
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 7,470,645 – 7,470,932 |
| Length | 287 |
| Max. P | 0.985092 |

| Location | 7,470,645 – 7,470,738 |
|---|---|
| Length | 93 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 85.38 |
| Mean single sequence MFE | -22.09 |
| Consensus MFE | -18.97 |
| Energy contribution | -19.37 |
| Covariance contribution | 0.40 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.31 |
| Structure conservation index | 0.86 |
| SVM decision value | 2.00 |
| SVM RNA-class probability | 0.985092 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 7470645 93 + 22224390 CUUUUCACUUCCU------------------CCCCCUG--GUCCUCGUACUU-------CUUCUUUCUCCACAACUCGUUAUUACGACGAGUACGAAGGACGACGCCUGCACUGGGUAUC .............------------------.......--((((((((((((-------................((((....)))).))))))).)))))((..(((.....)))..)) ( -21.03) >DroSec_CAF1 3904 93 + 1 CUCUUCACUUCCU------------------CCCCGUG--GUCCUCGUACUU-------CUUCUUUCUCCACAACUCGUUAUUACGACGAGUACGAAGGACGACGCCUGCACUGGGUAUC .............------------------....((.--((((((((((((-------................((((....)))).))))))).))))).))((((.....))))... ( -23.53) >DroSim_CAF1 5617 93 + 1 CUUUUCACUUCCU------------------CCCCGUG--GUCCUCGUACUU-------CUUCUUUCUCCACAACUCGUUAUUACGACGAGUACGAAGGACGACGCCUGCACUGGGUAUC .............------------------....((.--((((((((((((-------................((((....)))).))))))).))))).))((((.....))))... ( -23.53) >DroEre_CAF1 4245 97 + 1 CUUUUCACUACC--------------CCUCCCCCCAUG--GUCCUCCUACUU-------CUUCUUUCUCCACAACUCGUUAUUACGACGAGUACGAAGGACGACGCCUGCACUGGGUAUC ........((((--------------(..........(--(....))....(-------(.((((((......(((((((.....)))))))..)))))).))..........))))).. ( -19.10) >DroYak_CAF1 6328 120 + 1 CUUUUCACUUCCUCAUCUGCCCCCCCCCCCCCCCUUUGGUGUCCUCCUACUUCUCCCUUCUUCUUUCCCCACAACUCGUUAUUACGACGAGUACGAAGGACGACGCCUGCACUGGGUAUC ............................(((......((((((.((((.........................(((((((.....)))))))....)))).))))))......))).... ( -23.25) >consensus CUUUUCACUUCCU__________________CCCCGUG__GUCCUCGUACUU_______CUUCUUUCUCCACAACUCGUUAUUACGACGAGUACGAAGGACGACGCCUGCACUGGGUAUC ........................................((((((((((((.......................((((....)))).))))))).)))))((..(((.....)))..)) (-18.97 = -19.37 + 0.40)

