUAUGACACACUCGUUCUUUGGAAUUUUUAGACACUUGAGUGCGUCAGGAGUUCAACGGGUGUGAUGCUAUGUGGCAAAGUGUAUU ....(((((((.((.((....((((.((((((((((.((((.((((((......................)))))).)))).)))))).)))).))))......)).))))))))).... (-30.27 = -30.90 + 0.63)

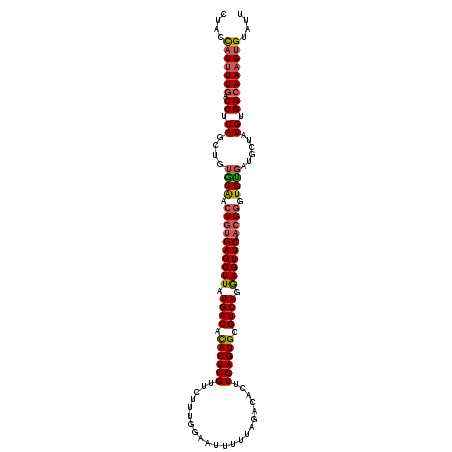
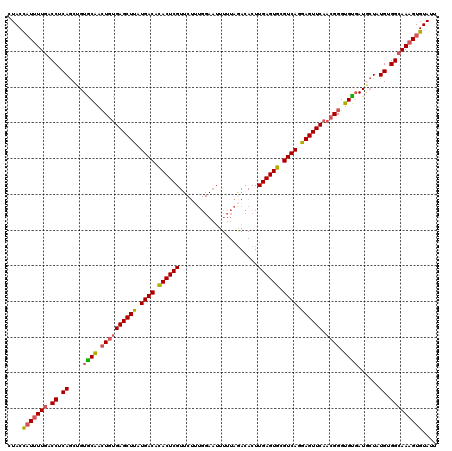
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:45:41 2006