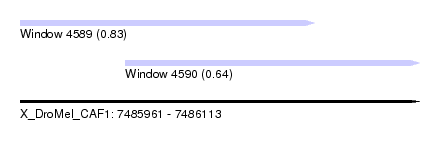

| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 7,485,961 – 7,486,113 |
| Length | 152 |
| Max. P | 0.829715 |

| Location | 7,485,961 – 7,486,073 |
|---|---|
| Length | 112 |
| Sequences | 4 |
| Columns | 113 |
| Reading direction | forward |
| Mean pairwise identity | 97.63 |
| Mean single sequence MFE | -21.42 |
| Consensus MFE | -19.72 |
| Energy contribution | -20.22 |
| Covariance contribution | 0.50 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.09 |
| Structure conservation index | 0.92 |
| SVM decision value | 0.71 |
| SVM RNA-class probability | 0.829715 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
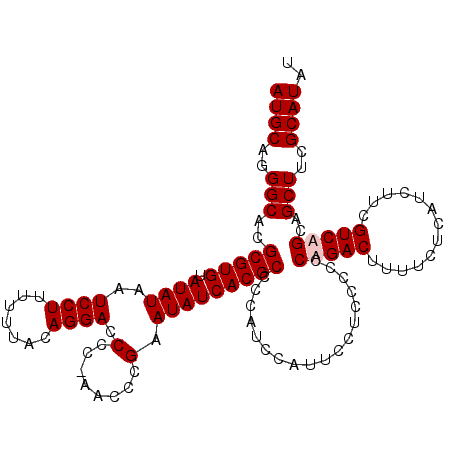
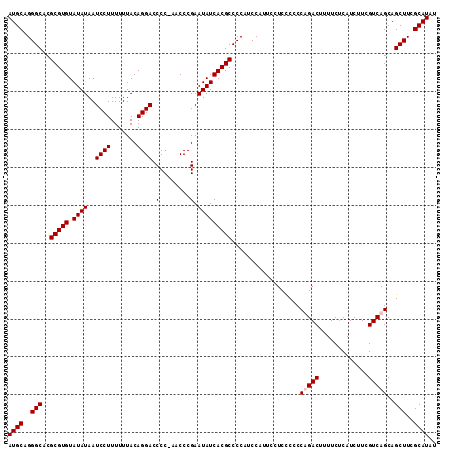
>X_DroMel_CAF1 7485961 112 + 22224390 AUGCAGGGCACGCGUGUAUAUAAUCCUUUUUUACAGGACCCC-AACCCGAAUAUCACGCCCCAUCCAUUCCUCCCCCCUGACUUUUCUCAUCUUCGUCAGCAGCUUCGCAUAU ((((..(((..(((((.((((..((((.......))))...(-.....).)))))))))..................(((((.............)))))..)))..)))).. ( -22.92) >DroSec_CAF1 19011 112 + 1 AUGCAGGGCACGCGUGUAUAUAAUCCUUUUUUACAGGACCCC-AACCCGAAUAUCACGCCCCAUCCAUUCCUCCCCCCAGACUUUUCUCAUCUUCGUCAGCAGCUUCGCAUAU ((((..(((..(((((.((((..((((.......))))...(-.....).)))))))))....................(((.............)))....)))..)))).. ( -19.92) >DroSim_CAF1 20983 112 + 1 AUGCAGGGCACGCGUGUAUAUAAUCCUUUUUUACAGGACCCC-AACCCGAAUAUCACGCCCCAUCCAUUCCUCCCCCCAGACUUUUCUCAUCUUCGUCAGCAGCUUCGCAUAU ((((..(((..(((((.((((..((((.......))))...(-.....).)))))))))....................(((.............)))....)))..)))).. ( -19.92) >DroEre_CAF1 20061 113 + 1 AUGCAGGGCACGCGUGUAUAUAAUCCUUUUUUACAGGACCCCCAACCCGAAUAUCACGCCCCAUCCGUUCCUCUCCCCUGACUUUUCCCAUCUUCGUCAGCAGCUUCGCAUAU ((((..(((..(((((.((((..((((.......))))....(.....).)))))))))..................(((((.............)))))..)))..)))).. ( -22.92) >consensus AUGCAGGGCACGCGUGUAUAUAAUCCUUUUUUACAGGACCCC_AACCCGAAUAUCACGCCCCAUCCAUUCCUCCCCCCAGACUUUUCUCAUCUUCGUCAGCAGCUUCGCAUAU ((((..(((..(((((.((((..((((.......)))).(........).)))))))))..................(((((.............)))))..)))..)))).. (-19.72 = -20.22 + 0.50)

