
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 6,572,438 – 6,572,591 |
| Length | 153 |
| Max. P | 0.999806 |

| Location | 6,572,438 – 6,572,558 |
|---|---|
| Length | 120 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 96.24 |
| Mean single sequence MFE | -46.51 |
| Consensus MFE | -44.26 |
| Energy contribution | -44.82 |
| Covariance contribution | 0.56 |
| Combinations/Pair | 1.03 |
| Mean z-score | -2.41 |
| Structure conservation index | 0.95 |
| SVM decision value | 4.13 |
| SVM RNA-class probability | 0.999806 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 6572438 120 + 22224390 ACUUCAAGCGGUGGAAAGUAUCUGAGCGGCGUUCGACGGCCGCUUAAAUACACAGAAAUCGCAUCCGCGGAACGGGCGACAGUGGAACGCGGCUUAUCACCCGCCCACCGCCCCCCUCCC .......((((((....((((.((((((((........)))))))).))))...(....).)).))))(((..(((((...((((...((((........)))))))))))))...))). ( -46.10) >DroSec_CAF1 17275 115 + 1 ACUUCAAGCGGUGGAAAGUAUCUGAGCGGCGUUCGACGGCCGCUUAAAUACACAGAAAUCGCAUCCGCGGAACGAGCGGCAUUGGAACGCGGCUUAUCACCCGCCCACCGCCCCC----- .......(((((((........(((((.((((((.(..(((((((........(....)(((....)))....)))))))..).)))))).)))))........)))))))....----- ( -46.09) >DroSim_CAF1 27850 115 + 1 ACUUCAAGCGGUGGAAAGUAUCUGAGCGGCGUUCGACGGCCGCUUAAAUACACAGAAAUCGCAUCCGCGGAACGAGCGGCAUUGGAACGCGGCUUAUCACCCGCCCACCGCCCCC----- .......(((((((........(((((.((((((.(..(((((((........(....)(((....)))....)))))))..).)))))).)))))........)))))))....----- ( -46.09) >DroEre_CAF1 16915 115 + 1 ACUUCAAGCGGUGGAAAGUAUGUGAGCGACGUUCGACGGCCGCUUAAAUACACAGAAAUCGCAUCCGCGGAACGGGCGGCAGUGGAACGCGGCUUAUCACCCGCCCACCGCCCCC----- .......(((((((...(..(((((((..(((((.((.(((((((........(....)(((....)))....))))))).)).)))))..))))).))..)..)))))))....----- ( -45.50) >DroYak_CAF1 26824 115 + 1 ACUUCAAGCGGUGGAAAGUAUCUGAGCGGCGUUCGACGGCCGCUUAAAUACACAGAAAUCGCAUCCGCGGAACGGGCGGCAGUGGAACGCGGCUUAUCACCCGCCCACCGCCCCC----- .......(((((((........(((((.((((((.((.(((((((........(....)(((....)))....))))))).)).)))))).)))))........)))))))....----- ( -48.79) >consensus ACUUCAAGCGGUGGAAAGUAUCUGAGCGGCGUUCGACGGCCGCUUAAAUACACAGAAAUCGCAUCCGCGGAACGGGCGGCAGUGGAACGCGGCUUAUCACCCGCCCACCGCCCCC_____ .......(((((((........(((((.((((((.((.(((((((.............((((....))))...))))))).)).)))))).)))))........)))))))......... (-44.26 = -44.82 + 0.56)



