
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 6,687,473 – 6,687,624 |
| Length | 151 |
| Max. P | 0.648116 |

| Location | 6,687,473 – 6,687,590 |
|---|---|
| Length | 117 |
| Sequences | 5 |
| Columns | 119 |
| Reading direction | reverse |
| Mean pairwise identity | 89.97 |
| Mean single sequence MFE | -22.52 |
| Consensus MFE | -17.04 |
| Energy contribution | -16.80 |
| Covariance contribution | -0.24 |
| Combinations/Pair | 1.14 |
| Mean z-score | -1.68 |
| Structure conservation index | 0.76 |
| SVM decision value | 0.24 |
| SVM RNA-class probability | 0.648116 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 6687473 117 - 22224390 CGAUAUCUAUUUUUGGAUUUACCAAGGGUAAUUCCAUUAAUAAGCACUUAAUAGUUGGGCAAAUAAGCCGAAUACCAAGAAUGUAUAUUCACAUUGAAUGU--ACUAAUCAACUUAUAU .......((((..((((.(((((...))))).))))..))))..........((((((((......))).............((((((((.....))))))--))....)))))..... ( -27.60) >DroSec_CAF1 598 115 - 1 CGAUAUCUAUUUUUGGAUUCACCAAGGGUAAUUCCAUUAAUAAGCACCUAAUAGUUGGGCAAAUAAGCCGAAUACCAAAAAUGU--AUUCACAUUGUAUGU--ACUAAUCACCUAAUAU .(((...((((..((((...(((...)))...))))..)))).(((..((((.....(((......)))((((((.......))--))))..))))..)))--....)))......... ( -20.30) >DroSim_CAF1 634 115 - 1 CGAUAUCUAUUUUUGGAUUCACCAAGGGUAAUUCCAUUAAUAAGCACUUAAUAGUUGGGCAAAUAAGCCGAAUACCAAAAAUGU--AUUCACAUUGUGUGU--ACUAAUCACCCAAUAU .(((...((((..((((...(((...)))...))))..)))).((((.((((.....(((......)))((((((.......))--))))..)))).))))--....)))......... ( -23.60) >DroEre_CAF1 592 117 - 1 CGAUAUCUAUUUUUGGAUUCACCAAGGGUAAUUCCAUUAAUAAGUACUUAAUACUUGGGUAAAUAAGCCGAAUACCAAGAAUCU--AAUCGCAUUGUAUGUAUAUUUAUCACUUUAUAU .((((..(((..((((((((......((((.(((......((((((.....))))))(((......))))))))))..))))))--))..((((...)))))))..))))......... ( -20.00) >DroYak_CAF1 653 115 - 1 CGAUAUCUAUUUUUGGAUUCAACAAGGGUAAUUCCAUUAAUAAGUACUUAAUAGUUGGGUAAAUAAGCCGAAUACCCAGAAUGC--AUUCACAUUGUAUGU--AUUUAUGACUAAAUAU .((.(((((....))))))).....(((....)))....((((((((.......(((((((...........))))))).((((--(.......)))))))--)))))).......... ( -21.10) >consensus CGAUAUCUAUUUUUGGAUUCACCAAGGGUAAUUCCAUUAAUAAGCACUUAAUAGUUGGGCAAAUAAGCCGAAUACCAAGAAUGU__AUUCACAUUGUAUGU__ACUAAUCACCUAAUAU .((....(((((((((((((.....(((....)))......................(((......))))))).))))))))).....))............................. (-17.04 = -16.80 + -0.24)

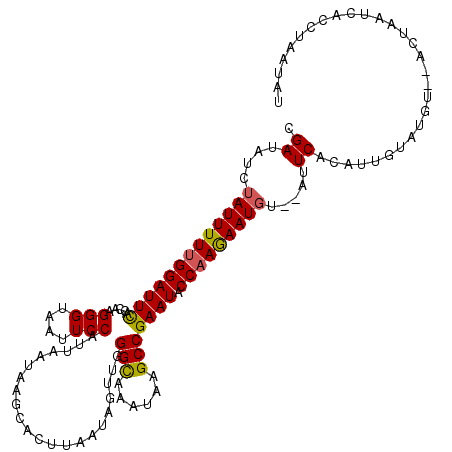

| Location | 6,687,511 – 6,687,624 |
|---|---|
| Length | 113 |
| Sequences | 5 |
| Columns | 119 |
| Reading direction | reverse |
| Mean pairwise identity | 91.07 |
| Mean single sequence MFE | -26.16 |
| Consensus MFE | -19.92 |
| Energy contribution | -19.84 |
| Covariance contribution | -0.08 |
| Combinations/Pair | 1.12 |
| Mean z-score | -1.52 |
| Structure conservation index | 0.76 |
| SVM decision value | -0.01 |
| SVM RNA-class probability | 0.526206 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 6687511 113 - 22224390 U------GCCUGUCUAAUGGAAGUCAUUGUCCAAUUUGUUCGAUAUCUAUUUUUGGAUUUACCAAGGGUAAUUCCAUUAAUAAGCACUUAAUAGUUGGGCAAAUAAGCCGAAUACCAAG (------((.(((.((((((((...((((..(.....)..))))...(((((((((.....))))))))).))))))))))).))).......(((.(((......))).)))...... ( -26.20) >DroSec_CAF1 634 113 - 1 U------GCCUGUCUAAUGGAAGCCAUUGUCCAAUUUGUUCGAUAUCUAUUUUUGGAUUCACCAAGGGUAAUUCCAUUAAUAAGCACCUAAUAGUUGGGCAAAUAAGCCGAAUACCAAA (------((.(((.((((((((...((((..(.....)..))))...(((((((((.....))))))))).))))))))))).))).......(((.(((......))).)))...... ( -26.20) >DroSim_CAF1 670 113 - 1 U------GCCUGUCUAAUGGAAGCCAUUGUCCAAUUUGUUCGAUAUCUAUUUUUGGAUUCACCAAGGGUAAUUCCAUUAAUAAGCACUUAAUAGUUGGGCAAAUAAGCCGAAUACCAAA (------((.(((.((((((((...((((..(.....)..))))...(((((((((.....))))))))).))))))))))).))).......(((.(((......))).)))...... ( -26.20) >DroEre_CAF1 630 119 - 1 UGUGUAUGUUUGUUUAAUGGAACCCAUUGUCCAAUUUGUUCGAUAUCUAUUUUUGGAUUCACCAAGGGUAAUUCCAUUAAUAAGUACUUAAUACUUGGGUAAAUAAGCCGAAUACCAAG ((.((((((((((((((((((((((((((..(.....)..))))........((((.....))))))))...)))))...((((((.....))))))..)))))))))...)))))).. ( -25.00) >DroYak_CAF1 689 113 - 1 U------GUCUGUCCAGUGGCAGGCAUUGUCCAACUUGUUCGAUAUCUAUUUUUGGAUUCAACAAGGGUAAUUCCAUUAAUAAGUACUUAAUAGUUGGGUAAAUAAGCCGAAUACCCAG (------(((((((....))))))))((..((((((((((.((.(((((....))))))))))))(((....)))..................)))))..))................. ( -27.20) >consensus U______GCCUGUCUAAUGGAAGCCAUUGUCCAAUUUGUUCGAUAUCUAUUUUUGGAUUCACCAAGGGUAAUUCCAUUAAUAAGCACUUAAUAGUUGGGCAAAUAAGCCGAAUACCAAG .......((.(((.((((((((...((((..(.....)..))))...(((((((((.....))))))))).))))))))))).))........(((.(((......))).)))...... (-19.92 = -19.84 + -0.08)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:36:43 2006