
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 6,578,468 – 6,578,585 |
| Length | 117 |
| Max. P | 0.587021 |

| Location | 6,578,468 – 6,578,585 |
|---|---|
| Length | 117 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 85.94 |
| Mean single sequence MFE | -31.75 |
| Consensus MFE | -20.98 |
| Energy contribution | -21.72 |
| Covariance contribution | 0.75 |
| Combinations/Pair | 1.21 |
| Mean z-score | -1.69 |
| Structure conservation index | 0.66 |
| SVM decision value | 0.11 |
| SVM RNA-class probability | 0.587021 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 6578468 117 + 22224390 UGGUAUCAUAUGAAAUGCAGGG-UAAAAUUUCAAGGUGUUUGGCCUGUAAUCCA--GCAUUAUUUGUAAAUUCGGUGGCUGGGUGCCGCCGAAUGAAAUGUAGAUCGGCUAAAAAUAAGU .((..((....))..((((((.-((((...(....)..)))).))))))..))(--((...(((((((.((((((((((.....))))))))))....)))))))..))).......... ( -31.20) >DroSec_CAF1 22956 119 + 1 UGGUAUCAUAUGAAAUGCAGUG-UAGAAUUUCAAGGUGUUCGCCCUGUAAUCCGGGGCCUUAUUUGUAAAUUCGGUGGCUGGUUGCUGCCGAAUGCAAUGUAGAUCGGCUAAAAAUAAGU ..........(((..((((.((-((((((((((((......((((((.....))))))....)))).))))))((..((.....))..))...)))).))))..)))............. ( -34.00) >DroEre_CAF1 22268 118 + 1 UAUUAUCAUAUAAAACACAGGGAGAAACUUUUUAGGUUUUUGCCCUAUAAUCCG--ACAAUAUUUGUAAAUUCGGUGGCUGGGUGCCGCCGACUGCAAUAUAGAUCGGCUAAAAAUAACU ..................(((((((((((.....))))))).)))).....(((--((.(((((.(((...((((((((.....)))))))).)))))))).).))))............ ( -33.90) >DroYak_CAF1 33489 117 + 1 UAUUAUCAUAUAAGACGCAGUG-GAGAAUUUCAAGGUGUUUGCCCUGUAAUCCG--GCAUUAUUUGUAAAUUCGGUGGCUGGCUGCCGUCGAAUGCAAUAUAGAUCGGCUAAAAAUAAGU .(((((......((((((..((-((....))))..)))))).....)))))..(--((...(((((((.(((((..(((.....)))..)))))....)))))))..))).......... ( -27.90) >consensus UAGUAUCAUAUAAAACGCAGGG_GAAAAUUUCAAGGUGUUUGCCCUGUAAUCCG__GCAUUAUUUGUAAAUUCGGUGGCUGGGUGCCGCCGAAUGCAAUAUAGAUCGGCUAAAAAUAAGU ................((((((...((((........)))).))))))........((...(((((((.((((((((((.....))))))))))....)))))))..))........... (-20.98 = -21.72 + 0.75)

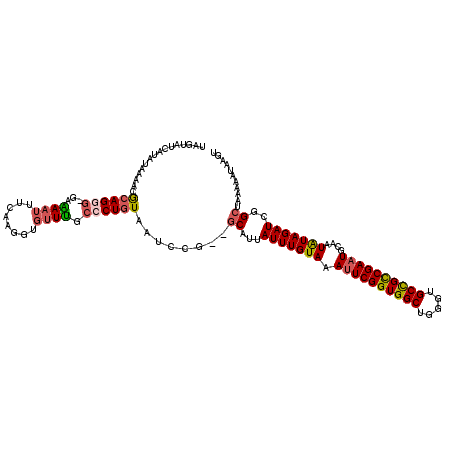

| Location | 6,578,468 – 6,578,585 |
|---|---|
| Length | 117 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 85.94 |
| Mean single sequence MFE | -25.08 |
| Consensus MFE | -15.93 |
| Energy contribution | -17.67 |
| Covariance contribution | 1.75 |
| Combinations/Pair | 1.14 |
| Mean z-score | -1.74 |
| Structure conservation index | 0.64 |
| SVM decision value | 0.00 |
| SVM RNA-class probability | 0.534595 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 6578468 117 - 22224390 ACUUAUUUUUAGCCGAUCUACAUUUCAUUCGGCGGCACCCAGCCACCGAAUUUACAAAUAAUGC--UGGAUUACAGGCCAAACACCUUGAAAUUUUA-CCCUGCAUUUCAUAUGAUACCA ...........(((((((((((((..((((((.(((.....))).))))))........)))).--))))))...)))....((...((((((.(..-....).))))))..))...... ( -23.10) >DroSec_CAF1 22956 119 - 1 ACUUAUUUUUAGCCGAUCUACAUUGCAUUCGGCAGCAACCAGCCACCGAAUUUACAAAUAAGGCCCCGGAUUACAGGGCGAACACCUUGAAAUUCUA-CACUGCAUUUCAUAUGAUACCA .(((((((.((((.(((....)))))((((((..((.....))..)))))).)).)))))))((((.(.....).))))...((...((((((.(..-....).))))))..))...... ( -24.30) >DroEre_CAF1 22268 118 - 1 AGUUAUUUUUAGCCGAUCUAUAUUGCAGUCGGCGGCACCCAGCCACCGAAUUUACAAAUAUUGU--CGGAUUAUAGGGCAAAAACCUAAAAAGUUUCUCCCUGUGUUUUAUAUGAUAAUA .((((((..((((((((..(((((....((((.(((.....))).)))).......))))).))--)))..(((((((...((((.......))))..)))))))..)))...)))))). ( -30.40) >DroYak_CAF1 33489 117 - 1 ACUUAUUUUUAGCCGAUCUAUAUUGCAUUCGACGGCAGCCAGCCACCGAAUUUACAAAUAAUGC--CGGAUUACAGGGCAAACACCUUGAAAUUCUC-CACUGCGUCUUAUAUGAUAAUA ...............(((((((.((((((((..(((.....)))..))))..............--.(((((.(((((......))))).)))))..-...))))...)))).))).... ( -22.50) >consensus ACUUAUUUUUAGCCGAUCUACAUUGCAUUCGGCGGCACCCAGCCACCGAAUUUACAAAUAAUGC__CGGAUUACAGGGCAAACACCUUGAAAUUCUA_CACUGCAUUUCAUAUGAUAACA ...............(((((((.(((((((((.(((.....))).))))).................(((((.(((((......))))).)))))......))))...)))).))).... (-15.93 = -17.67 + 1.75)

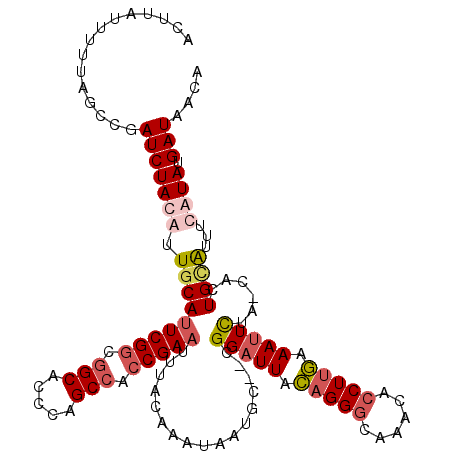

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:35:23 2006