| Location | 6,572,438 – 6,572,558 |
|---|---|
| Length | 120 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 96.24 |
| Mean single sequence MFE | -51.60 |
| Consensus MFE | -49.02 |
| Energy contribution | -49.10 |
| Covariance contribution | 0.08 |
| Combinations/Pair | 1.05 |
| Mean z-score | -1.96 |
| Structure conservation index | 0.95 |
| SVM decision value | 3.03 |
| SVM RNA-class probability | 0.998207 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 6572438 120 - 22224390 GGGAGGGGGGCGGUGGGCGGGUGAUAAGCCGCGUUCCACUGUCGCCCGUUCCGCGGAUGCGAUUUCUGUGUAUUUAAGCGGCCGUCGAACGCCGCUCAGAUACUUUCCACCGCUUGAAGU ((((.(((((((((((((((((.....))).)).))))))))).))).))))((((.((.((.......((((((.((((((........)))))).))))))..))))))))....... ( -56.60) >DroSec_CAF1 17275 115 - 1 -----GGGGGCGGUGGGCGGGUGAUAAGCCGCGUUCCAAUGCCGCUCGUUCCGCGGAUGCGAUUUCUGUGUAUUUAAGCGGCCGUCGAACGCCGCUCAGAUACUUUCCACCGCUUGAAGU -----..((((((((((.(((((...(((.((((((....((((((.....((((((.......))))))......))))))....)))))).)))....)))))))))))))))..... ( -49.70) >DroSim_CAF1 27850 115 - 1 -----GGGGGCGGUGGGCGGGUGAUAAGCCGCGUUCCAAUGCCGCUCGUUCCGCGGAUGCGAUUUCUGUGUAUUUAAGCGGCCGUCGAACGCCGCUCAGAUACUUUCCACCGCUUGAAGU -----..((((((((((.(((((...(((.((((((....((((((.....((((((.......))))))......))))))....)))))).)))....)))))))))))))))..... ( -49.70) >DroEre_CAF1 16915 115 - 1 -----GGGGGCGGUGGGCGGGUGAUAAGCCGCGUUCCACUGCCGCCCGUUCCGCGGAUGCGAUUUCUGUGUAUUUAAGCGGCCGUCGAACGUCGCUCACAUACUUUCCACCGCUUGAAGU -----..((((((((((.(((((...(((.((((((.((.(((((......((((((.......)))))).......))))).)).)))))).)))..))).)).))))))))))..... ( -49.72) >DroYak_CAF1 26824 115 - 1 -----GGGGGCGGUGGGCGGGUGAUAAGCCGCGUUCCACUGCCGCCCGUUCCGCGGAUGCGAUUUCUGUGUAUUUAAGCGGCCGUCGAACGCCGCUCAGAUACUUUCCACCGCUUGAAGU -----(((((((((((((((((.....))).)).))))))))).))).(((.((((.((.((.......((((((.((((((........)))))).))))))..))))))))..))).. ( -52.30) >consensus _____GGGGGCGGUGGGCGGGUGAUAAGCCGCGUUCCACUGCCGCCCGUUCCGCGGAUGCGAUUUCUGUGUAUUUAAGCGGCCGUCGAACGCCGCUCAGAUACUUUCCACCGCUUGAAGU .......((((((((((.(((((...(((.((((((.((.(((((......((((((.......)))))).......))))).)).)))))).)))....)))))))))))))))..... (-49.02 = -49.10 + 0.08)


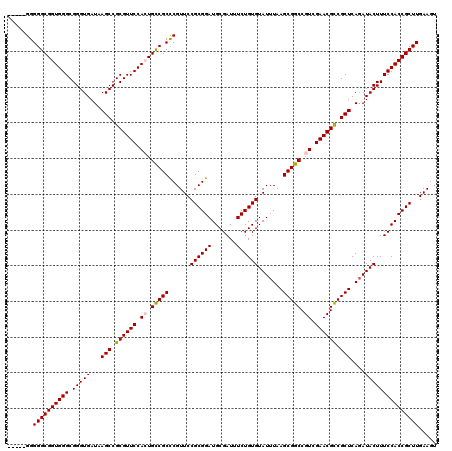
| Location | 6,572,478 – 6,572,591 |
