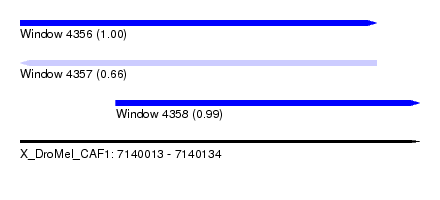

| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 7,140,013 – 7,140,134 |
| Length | 121 |
| Max. P | 0.997466 |

| Location | 7,140,013 – 7,140,121 |
|---|---|
| Length | 108 |
| Sequences | 5 |
| Columns | 109 |
| Reading direction | forward |
| Mean pairwise identity | 96.03 |
| Mean single sequence MFE | -33.58 |
| Consensus MFE | -30.24 |
| Energy contribution | -30.64 |
| Covariance contribution | 0.40 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.15 |
| Structure conservation index | 0.90 |
| SVM decision value | 2.86 |
| SVM RNA-class probability | 0.997466 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 7140013 108 + 22224390 AGUGGGCACAUUUUGUAGAGCUUGAACUUUACCUUUGAAAGGGUUGCUCUUGCUCC-CUUUUGAAACUCUGACCUUCAGAAGGAAAAGGAAACCCGGUGCUCAAGUCCC ..(((((((.....((((((......))))))........(((((..((((....(-((((((((.........)))))))))..)))).))))).)))))))...... ( -34.10) >DroSec_CAF1 7790 108 + 1 AGUGGGCACAUUUUGUCGAGCUUGAACUUUACCUUUCAAAGGGUUGCUCUUGCUCC-CUUUUGAAACUCUGACCUUCAGAAGGAAAAGGAAACCCGGUGCUCAAGUCCC ..(((((((......((((((..((.(...(((((....))))).).))..))))(-((((((((.........))))))))).....))......)))))))...... ( -32.90) >DroSim_CAF1 2157 108 + 1 AGUGGGCACAUUUUGUAGAGCUUGAACUUUACCUUUCAAAGGGUUGCUCUUGCUCC-CUUUUGAAACUCUGACCUUCAGAAGGAAAAGGAAACCCGGUGCUCAAGUCCC ..(((((((.....((((((......))))))........(((((..((((....(-((((((((.........)))))))))..)))).))))).)))))))...... ( -34.10) >DroEre_CAF1 8012 107 + 1 AGUGGGCACUU-UUGUAGAGCUUGAACUUUACCUUACAAAGGGUUGCUCUUGCUCC-CUUUUGAAACUCUGACCUUCAGAAGGAAAAGGAAACCCGGUGCUCAAGUCCC ..((((((((.-..((((((......))))))........(((((..((((....(-((((((((.........)))))))))..)))).)))))))))))))...... ( -34.40) >DroYak_CAF1 7585 109 + 1 AGUGGGCACUCGUUGUAGAGCUUGAACUUAACCAUUGAAAGGGUUGCUCUUGCUCCCCUUUUGAAACUCUGACCUUCAGAAGGAAAAGGAAACCCGGUGCUCAAGUCCC ..(((((((.((.....((((..((...(((((........))))).))..)))).(((((((((.........)))))))))....(....).)))))))))...... ( -32.40) >consensus AGUGGGCACAUUUUGUAGAGCUUGAACUUUACCUUUCAAAGGGUUGCUCUUGCUCC_CUUUUGAAACUCUGACCUUCAGAAGGAAAAGGAAACCCGGUGCUCAAGUCCC ..(((((((.....((((((......))))))........(((((..((((..(((..(((((((.........)))))))))).)))).))))).)))))))...... (-30.24 = -30.64 + 0.40)



| Location | 7,140,013 – 7,140,121 |
|---|---|
| Length | 108 |
| Sequences | 5 |
| Columns | 109 |
| Reading direction | reverse |
| Mean pairwise identity | 96.03 |
| Mean single sequence MFE | -31.82 |
| Consensus MFE | -26.77 |
| Energy contribution | -26.97 |
| Covariance contribution | 0.20 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.47 |
| Structure conservation index | 0.84 |
| SVM decision value | 0.27 |
| SVM RNA-class probability | 0.663293 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 7140013 108 - 22224390 GGGACUUGAGCACCGGGUUUCCUUUUCCUUCUGAAGGUCAGAGUUUCAAAAG-GGAGCAAGAGCAACCCUUUCAAAGGUAAAGUUCAAGCUCUACAAAAUGUGCCCACU (((....((((.....((((((((((..(((((.....)))))....)))))-)))))..((((.(((........)))...))))..))))(((.....))))))... ( -30.80) >DroSec_CAF1 7790 108 - 1 GGGACUUGAGCACCGGGUUUCCUUUUCCUUCUGAAGGUCAGAGUUUCAAAAG-GGAGCAAGAGCAACCCUUUGAAAGGUAAAGUUCAAGCUCGACAAAAUGUGCCCACU (((((((((((.....((((((((((..(((((.....)))))....)))))-)))))..((((.(((........)))...))))..))))))......)).)))... ( -32.00) >DroSim_CAF1 2157 108 - 1 GGGACUUGAGCACCGGGUUUCCUUUUCCUUCUGAAGGUCAGAGUUUCAAAAG-GGAGCAAGAGCAACCCUUUGAAAGGUAAAGUUCAAGCUCUACAAAAUGUGCCCACU (((.(((((((((((((........)))(((((.....)))))(((((((.(-(..((....))..)).))))))))))...)))))))...(((.....))))))... ( -30.80) >DroEre_CAF1 8012 107 - 1 GGGACUUGAGCACCGGGUUUCCUUUUCCUUCUGAAGGUCAGAGUUUCAAAAG-GGAGCAAGAGCAACCCUUUGUAAGGUAAAGUUCAAGCUCUACAA-AAGUGCCCACU (((((((((((.....((((((((((..(((((.....)))))....)))))-)))))..((((.(((........)))...))))..)))).....-)))).)))... ( -32.70) >DroYak_CAF1 7585 109 - 1 GGGACUUGAGCACCGGGUUUCCUUUUCCUUCUGAAGGUCAGAGUUUCAAAAGGGGAGCAAGAGCAACCCUUUCAAUGGUUAAGUUCAAGCUCUACAACGAGUGCCCACU ((((((((............((((((..(((((.....)))))....))))))(((((..((((((((........))))..))))..)))))....))))).)))... ( -32.80) >consensus GGGACUUGAGCACCGGGUUUCCUUUUCCUUCUGAAGGUCAGAGUUUCAAAAG_GGAGCAAGAGCAACCCUUUGAAAGGUAAAGUUCAAGCUCUACAAAAUGUGCCCACU (((.(((((((((((((((..(((((...((((.....))))(((((......)))))))))).))))).......)))...)))))))...(((.....))))))... (-26.77 = -26.97 + 0.20)


