
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 900,111 – 900,204 |
| Length | 93 |
| Max. P | 0.707833 |

| Location | 900,111 – 900,204 |
|---|---|
| Length | 93 |
| Sequences | 6 |
| Columns | 93 |
| Reading direction | forward |
| Mean pairwise identity | 82.76 |
| Mean single sequence MFE | -33.39 |
| Consensus MFE | -22.26 |
| Energy contribution | -24.57 |
| Covariance contribution | 2.31 |
| Combinations/Pair | 1.17 |
| Mean z-score | -2.02 |
| Structure conservation index | 0.67 |
| SVM decision value | 0.23 |
| SVM RNA-class probability | 0.643138 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 900111 93 + 22224390 CGGCAGCCUCCACUUCCAUCGGUGGCUCCUCAACGGCGACCUCCCAGCUCUCGCCGAACUCCGAGGAGAGCGAGCUGCCUUUCGUCCUUCCCU .(((((((.(((((......)))))((((((..((((((...........))))))......))))))...).)))))).............. ( -33.90) >DroSec_CAF1 15126 93 + 1 CGGCAGCCUCCACUUCCCUCGGUGGCUCCUCAACGGCGACCUCCCAGCUCUCGCCGAACUCCGAGGAGAGCGAGCUGCCCUUCGUCCUGCCUU .((((((.((..((..(((((((((....))).((((((...........))))))....))))))..)).)))))))).............. ( -34.40) >DroSim_CAF1 15818 93 + 1 CGGCAGCCUCCACUUCCCUCGGUGGCUCCUCAACGGCGACCUCCCAGCUCUCGCCGAACUCCGAGGAGAGCGAGCUGCCCUUCCUCCUGCCCU .((((((.((..((..(((((((((....))).((((((...........))))))....))))))..)).)))))))).............. ( -34.40) >DroEre_CAF1 9079 93 + 1 CGGCAGCCUCCACUUCCCUCAGUGGCUCCUCGACGGCGCCCUCCCAGCUCUCGCCGGAUUCCGAGGAGAGCGAGCUGCCCUUCGUCCUGCCCU .(((((((.(((((......)))))(((((((.(((((.............))))).....)))))))...).)))))).............. ( -35.02) >DroYak_CAF1 8912 84 + 1 CGGCAGCCUCCUCCAC---------UUCCUCCACGGCGCCCUCCCAGCUCUCGCCGAACUCCGAGGAGAGCGAGCUGCCCUUCGUCCUGCCCU .((((((.(((((...---------.(((((..(((((.............)))))......)))))))).)))))))).............. ( -27.12) >DroAna_CAF1 11031 90 + 1 CUGGAGCAACGGGAUC---AGCUGGGUCCUCCACCACGACGGCCCAGCUCUCGCCCGAUUCGGAGGAGAGCGAACAGCCCUUUGUCCUGCCAC ...(.(((.((((...---(((((((((.((......)).)))))))))....))))....(((((((.((.....)).)))).))))))).. ( -35.50) >consensus CGGCAGCCUCCACUUCCCUCGGUGGCUCCUCAACGGCGACCUCCCAGCUCUCGCCGAACUCCGAGGAGAGCGAGCUGCCCUUCGUCCUGCCCU .((((((..(((((......)))))((((((..((((((...........))))))......)))))).....)))))).............. (-22.26 = -24.57 + 2.31)

