
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 7,007,875 – 7,008,261 |
| Length | 386 |
| Max. P | 0.998335 |

| Location | 7,007,875 – 7,007,995 |
|---|---|
| Length | 120 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 83.54 |
| Mean single sequence MFE | -37.05 |
| Consensus MFE | -30.83 |
| Energy contribution | -31.70 |
| Covariance contribution | 0.87 |
| Combinations/Pair | 1.19 |
| Mean z-score | -2.79 |
| Structure conservation index | 0.83 |
| SVM decision value | 2.45 |
| SVM RNA-class probability | 0.994046 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 7007875 120 + 22224390 UAUAUUCCGAGUUUUAAAAAGUUAGAAUACUUCCUUUUCAAUUGACCACCAAACGCCACCUGUUGGCCGGGGCAGUUUCAGCCGUUGGCCCAUAGAUGGCGACAGUAUGCUUAUUGGACA .....((((((((...(((((..((....))..))))).....))).......(((((.((((.(((((((((.......))).)))))).)))).)))))............))))).. ( -35.10) >DroSec_CAF1 9081 119 + 1 -AUAUUCCGAGUUUUAAGAAGUUAGAACACUUCCUUAUCAAAUGACCACCAAACGCCACCUGUUGGCCGGGGCAGUUUCAGCCGUUGGCCCACAGAUGGCGACAGUAUGCUUAUUGGACA -....((((((((((((....))))))).........................(((((.((((.(((((((((.......))).)))))).)))).)))))............))))).. ( -37.80) >DroEre_CAF1 14938 104 + 1 -AUAUUCCGAGUUUUGAAGCCUUAA---------------AAUGACCACCAAACGCCACCUGUUGGCCAGGCCAGUUUCAGCCGUUGGCCAACAGAUGGCGACAGUAUGCUUAUUGGACA -....((((((((((((....))))---------------)))..........(((((.(((((((((((((........)))..)))))))))).)))))............))))).. ( -38.40) >DroYak_CAF1 23419 119 + 1 -AUAUUCAACAUUUUGAAGACUUUAAAUACAUUCUUAGCAAAUGACCACCAAACGCCACCUGUUGGCCGGGCCUGUUUCAGCCGUUGGCCAACAGAUGGCGACAGCAUGCUUAUUGGACA -........(((((..((((............))))...)))))....((((.(((((.(((((((((((((........)))..)))))))))).)))))..((....))..))))... ( -36.90) >consensus _AUAUUCCGAGUUUUAAAAACUUAAAAUACUUCCUUAUCAAAUGACCACCAAACGCCACCUGUUGGCCGGGCCAGUUUCAGCCGUUGGCCAACAGAUGGCGACAGUAUGCUUAUUGGACA ..........(((((((....)))))))....................((((.(((((.((((((((((((((.......))).))))))))))).)))))..((....))..))))... (-30.83 = -31.70 + 0.87)


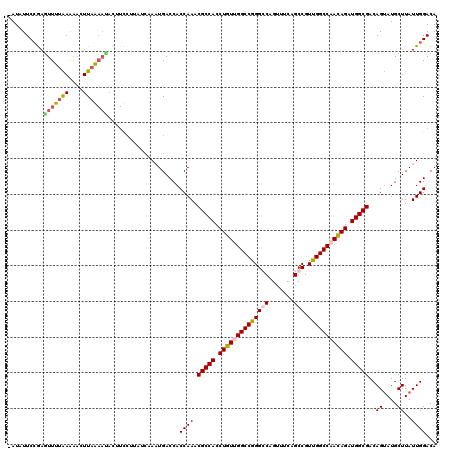
| Location | 7,007,875 – 7,007,995 |
|---|---|
| Length | 120 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 83.54 |
| Mean single sequence MFE | -42.95 |
| Consensus MFE | -32.90 |
| Energy contribution | -34.15 |
| Covariance contribution | 1.25 |
| Combinations/Pair | 1.00 |
| Mean z-score | -3.72 |
| Structure conservation index | 0.77 |
| SVM decision value | 3.07 |
| SVM RNA-class probability | 0.998335 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 7007875 120 - 22224390 UGUCCAAUAAGCAUACUGUCGCCAUCUAUGGGCCAACGGCUGAAACUGCCCCGGCCAACAGGUGGCGUUUGGUGGUCAAUUGAAAAGGAAGUAUUCUAACUUUUUAAAACUCGGAAUAUA ..(((.....((.(((((.((((((((...((((...(((.......)))..))))...))))))))..)))))))...((((((((..((....))..)))))))).....)))..... ( -38.90) >DroSec_CAF1 9081 119 - 1 UGUCCAAUAAGCAUACUGUCGCCAUCUGUGGGCCAACGGCUGAAACUGCCCCGGCCAACAGGUGGCGUUUGGUGGUCAUUUGAUAAGGAAGUGUUCUAACUUCUUAAAACUCGGAAUAU- ..(((........(((((.((((((((((.((((...(((.......)))..)))).))))))))))..)))))(((....))).(((((((......))))))).......)))....- ( -44.10) >DroEre_CAF1 14938 104 - 1 UGUCCAAUAAGCAUACUGUCGCCAUCUGUUGGCCAACGGCUGAAACUGGCCUGGCCAACAGGUGGCGUUUGGUGGUCAUU---------------UUAAGGCUUCAAAACUCGGAAUAU- ..(((...............(((((((((((((((..((((......)))))))))))))))))))((((...((((...---------------....))))...))))..)))....- ( -45.20) >DroYak_CAF1 23419 119 - 1 UGUCCAAUAAGCAUGCUGUCGCCAUCUGUUGGCCAACGGCUGAAACAGGCCCGGCCAACAGGUGGCGUUUGGUGGUCAUUUGCUAAGAAUGUAUUUAAAGUCUUCAAAAUGUUGAAUAU- .........(((((..((.(((((((((((((((...((((......)))).)))))))))))))))(((((..(....)..))))).................))..)))))......- ( -43.60) >consensus UGUCCAAUAAGCAUACUGUCGCCAUCUGUGGGCCAACGGCUGAAACUGCCCCGGCCAACAGGUGGCGUUUGGUGGUCAUUUGAUAAGGAAGUAUUCUAACUCUUCAAAACUCGGAAUAU_ ((.(((.(((((........((((((((((((((...((((......)))).))))))))))))))))))).))).)).......................................... (-32.90 = -34.15 + 1.25)


