
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 6,668,288 – 6,668,380 |
| Length | 92 |
| Max. P | 0.847629 |

| Location | 6,668,288 – 6,668,380 |
|---|---|
| Length | 92 |
| Sequences | 3 |
| Columns | 92 |
| Reading direction | forward |
| Mean pairwise identity | 82.61 |
| Mean single sequence MFE | -32.43 |
| Consensus MFE | -24.55 |
| Energy contribution | -25.63 |
| Covariance contribution | 1.09 |
| Combinations/Pair | 1.07 |
| Mean z-score | -1.51 |
| Structure conservation index | 0.76 |
| SVM decision value | -0.05 |
| SVM RNA-class probability | 0.508123 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
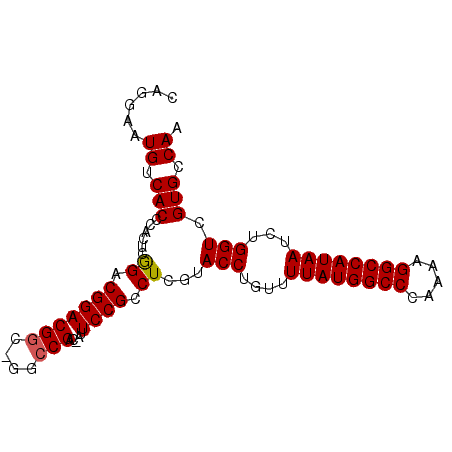
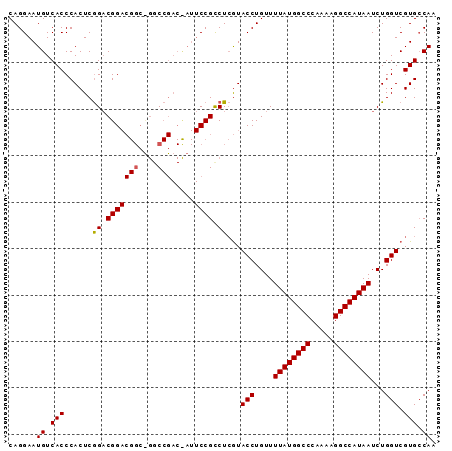
>X_DroMel_CAF1 6668288 92 + 22224390 UUGGCACGACCAGAUUAUGGCCUUUUGGGCCAUAAAACAGGUACGAGGCGGAAUAGUCGGCCGGCCGUCCGUCCGAGUGGGUGACAUUCCUG ...((.(((((...((((((((.....))))))))....))).))..))(((((.((((.((.((((......)).)).))))))))))).. ( -35.20) >DroSim_CAF1 173 92 + 1 UUGCCACGACCAGAUUAUGGCCUGUUGGGCCAUAAAACAGGUACGAGACGGAAUGGUCGGCCAGCCGUCCGUCCGAGUGGGUGACAUUCCUG ...((((.(((...((((((((.....))))))))....))).((.((((((.((((......)))))))))))).))))............ ( -36.50) >DroYak_CAF1 250 76 + 1 UUGGCACGACCAGAUUAUGGCCUUUUGGGCCAUAAAACAGGUAUCAGGCGG--------G--------CCGGCUGAGUGGGUGACAUUCCUG ((((.....)))).((((((((.....))))))))..((((..((((.((.--------.--------.)).))))...(....)...)))) ( -25.60) >consensus UUGGCACGACCAGAUUAUGGCCUUUUGGGCCAUAAAACAGGUACGAGGCGGAAU_GUCGGCC_GCCGUCCGUCCGAGUGGGUGACAUUCCUG ......(((((...((((((((.....))))))))....))).)).((((((.((((......)))))))))).(((((.....)))))... (-24.55 = -25.63 + 1.09)



| Location | 6,668,288 – 6,668,380 |
|---|---|
| Length | 92 |
| Sequences | 3 |
| Columns | 92 |
| Reading direction | reverse |
| Mean pairwise identity | 82.61 |
| Mean single sequence MFE | -30.50 |
| Consensus MFE | -21.07 |
| Energy contribution | -21.43 |
| Covariance contribution | 0.36 |
| Combinations/Pair | 1.04 |
| Mean z-score | -2.23 |
| Structure conservation index | 0.69 |
| SVM decision value | 0.78 |
| SVM RNA-class probability | 0.847629 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 6668288 92 - 22224390 CAGGAAUGUCACCCACUCGGACGGACGGCCGGCCGACUAUUCCGCCUCGUACCUGUUUUAUGGCCCAAAAGGCCAUAAUCUGGUCGUGCCAA ..((((((((..((....)).(((....)))...))).)))))((..((.(((....((((((((.....))))))))...))))).))... ( -32.30) >DroSim_CAF1 173 92 - 1 CAGGAAUGUCACCCACUCGGACGGACGGCUGGCCGACCAUUCCGUCUCGUACCUGUUUUAUGGCCCAACAGGCCAUAAUCUGGUCGUGGCAA ..((((((((..(((..((......))..)))..)).))))))(((.((.(((....((((((((.....))))))))...))))).))).. ( -37.30) >DroYak_CAF1 250 76 - 1 CAGGAAUGUCACCCACUCAGCCGG--------C--------CCGCCUGAUACCUGUUUUAUGGCCCAAAAGGCCAUAAUCUGGUCGUGCCAA ..((........))........((--------(--------.((((.(((........(((((((.....)))))))))).)).)).))).. ( -21.90) >consensus CAGGAAUGUCACCCACUCGGACGGACGGC_GGCCGAC_AUUCCGCCUCGUACCUGUUUUAUGGCCCAAAAGGCCAUAAUCUGGUCGUGCCAA ......((.(((......((.(((((((....))).....)))).))...(((....((((((((.....))))))))...))).))).)). (-21.07 = -21.43 + 0.36)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:36:31 2006