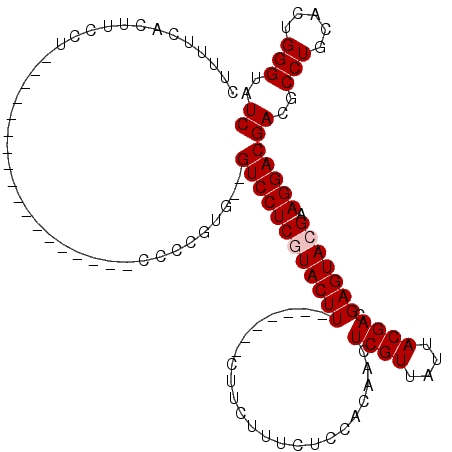

| Location | 7,470,665 – 7,470,774 |
|---|---|
| Length | 109 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 93.21 |
| Mean single sequence MFE | -24.57 |
| Consensus MFE | -23.03 |
| Energy contribution | -23.43 |
| Covariance contribution | 0.40 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.14 |
| Structure conservation index | 0.94 |
| SVM decision value | 0.31 |
| SVM RNA-class probability | 0.681917 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 7470665 109 + 22224390 GUCCUCGUACUU-------CUUCUUUCUCCACAACUCGUUAUUACGACGAGUACGAAGGACGACGCCUGCACUGGGUAUCCUCUUUUCUCUUCUCCGGCUGUCCAUGUCCGUUCCG---- ((((((((((((-------................((((....)))).))))))).)))))((((...(((.(((..(.((...............)).)..)))))).))))...---- ( -25.09) >DroSec_CAF1 3924 109 + 1 GUCCUCGUACUU-------CUUCUUUCUCCACAACUCGUUAUUACGACGAGUACGAAGGACGACGCCUGCACUGGGUAUCCUCUUUUCUCUUCUCCGGCUGUCCAUGUCCGUUCCG---- ((((((((((((-------................((((....)))).))))))).)))))((((...(((.(((..(.((...............)).)..)))))).))))...---- ( -25.09) >DroSim_CAF1 5637 109 + 1 GUCCUCGUACUU-------CUUCUUUCUCCACAACUCGUUAUUACGACGAGUACGAAGGACGACGCCUGCACUGGGUAUCCUCUUUUCUCUUCUCCGGCUGUCCAUGUCCGUUCAG---- ((((((((((((-------................((((....)))).))))))).)))))((((...(((.(((..(.((...............)).)..)))))).))))...---- ( -25.09) >DroEre_CAF1 4269 109 + 1 GUCCUCCUACUU-------CUUCUUUCUCCACAACUCGUUAUUACGACGAGUACGAAGGACGACGCCUGCACUGGGUAUCCUCCUCACUCUUCUCCGGCUGUCCAUGUCCGUUCCG---- ............-------..............(((((((.....)))))))..(((((((((((((.(....((((.........))))....).))).)))...)))).)))..---- ( -22.80) >DroYak_CAF1 6368 120 + 1 GUCCUCCUACUUCUCCCUUCUUCUUUCCCCACAACUCGUUAUUACGACGAGUACGAAGGACGACGCCUGCACUGGGUAUCCUCUUUUCUCUGCUCCGGCUGUCCAUGUCCGUUCUGUUCC .................................(((((((.....)))))))..(((((((((((((.(((..(((.........)))..)))...))).)))...)))).)))...... ( -24.80) >consensus GUCCUCGUACUU_______CUUCUUUCUCCACAACUCGUUAUUACGACGAGUACGAAGGACGACGCCUGCACUGGGUAUCCUCUUUUCUCUUCUCCGGCUGUCCAUGUCCGUUCCG____ ((((((((((((.......................((((....)))).))))))).)))))((((...(((.(((..(.((...............)).)..)))))).))))....... (-23.03 = -23.43 + 0.40)

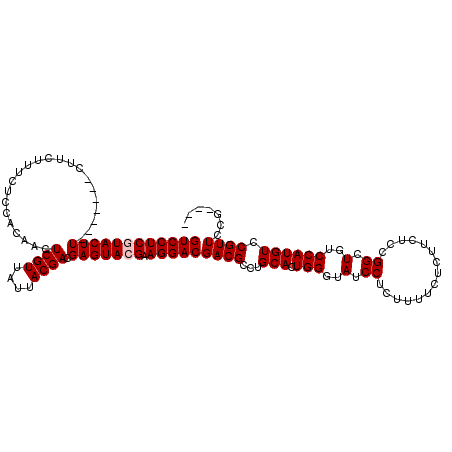

| Location | 7,470,774 – 7,470,869 |
|---|---|
| Length | 95 |
| Sequences | 4 |
| Columns | 104 |
| Reading direction | reverse |
| Mean pairwise identity | 87.60 |
| Mean single sequence MFE | -31.62 |
| Consensus MFE | -22.14 |
| Energy contribution | -21.89 |
| Covariance contribution | -0.25 |
| Combinations/Pair | 1.14 |
| Mean z-score | -2.60 |
| Structure conservation index | 0.70 |
| SVM decision value | 0.99 |
| SVM RNA-class probability | 0.895931 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 7470774 95 - 22224390 GUGUUGAAGGACCAAAAGGAUGGGACUUUUAUAGGAGGCCC---UAUGUAUGGGGUAGGGUCUUUC------GGAGAGUCAGUGUCGACGCACUCAAAACUGAA (((((((....(((......)))(((((((...((((((((---(((.......))))))))))).------.)))))))....)))))))............. ( -33.60) >DroSec_CAF1 4033 95 - 1 GUGUUGAAGGACCAAAAGGAUGGGACUUUUAUAGGAGGCCC---UAUAUAUGGGGUAGGGUCUUUC------GGGGAGUCAGUGUCGACGCACUCAGAACUGAA (((((((....(((......)))(((((((...((((((((---(((.......))))))))))).------.)))))))....)))))))..(((....))). ( -33.50) >DroSim_CAF1 5746 95 - 1 GUGUUGAAGGACCAAAAGGAUGGGACUUUUAUAGGAGGCCC---UAUAUAUGGGGUAGGGUCUUUC------GGGGAGUCAGUGUCGACGCACUCAGAACUGAA (((((((....(((......)))(((((((...((((((((---(((.......))))))))))).------.)))))))....)))))))..(((....))). ( -33.50) >DroEre_CAF1 4378 99 - 1 GUGUUGAAGGACCAAAAGGAUGGGAGUUUUAUAGGGGGC-UUCU---CU-AUGGGUAGGGUCUUCAGUGGGGGGGGAGUCAGUGUCGACGCACUCAGAACUGAA ........(((((........(((((((((....)))))-))))---((-(....))))))))(((((...(((...(((......)))...)))...))))). ( -25.90) >consensus GUGUUGAAGGACCAAAAGGAUGGGACUUUUAUAGGAGGCCC___UAUAUAUGGGGUAGGGUCUUUC______GGGGAGUCAGUGUCGACGCACUCAGAACUGAA (((((((....(((......)))(((((((...((((((((................))))))))........)))))))....)))))))..(((....))). (-22.14 = -21.89 + -0.25)


