
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 6,591,279 – 6,591,432 |
| Length | 153 |
| Max. P | 0.761223 |

| Location | 6,591,279 – 6,591,392 |
|---|---|
| Length | 113 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 86.14 |
| Mean single sequence MFE | -37.50 |
| Consensus MFE | -28.59 |
| Energy contribution | -28.34 |
| Covariance contribution | -0.25 |
| Combinations/Pair | 1.11 |
| Mean z-score | -1.77 |
| Structure conservation index | 0.76 |
| SVM decision value | 0.50 |
| SVM RNA-class probability | 0.761223 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
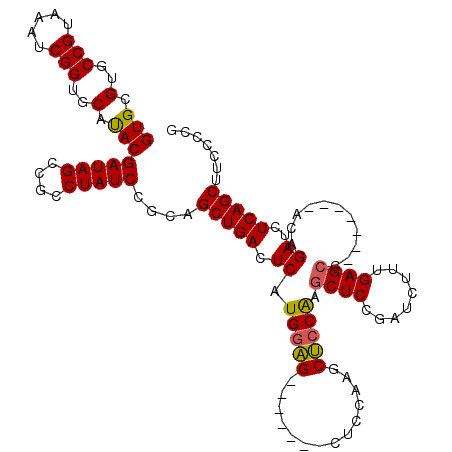
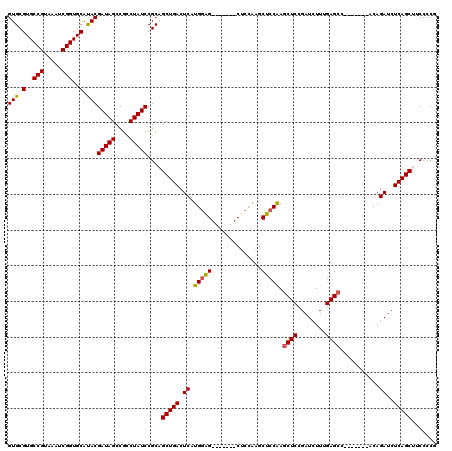
>X_DroMel_CAF1 6591279 113 - 22224390 GUGCGUGCCGUAAAUCGGUGCAUACGAUAGCCGCCUAUCCGCCGCUGACUCAUGGAGCUACAAGCUCCAAGCUCCAAGCUCCGAUGUUCGAGCC-------ACAGAUCUCAGCUUCCCCG (.(((..(((.....)))..)....(((((....))))).)))(((((.((.((((((.....)))))).((((..(((......))).)))).-------...))..)))))....... ( -37.00) >DroSec_CAF1 6953 106 - 1 GUGCGUGCCGUAAAUCGGUGCAUACGAUAGCCGCCUAUCCGCCGCUGACUCAUGCAG-------CUCCAAGCUCCAAGCUCCGAUCUUUGAGCC-------ACAGAUCUCAGCUUCCCCG (.(((..(((.....)))..)....(((((....))))).)))(((((....((..(-------(((.(((.((........)).))).)))).-------.))....)))))....... ( -30.00) >DroEre_CAF1 7093 113 - 1 GUGCGUGCCGUAAAUCGGUGCAUACGAUAGCCGUCUAUCCGCAGCUGACUCAUGGGG-------CUCAGAUCCCCGAUCUCCGAUCUUGGAGCCUCGGAUCACAGAUCUCAGCUGCCCCU (((.(..(((.....)))..).)))(((((....))))).((((((((....(((((-------((((((((..........)))))..))))))))((((...)))))))))))).... ( -44.70) >DroYak_CAF1 6209 106 - 1 GUGCGUGCCGUAAAUCGGUGCACACGAUAGCCGCCUAUCCGCAGCUGACUCAUGGAG-------CACCGAGCUCCAAGCUCCGAUCUUUGAGCC-------ACAGAUCUCAGCUUCCCCG (((.(..(((.....)))..).)))(((((....))))).(.((((((.((.(((((-------(.....)))))).((((........)))).-------...))..)))))).).... ( -38.30) >consensus GUGCGUGCCGUAAAUCGGUGCAUACGAUAGCCGCCUAUCCGCAGCUGACUCAUGGAG_______CUCCAAGCUCCAAGCUCCGAUCUUUGAGCC_______ACAGAUCUCAGCUUCCCCG (((.(..(((.....)))..).)))(((((....)))))....(((((.((.(((((..............))))).((((........))))...........))..)))))....... (-28.59 = -28.34 + -0.25)

