
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 7,421,471 – 7,421,582 |
| Length | 111 |
| Max. P | 0.996874 |

| Location | 7,421,471 – 7,421,565 |
|---|---|
| Length | 94 |
| Sequences | 5 |
| Columns | 95 |
| Reading direction | forward |
| Mean pairwise identity | 93.22 |
| Mean single sequence MFE | -34.60 |
| Consensus MFE | -29.82 |
| Energy contribution | -30.22 |
| Covariance contribution | 0.40 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.73 |
| Structure conservation index | 0.86 |
| SVM decision value | 1.50 |
| SVM RNA-class probability | 0.959245 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 7421471 94 + 22224390 CCUGAA-AAGGAGCAGGAUCAUCGUUCUGUGGAGCGCGUGAGCUCGCGAAUCAAGCGUUUGCCAAAGGGAAGACGCUAUCGCCAGCCCAUCGCCA (((...-.))).(((((((....))))))).(.(((.(((.(((.((((....(((((((.((....)).))))))).)))).))).))))))). ( -35.50) >DroSec_CAF1 170285 94 + 1 CCGAAA-AAGGAGCAGGAUCAUCGUUCUGUGGAGCGCGUGAGCUCGCGAAUCAAGCGUUUGCCAAAGGGAAGACGCUAUCGCCAGCCCAUCGCCA ((....-..)).(((((((....))))))).(.(((.(((.(((.((((....(((((((.((....)).))))))).)))).))).))))))). ( -32.90) >DroSim_CAF1 172220 94 + 1 CCGAAA-AAGGAGCAGGAUCAUCGUUCUGUGGAGCGCGUGAGCUCGCGAAUCAAGCGUUUGCCAAAGGGAAGACGCUAUCGCCAGCCCAUCGCCA ((....-..)).(((((((....))))))).(.(((.(((.(((.((((....(((((((.((....)).))))))).)))).))).))))))). ( -32.90) >DroEre_CAF1 184577 94 + 1 CCGAAA-AAGGAGCAGGAUCAUCGUUCUGUGGAGCGCGUGAGCUCGCGAAUCAAGCGUUCGCCAAGUGGCAGACGCUAUCGCCAGCCCAACGCCA ((....-..)).(((((((....)))))))...(.((((..(((.((((....(((((..(((....)))..))))).)))).)))...))))). ( -35.90) >DroYak_CAF1 210834 95 + 1 UCAACAGGAUCAGCAGGAUCAUCGUUCUGUGGAGCGCGUGAGCUCGCGAAUCAAGCGUUUGCCAACGUGCAGACGCUAUCGCCAGCCCAUCGCCA .........((.(((((((....))))))).))(((.(((.(((.((((....(((((((((......))))))))).)))).))).)))))).. ( -35.80) >consensus CCGAAA_AAGGAGCAGGAUCAUCGUUCUGUGGAGCGCGUGAGCUCGCGAAUCAAGCGUUUGCCAAAGGGAAGACGCUAUCGCCAGCCCAUCGCCA ............(((((((....)))))))...(.(((((.(((.((((....(((((((.((....)).))))))).)))).))).)).)))). (-29.82 = -30.22 + 0.40)