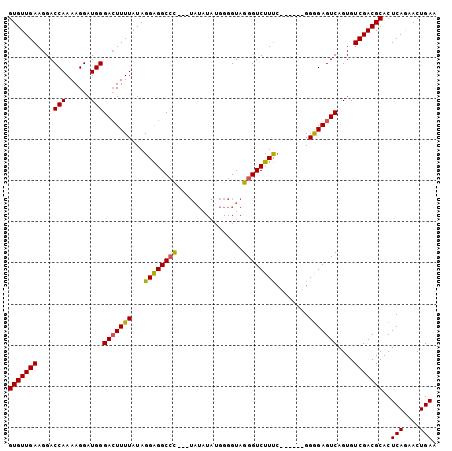
| Location | 7,470,829 – 7,470,932 |
|---|---|
| Length | 103 |
| Sequences | 4 |
| Columns | 116 |
| Reading direction | reverse |
| Mean pairwise identity | 86.45 |
| Mean single sequence MFE | -29.65 |
| Consensus MFE | -21.67 |
| Energy contribution | -21.80 |
| Covariance contribution | 0.13 |
| Combinations/Pair | 1.08 |
| Mean z-score | -1.54 |
| Structure conservation index | 0.73 |
| SVM decision value | 0.14 |
| SVM RNA-class probability | 0.604710 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 7470829 103 - 22224390 UGCAGAGGCAUGCCCGCAAGGUCUUUGGAAGGA-------------AGGAGGGAUUCCGAUUGUGAGGAGAAGGGUGUGUUGAAGGACCAAAAGGAUGGGACUUUUAUAGGAGGCC .((...(((((((((......((((.....(((-------------(.......))))......))))....)))))))))((((..(((......)))..))))........)). ( -26.80) >DroSec_CAF1 4088 103 - 1 UGCAGAGGCAUGCCCGCAAGGUCUUUAGAAGGC-------------AGGAGGGAUUCCGAUUGUGAGGAGAAGGGUGUGUUGAAGGACCAAAAGGAUGGGACUUUUAUAGGAGGCC .((...(((((((((......((((((.((...-------------.((((...))))..)).))))))...)))))))))((((..(((......)))..))))........)). ( -27.90) >DroSim_CAF1 5801 103 - 1 UGCAGAGGCAUGCCCGCAAGGUCUUUAGAAGGC-------------AGGAGGGAUUCCGAUUGUGAGGAGAAGGGUGUGUUGAAGGACCAAAAGGAUGGGACUUUUAUAGGAGGCC .((...(((((((((......((((((.((...-------------.((((...))))..)).))))))...)))))))))((((..(((......)))..))))........)). ( -27.90) >DroYak_CAF1 6562 116 - 1 UGCAGAGGCAUACCCGCAAGGUCCUUGGAAGGAAGAAGGGAUCGGAAGGAGGGAUUCCGAUUGCUAGGAGAUGGGUGUGUUGAAGGACCAAACGGAUGGGAGCUUCAAAAGGGGGG ......(((((((((((....((((....))))...((.((((((((.......)))))))).))....).))))))))))((((..(((......)))...)))).......... ( -36.00) >consensus UGCAGAGGCAUGCCCGCAAGGUCUUUAGAAGGA_____________AGGAGGGAUUCCGAUUGUGAGGAGAAGGGUGUGUUGAAGGACCAAAAGGAUGGGACUUUUAUAGGAGGCC ......(((((((((.....((((((.......................))))))(((........)))...)))))))))((((..(((......)))..))))........... (-21.67 = -21.80 + 0.13)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:43:09 2006