| Location | 7,486,001 – 7,486,113 |
|---|---|
| Length | 112 |
| Sequences | 4 |
| Columns | 113 |
| Reading direction | forward |
| Mean pairwise identity | 97.19 |
| Mean single sequence MFE | -17.57 |
| Consensus MFE | -16.42 |
| Energy contribution | -16.42 |
| Covariance contribution | -0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.67 |
| Structure conservation index | 0.93 |
| SVM decision value | 0.23 |
| SVM RNA-class probability | 0.644647 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
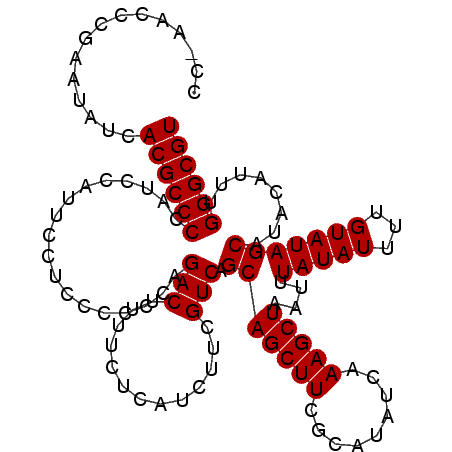
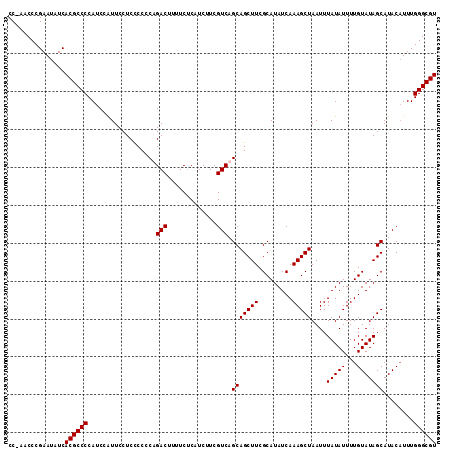
>X_DroMel_CAF1 7486001 112 + 22224390 CC-AACCCGAAUAUCACGCCCCAUCCAUUCCUCCCCCCUGACUUUUCUCAUCUUCGUCAGCAGCUUCGCAUAUCAAAGCUAAUUUAUAUUUUGUAUAGCAUACUUUUGGGCGU ..-............((((((................(((((.............)))))..(((..(((.....................)))..)))........)))))) ( -18.62) >DroSec_CAF1 19051 112 + 1 CC-AACCCGAAUAUCACGCCCCAUCCAUUCCUCCCCCCAGACUUUUCUCAUCUUCGUCAGCAGCUUCGCAUAUCAAAGCUAAUUUAUAUUUUGUAUAGCAUACAUUUGGGCGU ..-............((((((..................(((.............))).(((((((.........)))))....(((((...)))))))........)))))) ( -16.52) >DroSim_CAF1 21023 112 + 1 CC-AACCCGAAUAUCACGCCCCAUCCAUUCCUCCCCCCAGACUUUUCUCAUCUUCGUCAGCAGCUUCGCAUAUCAAAGCUAAUUUAUAUUUUGUAUAGCAUACAUUUGGGCGU ..-............((((((..................(((.............))).(((((((.........)))))....(((((...)))))))........)))))) ( -16.52) >DroEre_CAF1 20101 113 + 1 CCCAACCCGAAUAUCACGCCCCAUCCGUUCCUCUCCCCUGACUUUUCCCAUCUUCGUCAGCAGCUUCGCAUAUCAAAGCUAAUUUAUAUUUUGUAUAGCAUACAUUUGGGCGU ...............((((((................(((((.............)))))..(((..(((.....................)))..)))........)))))) ( -18.62) >consensus CC_AACCCGAAUAUCACGCCCCAUCCAUUCCUCCCCCCAGACUUUUCUCAUCUUCGUCAGCAGCUUCGCAUAUCAAAGCUAAUUUAUAUUUUGUAUAGCAUACAUUUGGGCGU ...............((((((..................(((.............))).(((((((.........)))))....(((((...)))))))........)))))) (-16.42 = -16.42 + -0.00)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:43:45 2006