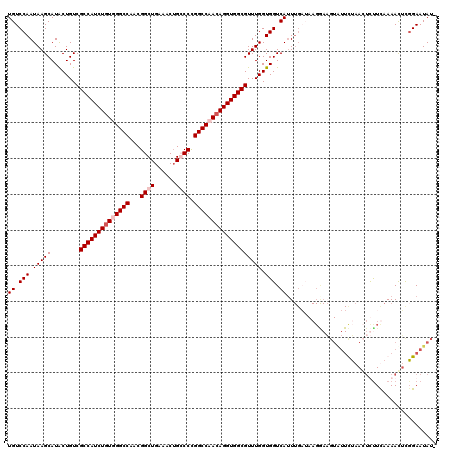
| Location | 7,007,915 – 7,008,035 |
|---|---|
| Length | 120 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 96.39 |
| Mean single sequence MFE | -42.97 |
| Consensus MFE | -40.47 |
| Energy contribution | -41.10 |
| Covariance contribution | 0.63 |
| Combinations/Pair | 1.05 |
| Mean z-score | -2.28 |
| Structure conservation index | 0.94 |
| SVM decision value | 1.42 |
| SVM RNA-class probability | 0.952404 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 7007915 120 + 22224390 AUUGACCACCAAACGCCACCUGUUGGCCGGGGCAGUUUCAGCCGUUGGCCCAUAGAUGGCGACAGUAUGCUUAUUGGACACGCGCAAACGUAAAUGCCAAUUGGAAGCCUCGAAUUGGUU .......(((((.(((((.((((.(((((((((.......))).)))))).)))).))))).......(((((((((.((.(((....)))...)))))))...))))......))))). ( -40.40) >DroSec_CAF1 9120 120 + 1 AAUGACCACCAAACGCCACCUGUUGGCCGGGGCAGUUUCAGCCGUUGGCCCACAGAUGGCGACAGUAUGCUUAUUGGACACGCGCAAACGUAAAUGCCAAUUGGAAGCCUCGAAUUGGUU .......(((((.(((((.((((.(((((((((.......))).)))))).)))).))))).......(((((((((.((.(((....)))...)))))))...))))......))))). ( -42.30) >DroEre_CAF1 14962 120 + 1 AAUGACCACCAAACGCCACCUGUUGGCCAGGCCAGUUUCAGCCGUUGGCCAACAGAUGGCGACAGUAUGCUUAUUGGACACGCGCAAACGUAAAUGCCAAUUGGAAGCCUCGAACUGGUU ...(((((.....(((((.(((((((((((((........)))..)))))))))).))))).......(((((((((.((.(((....)))...)))))))...)))).......))))) ( -44.50) >DroYak_CAF1 23458 120 + 1 AAUGACCACCAAACGCCACCUGUUGGCCGGGCCUGUUUCAGCCGUUGGCCAACAGAUGGCGACAGCAUGCUUAUUGGACACGCGCAAACGUAAAUGCCAAUUGGAAGCCUCGAAUUGGUU .......(((((.(((((.(((((((((((((........)))..)))))))))).)))))...((...((.(((((.((.(((....)))...))))))).))..))......))))). ( -44.70) >consensus AAUGACCACCAAACGCCACCUGUUGGCCGGGCCAGUUUCAGCCGUUGGCCAACAGAUGGCGACAGUAUGCUUAUUGGACACGCGCAAACGUAAAUGCCAAUUGGAAGCCUCGAAUUGGUU ...(((((.....(((((.((((((((((((((.......))).))))))))))).))))).......(((((((((.((.(((....)))...)))))))...)))).......))))) (-40.47 = -41.10 + 0.63)



