
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 7,181,041 – 7,181,149 |
| Length | 108 |
| Max. P | 0.751414 |

| Location | 7,181,041 – 7,181,149 |
|---|---|
| Length | 108 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 63.22 |
| Mean single sequence MFE | -28.93 |
| Consensus MFE | -12.73 |
| Energy contribution | -14.73 |
| Covariance contribution | 2.01 |
| Combinations/Pair | 1.17 |
| Mean z-score | -1.99 |
| Structure conservation index | 0.44 |
| SVM decision value | 0.48 |
| SVM RNA-class probability | 0.751414 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 7181041 108 - 22224390 GUACAAAAUAA-------AAAAAAUCGAAGGCCCC-----CCACUUUACCGUUUCGAAUGCGGCAAGCGGAGUGCACUGCACUCUGCGGAGGCAAAAACUUGAAAAUACAGACGGAAAAC ...........-------......(((...(((.(-----(.......((((.......))))...(((((((((...))))))))))).)))......(((......))).)))..... ( -27.50) >DroEre_CAF1 8225 115 - 1 GUACAAAAUAACAAAAAGAAAAAGUCGGAGGCCCC-----CCACUUUACCGUUUCGAAUGCGGCGAGCGGAGUGCACUGCACUCUGCGGAGGCAAAAACUCGAAAAUACAGACAGAAAAC .......................(((.((((((.(-----(....(..((((.......))))..)(((((((((...))))))))))).))).....)))(......).)))....... ( -29.70) >DroAna_CAF1 12749 94 - 1 GUGCAGCGUGG-------AG----CUCAAGCUCUCUCGCGCCAC--UACAGUUGU----------AACUGGGAGCACUGCACUCUGGGGAGGAAAACUUUUCAAA---CCAAAAGAAAAC ((((((((.((-------((----(....)))))..)))(((..--..((((...----------.)))).).))..)))))..((..(((.....)))..))..---............ ( -29.60) >consensus GUACAAAAUAA_______AAAAA_UCGAAGGCCCC_____CCACUUUACCGUUUCGAAUGCGGC_AGCGGAGUGCACUGCACUCUGCGGAGGCAAAAACUUGAAAAUACAGACAGAAAAC ..............................(((((.............((((.......))))...(((((((((...))))))))))).)))........................... (-12.73 = -14.73 + 2.01)

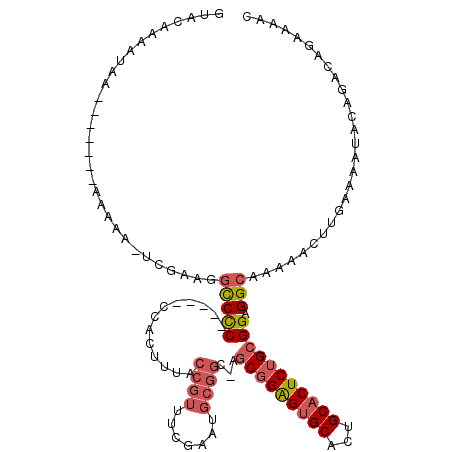

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:40:48 2006