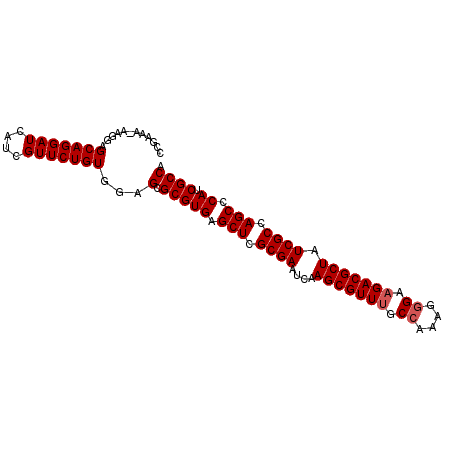

| Location | 7,421,487 – 7,421,582 |
|---|---|
| Length | 95 |
| Sequences | 5 |
| Columns | 95 |
| Reading direction | forward |
| Mean pairwise identity | 96.95 |
| Mean single sequence MFE | -37.68 |
| Consensus MFE | -35.02 |
| Energy contribution | -35.42 |
| Covariance contribution | 0.40 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.01 |
| Structure conservation index | 0.93 |
| SVM decision value | 2.76 |
| SVM RNA-class probability | 0.996874 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 7421487 95 + 22224390 AUCAUCGUUCUGUGGAGCGCGUGAGCUCGCGAAUCAAGCGUUUGCCAAAGGGAAGACGCUAUCGCCAGCCCAUCGCCACCAGCGAGCGGAUCAUG ....((((((..(((.(.(((((.(((.((((....(((((((.((....)).))))))).)))).))).)).)))).)))..))))))...... ( -36.40) >DroSec_CAF1 170301 95 + 1 AUCAUCGUUCUGUGGAGCGCGUGAGCUCGCGAAUCAAGCGUUUGCCAAAGGGAAGACGCUAUCGCCAGCCCAUCGCCACCAGCGAGCGGAUCAUG ....((((((..(((.(.(((((.(((.((((....(((((((.((....)).))))))).)))).))).)).)))).)))..))))))...... ( -36.40) >DroSim_CAF1 172236 95 + 1 AUCAUCGUUCUGUGGAGCGCGUGAGCUCGCGAAUCAAGCGUUUGCCAAAGGGAAGACGCUAUCGCCAGCCCAUCGCCACCAGCGAGCGGAUCAUG ....((((((..(((.(.(((((.(((.((((....(((((((.((....)).))))))).)))).))).)).)))).)))..))))))...... ( -36.40) >DroEre_CAF1 184593 95 + 1 AUCAUCGUUCUGUGGAGCGCGUGAGCUCGCGAAUCAAGCGUUCGCCAAGUGGCAGACGCUAUCGCCAGCCCAACGCCACCAGCGAGCGGAUCAUG ....((((((..(((.(.((((..(((.((((....(((((..(((....)))..))))).)))).)))...))))).)))..))))))...... ( -39.90) >DroYak_CAF1 210851 95 + 1 AUCAUCGUUCUGUGGAGCGCGUGAGCUCGCGAAUCAAGCGUUUGCCAACGUGCAGACGCUAUCGCCAGCCCAUCGCCACCAGCGAGCGGAUCAUG ....((((((..(((.(.(((((.(((.((((....(((((((((......))))))))).)))).))).)).)))).)))..))))))...... ( -39.30) >consensus AUCAUCGUUCUGUGGAGCGCGUGAGCUCGCGAAUCAAGCGUUUGCCAAAGGGAAGACGCUAUCGCCAGCCCAUCGCCACCAGCGAGCGGAUCAUG ....((((((..(((.(.(((((.(((.((((....(((((((.((....)).))))))).)))).))).)).)))).)))..))))))...... (-35.02 = -35.42 + 0.40)
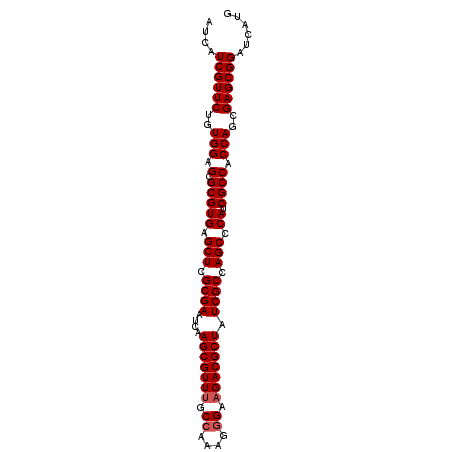



| Location | 7,421,487 – 7,421,582 |
|---|---|
| Length | 95 |
| Sequences | 5 |
| Columns | 95 |
| Reading direction | reverse |
| Mean pairwise identity | 96.95 |
| Mean single sequence MFE | -38.14 |
| Consensus MFE | -35.04 |
| Energy contribution | -35.08 |
| Covariance contribution | 0.04 |
| Combinations/Pair | 1.03 |
| Mean z-score | -1.66 |
| Structure conservation index | 0.92 |
| SVM decision value | 1.66 |
| SVM RNA-class probability | 0.970340 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 7421487 95 - 22224390 CAUGAUCCGCUCGCUGGUGGCGAUGGGCUGGCGAUAGCGUCUUCCCUUUGGCAAACGCUUGAUUCGCGAGCUCACGCGCUCCACAGAACGAUGAU .......((.((..(((.((((.((((((.((((.(((((...((....))...)))))....)))).))))))))).).)))..)).))..... ( -35.80) >DroSec_CAF1 170301 95 - 1 CAUGAUCCGCUCGCUGGUGGCGAUGGGCUGGCGAUAGCGUCUUCCCUUUGGCAAACGCUUGAUUCGCGAGCUCACGCGCUCCACAGAACGAUGAU .......((.((..(((.((((.((((((.((((.(((((...((....))...)))))....)))).))))))))).).)))..)).))..... ( -35.80) >DroSim_CAF1 172236 95 - 1 CAUGAUCCGCUCGCUGGUGGCGAUGGGCUGGCGAUAGCGUCUUCCCUUUGGCAAACGCUUGAUUCGCGAGCUCACGCGCUCCACAGAACGAUGAU .......((.((..(((.((((.((((((.((((.(((((...((....))...)))))....)))).))))))))).).)))..)).))..... ( -35.80) >DroEre_CAF1 184593 95 - 1 CAUGAUCCGCUCGCUGGUGGCGUUGGGCUGGCGAUAGCGUCUGCCACUUGGCGAACGCUUGAUUCGCGAGCUCACGCGCUCCACAGAACGAUGAU .......((.((..(((.(((((((((((.((((.(((((.((((....)))).)))))....)))).)))))).))))))))..)).))..... ( -43.50) >DroYak_CAF1 210851 95 - 1 CAUGAUCCGCUCGCUGGUGGCGAUGGGCUGGCGAUAGCGUCUGCACGUUGGCAAACGCUUGAUUCGCGAGCUCACGCGCUCCACAGAACGAUGAU .......((.((..(((.((((.((((((.((((.(((((.(((......))).)))))....)))).))))))))).).)))..)).))..... ( -39.80) >consensus CAUGAUCCGCUCGCUGGUGGCGAUGGGCUGGCGAUAGCGUCUUCCCUUUGGCAAACGCUUGAUUCGCGAGCUCACGCGCUCCACAGAACGAUGAU .......((.((..(((.((((.((((((.((((.(((((.(.((....)).).)))))....)))).))))))))).).)))..)).))..... (-35.04 = -35.08 + 0.04)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:42:23 2006