|---|---|
| Length | 113 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 89.64 |
| Mean single sequence MFE | -42.08 |
| Consensus MFE | -40.20 |
| Energy contribution | -39.80 |
| Covariance contribution | -0.40 |
| Combinations/Pair | 1.12 |
| Mean z-score | -1.52 |
| Structure conservation index | 0.96 |
| SVM decision value | 1.36 |
| SVM RNA-class probability | 0.946671 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
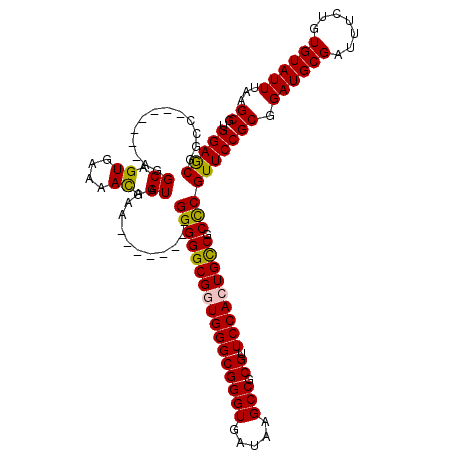
>X_DroMel_CAF1 6572478 113 - 22224390 UGGAGCGGCC-------AGGCAGUGAAAACGGUAAAAAGGGGGAGGGGGGCGGUGGGCGGGUGAUAAGCCGCGUUCCACUGUCGCCCGUUCCGCGGAUGCGAUUUCUGUGUAUUUAAGCG .(((((..((-------..((.((....)).))......))...(((.((((((((((((((.....))).)).))))))))).))))))))((.((((((.......))))))...)). ( -44.70) >DroSec_CAF1 17315 105 - 1 UGGAGCGGCC-------AAGCAGUGAAAACGGUAAAA--------GGGGGCGGUGGGCGGGUGAUAAGCCGCGUUCCAAUGCCGCUCGUUCCGCGGAUGCGAUUUCUGUGUAUUUAAGCG .((((((((.-------..((.((....)).))....--------...((((.(((((((((.....))).)).)))).)))))).))))))((.((((((.......))))))...)). ( -35.80) >DroSim_CAF1 27890 105 - 1 UGGAGCGGGC-------AAGCAGUGAAAAUGGUAAAA--------GGGGGCGGUGGGCGGGUGAUAAGCCGCGUUCCAAUGCCGCUCGUUCCGCGGAUGCGAUUUCUGUGUAUUUAAGCG .(((((((((-------..((.((....)).))....--------...((((.(((((((((.....))).)).)))).)))))))))))))((.((((((.......))))))...)). ( -39.30) >DroEre_CAF1 16955 105 - 1 UGGAACAGCG-------AGGCAGUGAAAACGGUAAAA--------GGGGGCGGUGGGCGGGUGAUAAGCCGCGUUCCACUGCCGCCCGUUCCGCGGAUGCGAUUUCUGUGUAUUUAAGCG .(((((....-------..((.((....)).))....--------(((((((((((((((((.....))).)).))))))))).))))))))((.((((((.......))))))...)). ( -43.80) >DroYak_CAF1 26864 112 - 1 UGGAACAGUGCGGCGUGCGGCAGUGAAAACGGUAAAA--------GGGGGCGGUGGGCGGGUGAUAAGCCGCGUUCCACUGCCGCCCGUUCCGCGGAUGCGAUUUCUGUGUAUUUAAGCG .(((((..(((.(....).)))((....)).......--------(((((((((((((((((.....))).)).))))))))).))))))))((.((((((.......))))))...)). ( -46.80) >consensus UGGAGCGGCC_______AGGCAGUGAAAACGGUAAAA________GGGGGCGGUGGGCGGGUGAUAAGCCGCGUUCCACUGCCGCCCGUUCCGCGGAUGCGAUUUCUGUGUAUUUAAGCG .(((((.............((.((....)).))............(((((((((((((((((.....))).)).))))))))).))))))))((.((((((.......))))))...)). (-40.20 = -39.80 + -0.40)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:35:14 2006