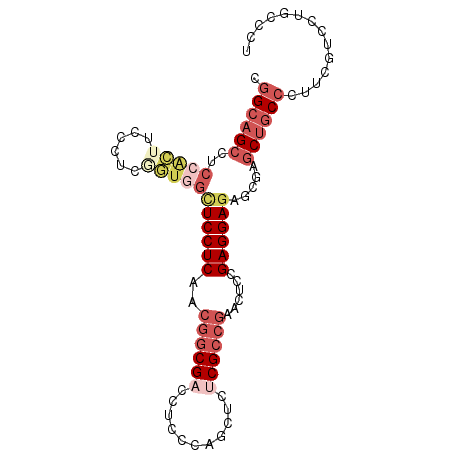
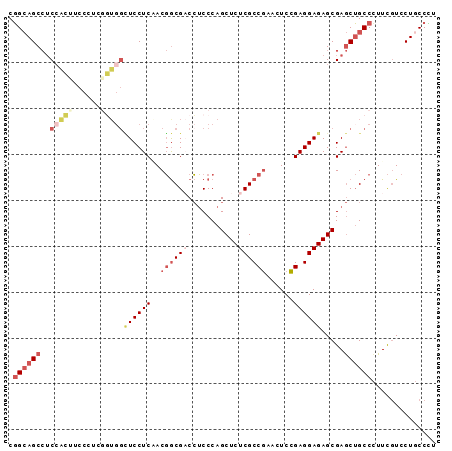
| Location | 900,111 – 900,204 |
|---|---|
| Length | 93 |
| Sequences | 6 |
| Columns | 93 |
| Reading direction | reverse |
| Mean pairwise identity | 82.76 |
| Mean single sequence MFE | -42.02 |
| Consensus MFE | -28.15 |
| Energy contribution | -29.18 |
| Covariance contribution | 1.03 |
| Combinations/Pair | 1.26 |
| Mean z-score | -2.22 |
| Structure conservation index | 0.67 |
| SVM decision value | 0.37 |
| SVM RNA-class probability | 0.707833 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
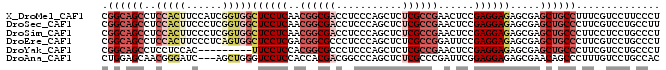
>X_DroMel_CAF1 900111 93 - 22224390 AGGGAAGGACGAAAGGCAGCUCGCUCUCCUCGGAGUUCGGCGAGAGCUGGGAGGUCGCCGUUGAGGAGCCACCGAUGGAAGUGGAGGCUGCCG ..............(((((((((((.(((((((..((((((....)))))).(((..((.....)).))).)))).)))))))..))))))). ( -40.40) >DroSec_CAF1 15126 93 - 1 AAGGCAGGACGAAGGGCAGCUCGCUCUCCUCGGAGUUCGGCGAGAGCUGGGAGGUCGCCGUUGAGGAGCCACCGAGGGAAGUGGAGGCUGCCG ..............(((((((((((..((((((..((((((....)))))).(((..((.....)).))).))))))..))))..))))))). ( -45.60) >DroSim_CAF1 15818 93 - 1 AGGGCAGGAGGAAGGGCAGCUCGCUCUCCUCGGAGUUCGGCGAGAGCUGGGAGGUCGCCGUUGAGGAGCCACCGAGGGAAGUGGAGGCUGCCG ..............(((((((((((..((((((..((((((....)))))).(((..((.....)).))).))))))..))))..))))))). ( -45.60) >DroEre_CAF1 9079 93 - 1 AGGGCAGGACGAAGGGCAGCUCGCUCUCCUCGGAAUCCGGCGAGAGCUGGGAGGGCGCCGUCGAGGAGCCACUGAGGGAAGUGGAGGCUGCCG ..............(((((((((((..((((((..((((((....))))))..(((.((.....)).))).))))))..))))..))))))). ( -49.50) >DroYak_CAF1 8912 84 - 1 AGGGCAGGACGAAGGGCAGCUCGCUCUCCUCGGAGUUCGGCGAGAGCUGGGAGGGCGCCGUGGAGGAA---------GUGGAGGAGGCUGCCG ..............(((((((((((((((.(((.(((((((....)))))....)).))).))))..)---------))).....))))))). ( -35.10) >DroAna_CAF1 11031 90 - 1 GUGGCAGGACAAAGGGCUGUUCGCUCUCCUCCGAAUCGGGCGAGAGCUGGGCCGUCGUGGUGGAGGACCCAGCU---GAUCCCGUUGCUCCAG (..((((((...(((((.....)))))..)))....((((....(((((((((.((......)))).)))))))---...)))).)))..).. ( -35.90) >consensus AGGGCAGGACGAAGGGCAGCUCGCUCUCCUCGGAGUUCGGCGAGAGCUGGGAGGUCGCCGUUGAGGAGCCACCGAGGGAAGUGGAGGCUGCCG ..............(((((((....((((((((..((((((....)))))).((...)).))))))))((((........)))).))))))). (-28.15 = -29.18 + 1.03)

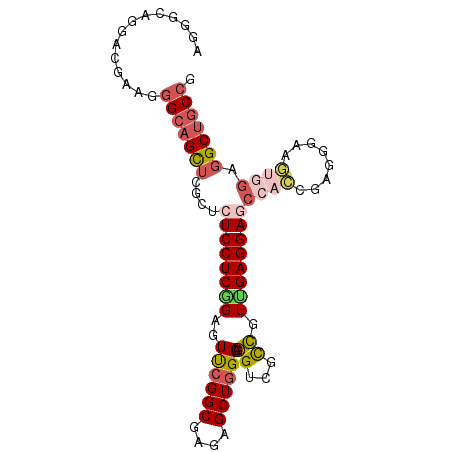
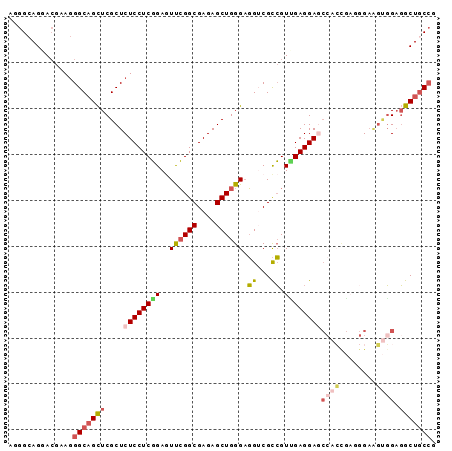
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 09:40:25 2006