
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 6,757,868 – 6,757,980 |
| Length | 112 |
| Max. P | 0.808230 |

| Location | 6,757,868 – 6,757,980 |
|---|---|
| Length | 112 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 79.27 |
| Mean single sequence MFE | -33.67 |
| Consensus MFE | -23.04 |
| Energy contribution | -24.49 |
| Covariance contribution | 1.45 |
| Combinations/Pair | 1.17 |
| Mean z-score | -1.80 |
| Structure conservation index | 0.68 |
| SVM decision value | 0.64 |
| SVM RNA-class probability | 0.808230 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
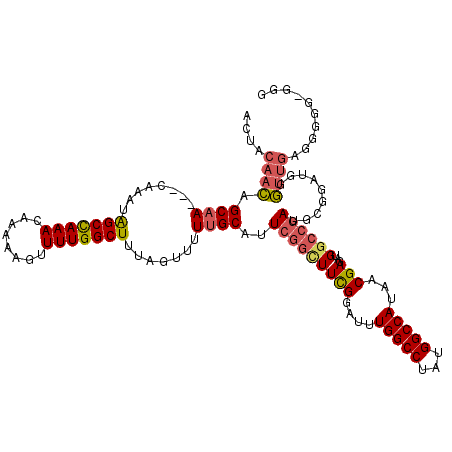
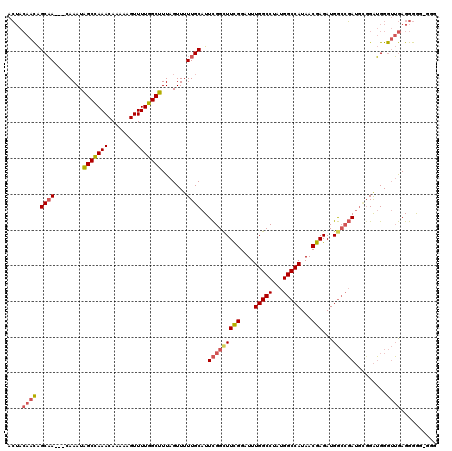
>X_DroMel_CAF1 6757868 112 + 22224390 ACUACAACAGCAA---CAAAUAGCCAAACAAAAAGUUUUGGCUUUAGUUUUUGCAUUCGGCUUCGGAUUUGGCCUAUGGCCAUAACGAGAUGGCCGAUUUCGAUGGGUUGAGGGA----- .((.((((.((((---.((..(((((((........)))))))....)).))))..(((((((((....(((((...)))))...)))...)))))).........)))).))..----- ( -36.60) >DroEre_CAF1 46575 117 + 1 ACUACAACAGCAA---CAAAUAGCCAAACAAAAAGUUUUGGCUUUAGUUUUUGCAUUCGGUUUCGGAUUUGGCCUAUGGCCAUAACGAGAUGGCCGAUGCGGAUGGGUUGAGGGGGAGGG .((.(..(..(((---(....(((((((........)))))))......(((((((.((((((((....(((((...)))))...))))...)))))))))))...))))..)..).)). ( -37.40) >DroAna_CAF1 53557 120 + 1 AGGAAAAUGGCCAAGCGAAAUGGCUAAACAAAAAGUUUUGGCUUUAGUUUUUGCAUUUGGCUUUGGAUUUGGCCUAAGGCCAUAACGAGAUGAGACAAGGGGGGGGAAGAAGGGGGUGGG ......((((((..((((((.(((((((........))))))).....))))))....((((........))))...))))))..................................... ( -27.00) >consensus ACUACAACAGCAA___CAAAUAGCCAAACAAAAAGUUUUGGCUUUAGUUUUUGCAUUCGGCUUCGGAUUUGGCCUAUGGCCAUAACGAGAUGGCCGAUGCGGAUGGGUUGAGGGGG_GGG ....((((.((((........(((((((........))))))).......))))..(((((((((....(((((...)))))...)))...)))))).........)))).......... (-23.04 = -24.49 + 1.45)

| Location | 6,757,868 – 6,757,980 |
|---|---|
| Length | 112 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 79.27 |
| Mean single sequence MFE | -25.77 |
| Consensus MFE | -15.09 |
| Energy contribution | -15.77 |
| Covariance contribution | 0.67 |
| Combinations/Pair | 1.15 |
| Mean z-score | -2.24 |
| Structure conservation index | 0.59 |
| SVM decision value | 0.62 |
| SVM RNA-class probability | 0.800759 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
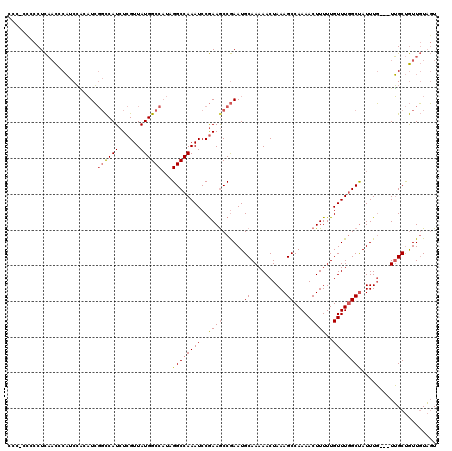
>X_DroMel_CAF1 6757868 112 - 22224390 -----UCCCUCAACCCAUCGAAAUCGGCCAUCUCGUUAUGGCCAUAGGCCAAAUCCGAAGCCGAAUGCAAAAACUAAAGCCAAAACUUUUUGUUUGGCUAUUUG---UUGCUGUUGUAGU -----...((((((.........(((((....(((...(((((...)))))....))).)))))..((((.......(((((((.(.....)))))))).....---)))).)))).)). ( -30.00) >DroEre_CAF1 46575 117 - 1 CCCUCCCCCUCAACCCAUCCGCAUCGGCCAUCUCGUUAUGGCCAUAGGCCAAAUCCGAAACCGAAUGCAAAAACUAAAGCCAAAACUUUUUGUUUGGCUAUUUG---UUGCUGUUGUAGU ........((((((......(((((((.....(((...(((((...)))))....)))..))).))))...(((...(((((((.(.....))))))))....)---))...)))).)). ( -27.70) >DroAna_CAF1 53557 120 - 1 CCCACCCCCUUCUUCCCCCCCCUUGUCUCAUCUCGUUAUGGCCUUAGGCCAAAUCCAAAGCCAAAUGCAAAAACUAAAGCCAAAACUUUUUGUUUAGCCAUUUCGCUUGGCCAUUUUCCU .....................................((((((...((......)).((((.(((((...((((.((((......))))..))))...))))).))))))))))...... ( -19.60) >consensus CCC_CCCCCUCAACCCAUCCACAUCGGCCAUCUCGUUAUGGCCAUAGGCCAAAUCCGAAGCCGAAUGCAAAAACUAAAGCCAAAACUUUUUGUUUGGCUAUUUG___UUGCUGUUGUAGU .........................((((((......))))))...((((((((..((((......((..........)).....))))..))))))))..................... (-15.09 = -15.77 + 0.67)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:37:04 2006