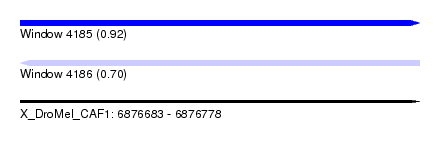

| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 6,876,683 – 6,876,778 |
| Length | 95 |
| Max. P | 0.920166 |

| Location | 6,876,683 – 6,876,778 |
|---|---|
| Length | 95 |
| Sequences | 4 |
| Columns | 98 |
| Reading direction | forward |
| Mean pairwise identity | 81.17 |
| Mean single sequence MFE | -25.65 |
| Consensus MFE | -17.99 |
| Energy contribution | -19.30 |
| Covariance contribution | 1.31 |
| Combinations/Pair | 1.14 |
| Mean z-score | -2.11 |
| Structure conservation index | 0.70 |
| SVM decision value | 1.13 |
| SVM RNA-class probability | 0.920166 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
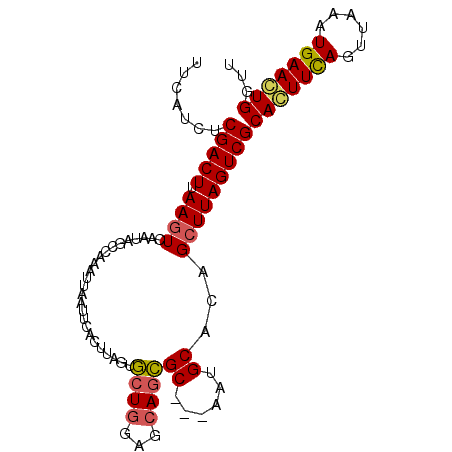
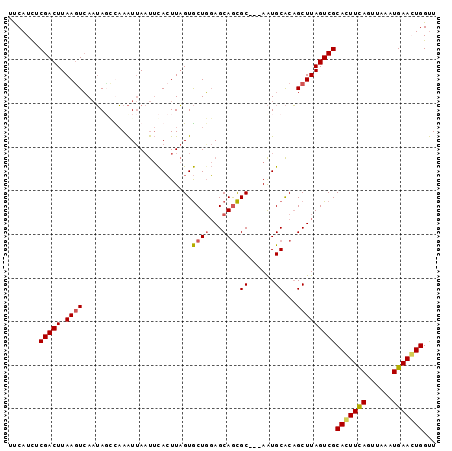
>X_DroMel_CAF1 6876683 95 + 22224390 AACCAGUUCAUUUAACUGAAGUGCGACUAAGCUGUGCAUU---GCGCUGCUCGAGCACUAAGUGAAUUAAUUUGCCUAUUCAGUUAAGUCGAAAUGAA ...((.((((((((((((((((((.....(((.((((...---)))).)))...)))))..(..((....))..)....)))))))))).))).)).. ( -27.80) >DroSec_CAF1 1185 95 + 1 AACCACUUCAUUUAACUGAAGUGCGACUAAGCUGUGCAUU---GCGCUGCUCCAGCACUAAGUGAAUUAAUUUGGAUAUUGACUUAAGUCGAGAUGAA ...(((((((......)))))))(((((..((((.(((..---....)))..)))).((((((......))))))...........)))))....... ( -25.50) >DroSim_CAF1 1215 95 + 1 AACCACUUCAUUUAACUGAAGUGCGACUAAGCUGUGCAUU---GCGCUGCUCCAGCACUAAGUGAAUUAAUUUAGCUAUUGACUUAAGUCGAGAUGAA ...(((((((......)))))))(((((..((((.(((..---....)))..))))..(((((.(((..........))).))))))))))....... ( -25.50) >DroYak_CAF1 1347 96 + 1 AGCCAGUUCAUUUAACUAAACUGCGACUAAGCGGCGCACUCGAGCAUUGCUACAGUACAAAUUA-G-UGACUAAUGUCGUGACUUAAGUCGAAAUGGC .((((.(((.............(((((..(((((.((......)).)))))..(((((......-)-).)))...)))))(((....)))))).)))) ( -23.80) >consensus AACCACUUCAUUUAACUGAAGUGCGACUAAGCUGUGCAUU___GCGCUGCUCCAGCACUAAGUGAAUUAAUUUAGCUAUUGACUUAAGUCGAAAUGAA ...(((((((......)))))))(((((.(((.((((......)))).))).(((..((((((......))))))...))).....)))))....... (-17.99 = -19.30 + 1.31)



| Location | 6,876,683 – 6,876,778 |
|---|---|
| Length | 95 |
| Sequences | 4 |
| Columns | 98 |
| Reading direction | reverse |
| Mean pairwise identity | 81.17 |
| Mean single sequence MFE | -24.45 |
| Consensus MFE | -15.76 |
| Energy contribution | -15.95 |
| Covariance contribution | 0.19 |
| Combinations/Pair | 1.14 |
| Mean z-score | -1.85 |
| Structure conservation index | 0.64 |
| SVM decision value | 0.36 |
| SVM RNA-class probability | 0.704086 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 6876683 95 - 22224390 UUCAUUUCGACUUAACUGAAUAGGCAAAUUAAUUCACUUAGUGCUCGAGCAGCGC---AAUGCACAGCUUAGUCGCACUUCAGUUAAAUGAACUGGUU .(((.((((..(((((((((...((..(((((.....)))))(((.(.(((....---..))).))))......))..))))))))).)))).))).. ( -20.10) >DroSec_CAF1 1185 95 - 1 UUCAUCUCGACUUAAGUCAAUAUCCAAAUUAAUUCACUUAGUGCUGGAGCAGCGC---AAUGCACAGCUUAGUCGCACUUCAGUUAAAUGAAGUGGUU .......(((((.((((.....((((.(((((.....)))))..))))(((....---..)))...)))))))))(((((((......)))))))... ( -25.20) >DroSim_CAF1 1215 95 - 1 UUCAUCUCGACUUAAGUCAAUAGCUAAAUUAAUUCACUUAGUGCUGGAGCAGCGC---AAUGCACAGCUUAGUCGCACUUCAGUUAAAUGAAGUGGUU .......((((((((((.(((..........))).)))))..((((..(((....---..))).))))..)))))(((((((......)))))))... ( -24.30) >DroYak_CAF1 1347 96 - 1 GCCAUUUCGACUUAAGUCACGACAUUAGUCA-C-UAAUUUGUACUGUAGCAAUGCUCGAGUGCGCCGCUUAGUCGCAGUUUAGUUAAAUGAACUGGCU ((((.((((((....)))..(((....)))(-(-(((.((((((((.(((..(((......)))..))))))).)))).))))).....))).)))). ( -28.20) >consensus UUCAUCUCGACUUAAGUCAAUAGCCAAAUUAAUUCACUUAGUGCUGGAGCAGCGC___AAUGCACAGCUUAGUCGCACUUCAGUUAAAUGAACUGGUU .......(((((.((((.........................((((...))))((......))...)))))))))(((((((......)))))))... (-15.76 = -15.95 + 0.19)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:37:47 2006