| Location | 7,007,915 – 7,008,035 |
|---|---|
| Length | 120 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 96.39 |
| Mean single sequence MFE | -48.30 |
| Consensus MFE | -44.46 |
| Energy contribution | -45.52 |
| Covariance contribution | 1.06 |
| Combinations/Pair | 1.03 |
| Mean z-score | -3.19 |
| Structure conservation index | 0.92 |
| SVM decision value | 1.79 |
| SVM RNA-class probability | 0.977509 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 7007915 120 - 22224390 AACCAAUUCGAGGCUUCCAAUUGGCAUUUACGUUUGCGCGUGUCCAAUAAGCAUACUGUCGCCAUCUAUGGGCCAACGGCUGAAACUGCCCCGGCCAACAGGUGGCGUUUGGUGGUCAAU ...........((((.((((((((....(((((....))))).)))))...........((((((((...((((...(((.......)))..))))...))))))))..))).))))... ( -41.20) >DroSec_CAF1 9120 120 - 1 AACCAAUUCGAGGCUUCCAAUUGGCAUUUACGUUUGCGCGUGUCCAAUAAGCAUACUGUCGCCAUCUGUGGGCCAACGGCUGAAACUGCCCCGGCCAACAGGUGGCGUUUGGUGGUCAUU ...........((((.((((((((....(((((....))))).)))))...........((((((((((.((((...(((.......)))..)))).))))))))))..))).))))... ( -46.00) >DroEre_CAF1 14962 120 - 1 AACCAGUUCGAGGCUUCCAAUUGGCAUUUACGUUUGCGCGUGUCCAAUAAGCAUACUGUCGCCAUCUGUUGGCCAACGGCUGAAACUGGCCUGGCCAACAGGUGGCGUUUGGUGGUCAUU ...........((((.((((((((....(((((....))))).)))))...........((((((((((((((((..((((......))))))))))))))))))))..))).))))... ( -52.50) >DroYak_CAF1 23458 120 - 1 AACCAAUUCGAGGCUUCCAAUUGGCAUUUACGUUUGCGCGUGUCCAAUAAGCAUGCUGUCGCCAUCUGUUGGCCAACGGCUGAAACAGGCCCGGCCAACAGGUGGCGUUUGGUGGUCAUU ...........((((.((((...(((........)))((((((.......))))))...(((((((((((((((...((((......)))).))))))))))))))).)))).))))... ( -53.50) >consensus AACCAAUUCGAGGCUUCCAAUUGGCAUUUACGUUUGCGCGUGUCCAAUAAGCAUACUGUCGCCAUCUGUGGGCCAACGGCUGAAACUGCCCCGGCCAACAGGUGGCGUUUGGUGGUCAUU .(((((......((((...(((((....(((((....))))).))))))))).......(((((((((((((((...((((......)))).))))))))))))))).)))))....... (-44.46 = -45.52 + 1.06)