| Location | 7,140,042 – 7,140,134 |
|---|---|
| Length | 92 |
| Sequences | 5 |
| Columns | 93 |
| Reading direction | forward |
| Mean pairwise identity | 97.62 |
| Mean single sequence MFE | -32.56 |
| Consensus MFE | -26.60 |
| Energy contribution | -26.60 |
| Covariance contribution | -0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -3.63 |
| Structure conservation index | 0.82 |
| SVM decision value | 2.14 |
| SVM RNA-class probability | 0.988853 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 7140042 92 + 22224390 UACCUUUGAAAGGGUUGCUCUUGCUCC-CUUUUGAAACUCUGACCUUCAGAAGGAAAAGGAAACCCGGUGCUCAAGUCCCGGAGUGCCUUUAA ..((((..((((((..((....)).))-))))..))..((((.....)))).)).(((((..((((((..(....)..)))).)).))))).. ( -32.40) >DroSec_CAF1 7819 92 + 1 UACCUUUCAAAGGGUUGCUCUUGCUCC-CUUUUGAAACUCUGACCUUCAGAAGGAAAAGGAAACCCGGUGCUCAAGUCCCGGAGUGCCUUUAA ..((((((((((((..((....))..)-))))))))..((((.....))))))).(((((..((((((..(....)..)))).)).))))).. ( -33.10) >DroSim_CAF1 2186 92 + 1 UACCUUUCAAAGGGUUGCUCUUGCUCC-CUUUUGAAACUCUGACCUUCAGAAGGAAAAGGAAACCCGGUGCUCAAGUCCCGGAGUGCCUUUAA ..((((((((((((..((....))..)-))))))))..((((.....))))))).(((((..((((((..(....)..)))).)).))))).. ( -33.10) >DroEre_CAF1 8040 92 + 1 UACCUUACAAAGGGUUGCUCUUGCUCC-CUUUUGAAACUCUGACCUUCAGAAGGAAAAGGAAACCCGGUGCUCAAGUCCCGGAGUGCCUUUAA ..((((.(((((((..((....))..)-)))))).....(((.....))))))).(((((..((((((..(....)..)))).)).))))).. ( -30.90) >DroYak_CAF1 7614 93 + 1 AACCAUUGAAAGGGUUGCUCUUGCUCCCCUUUUGAAACUCUGACCUUCAGAAGGAAAAGGAAACCCGGUGCUCAAGUCCCGGAGUGCCUUUAA ..((.(..((((((..((....))..))))))..)...((((.....)))).)).(((((..((((((..(....)..)))).)).))))).. ( -33.30) >consensus UACCUUUCAAAGGGUUGCUCUUGCUCC_CUUUUGAAACUCUGACCUUCAGAAGGAAAAGGAAACCCGGUGCUCAAGUCCCGGAGUGCCUUUAA ...........(((((..((((..(((..(((((((.........)))))))))).)))).)))))((..(((........)))..))..... (-26.60 = -26.60 + -0.00)
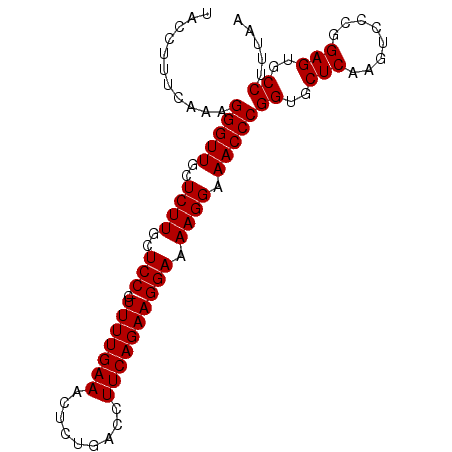


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:40:20 2006