| Location | 6,591,312 – 6,591,432 |
|---|---|
| Length | 120 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 82.32 |
| Mean single sequence MFE | -31.33 |
| Consensus MFE | -22.00 |
| Energy contribution | -21.56 |
| Covariance contribution | -0.44 |
| Combinations/Pair | 1.18 |
| Mean z-score | -1.67 |
| Structure conservation index | 0.70 |
| SVM decision value | 0.25 |
| SVM RNA-class probability | 0.651572 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
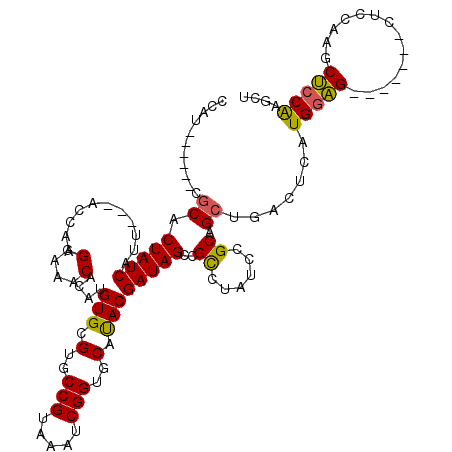
>X_DroMel_CAF1 6591312 120 - 22224390 CCAUCCAUCGCGCACUAUCAUUACUACCAAGGAAACACAUGUGCGUGCCGUAAAUCGGUGCAUACGAUAGCCGCCUAUCCGCCGCUGACUCAUGGAGCUACAAGCUCCAAGCUCCAAGCU .........(((..(((((...........(....)....(((.(..(((.....)))..).)))))))).))).........(((......((((((.....)))))).......))). ( -33.82) >DroSec_CAF1 6986 113 - 1 CCAUCCAUCGCACACUAUCAUUAAUACCAAGGAAACUCAUGUGCGUGCCGUAAAUCGGUGCAUACGAUAGCCGCCUAUCCGCCGCUGACUCAUGCAG-------CUCCAAGCUCCAAGCU .........((...(((((..........((....))...(((.(..(((.....)))..).))))))))..)).........((((.(....))))-------)....(((.....))) ( -25.10) >DroEre_CAF1 7133 104 - 1 CUAU------CGCACUAUCAUU---ACCAUGGAAACACAUGUGCGUGCCGUAAAUCGGUGCAUACGAUAGCCGUCUAUCCGCAGCUGACUCAUGGGG-------CUCAGAUCCCCGAUCU ..((------(((.........---..((((......))))((((..(((.....)))..)....(((((....))))).)))))........((((-------.......))))))).. ( -27.10) >DroYak_CAF1 6242 104 - 1 CUUC------GGCACUAUCAUU---ACCAAGGAAACUCGUGUGCGUGCCGUAAAUCGGUGCACACGAUAGCCGCCUAUCCGCAGCUGACUCAUGGAG-------CACCGAGCUCCAAGCU ..((------(((.(((((...---....((....)).((((((..((((.....)))))))))))))))..((......)).)))))....(((((-------(.....)))))).... ( -39.30) >consensus CCAU______CGCACUAUCAUU___ACCAAGGAAACACAUGUGCGUGCCGUAAAUCGGUGCAUACGAUAGCCGCCUAUCCGCAGCUGACUCAUGGAG_______CUCCAAGCUCCAAGCU ...........((.(((((...........(....)....(((.(..(((.....)))..).))))))))..((......)).)).......(((((..............))))).... (-22.00 = -21.56 + -0.44)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:35:40 2006