| Location | 7,008,035 – 7,008,150 |
|---|---|
| Length | 115 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 91.21 |
| Mean single sequence MFE | -37.38 |
| Consensus MFE | -31.60 |
| Energy contribution | -32.10 |
| Covariance contribution | 0.50 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.23 |
| Structure conservation index | 0.85 |
| SVM decision value | 1.34 |
| SVM RNA-class probability | 0.943555 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 7008035 115 - 22224390 CCACUAUGGUGUAUGUAUAUCCAGAUUUGCAUAUGUGUA----AGCCCCGUAUGUGUGUGAGUGUGCCACUCACAUGUGUGCACUGGUGUGCGUUUGUCCAUAAAACCACCCCA-AUGGC ....(((((...(((((((.((((.((((((....))))----))....(((((..((((((((...))))))))..))))).)))))))))))....)))))...(((.....-.))). ( -39.40) >DroSec_CAF1 9240 115 - 1 CCAGUAUGGUGUAUGUAUAUCCAAAUUUGCAUAUGUGUA----AGCCCCGUAUGUGUGUGAGUGUGCCACUCACAUGUGUGCACUGGUGUGCGUUUGUCCAUAAAACCACCCCA-AUGGC (((((..(((.((..(((((.((....)).)))))..))----.)))..(((((..((((((((...))))))))..)))))))))).(((.((((.......)))))))....-..... ( -37.00) >DroEre_CAF1 15082 114 - 1 CUAGU--AUGGUAUGUAUAUCCAGAUUUGCAUAUGUGUA----AGCCUAGUAUGUGUGUGAGUGUGCCACUCACAUGUGUGAACUGGUGUGCGUUUGUCCAUAAAACCACCCCAAAUGGC ....(--((((.(((((((.((((.((((((....))))----)).....((((..((((((((...))))))))..))))..)))))))))))....)))))...(((.......))). ( -34.40) >DroYak_CAF1 23578 119 - 1 CCAGUAUAUGGUAUGUAUAUCCUGAUUUGCAUAUGUGUACAUAUGCCCUGUAUGUGUGUGAGUGUGCCACUCACAUGUGUGCACUGGUGUGCGAUUGUCCAUAAAACCACCCCA-AUGGC ......(((((..((((((((.......(((((((....)))))))..((((((..((((((((...))))))))..))))))..)))))))).....)))))...(((.....-.))). ( -38.70) >consensus CCAGUAUAGGGUAUGUAUAUCCAGAUUUGCAUAUGUGUA____AGCCCCGUAUGUGUGUGAGUGUGCCACUCACAUGUGUGCACUGGUGUGCGUUUGUCCAUAAAACCACCCCA_AUGGC ..........(((((((.((....)).)))))))..........(((..(((((..((((((((...))))))))..)))))..(((.(((.((((.......))))))).)))...))) (-31.60 = -32.10 + 0.50)



| Location | 7,008,150 – 7,008,261 |
|---|---|
| Length | 111 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 89.71 |
| Mean single sequence MFE | -19.60 |
| Consensus MFE | -16.86 |
| Energy contribution | -16.42 |
| Covariance contribution | -0.44 |
| Combinations/Pair | 1.10 |
| Mean z-score | -1.83 |
| Structure conservation index | 0.86 |
| SVM decision value | 0.17 |
| SVM RNA-class probability | 0.615031 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 7008150 111 - 22224390 UCU---GUGACAAUUGACCCCUUGGAUCCUAUCAAAUUUCCACUUUUCCGAACGACUUACAAGGAAAUCGGGUUCCUUGACCAAUUCUCGACCCUUUUUCCU------UGCAUUUUCCCU ...---...............(((((....................)))))........((((((((..(((((....((.....))..)))))..))))))------)).......... ( -19.25) >DroSec_CAF1 9355 110 - 1 CCU---GUGACAAUUGACCCCUUGGAUCCUAUCAAAUGUCCACUUUUCCGAACGACUUCCAAGGAAAUCGGGUUCCUUGACCAAUUCUCGACCCUU-UUCCU------UGCAUUUUCCCU ..(---(.((((.((((((....))......)))).)))))).................((((((((..(((((....((.....))..))))).)-)))))------)).......... ( -20.60) >DroYak_CAF1 23697 119 - 1 UCUGCUGCGACAAUUGACCCCUCGGAUCAUAUCAAAUUUCCACUUUUCCGAACAACUUCCAAGGAAAUCGGGUUCCUUGACCAAUUCUCGACCCUU-UUCUUUUUUCCUGCAUUUUCCCU .......(((.(((((.....(((((....................)))))........(((((((......)))))))..))))).)))......-....................... ( -18.95) >consensus UCU___GUGACAAUUGACCCCUUGGAUCCUAUCAAAUUUCCACUUUUCCGAACGACUUCCAAGGAAAUCGGGUUCCUUGACCAAUUCUCGACCCUU_UUCCU______UGCAUUUUCCCU .......(((.(((((.....(((((....................)))))........(((((((......)))))))..))))).))).............................. (-16.86 = -16.42 + -0.44)

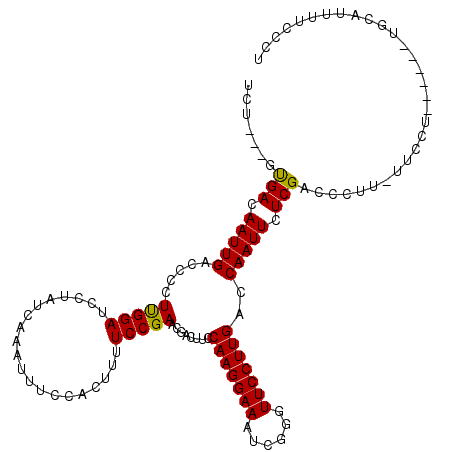

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:39:38 2006