
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 6,549,570 – 6,549,730 |
| Length | 160 |
| Max. P | 0.870119 |

| Location | 6,549,570 – 6,549,690 |
|---|---|
| Length | 120 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 92.22 |
| Mean single sequence MFE | -16.94 |
| Consensus MFE | -16.55 |
| Energy contribution | -16.33 |
| Covariance contribution | -0.22 |
| Combinations/Pair | 1.05 |
| Mean z-score | -1.74 |
| Structure conservation index | 0.98 |
| SVM decision value | 0.87 |
| SVM RNA-class probability | 0.870119 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 6549570 120 - 22224390 AUAUGCAGCAAACUUGCCCAUAAAAGCAUCAAAGAUGCAGCGCCCUCUCUUCACAUCUUACCAACAGCGUUGUAAAAAGGAAAAAAAAAACACACACACACAGAUGUGUUCAACAUAAUA .((((..(((....)))((......(((((...)))))(((((.......................))))).......))........(((((((.......).))))))...))))... ( -15.80) >DroSim_CAF1 4574 118 - 1 AUAUGCAGCAAACUUGCCCAUAAAAGCAUCAAAGAUGCAGCGCCCUCUCUUCACAUCUUCCCAACAGCGUUGCAAAAA--AAAAAAAACACACACACACACAGAUGUGUUCAACAUAAUA ...((((((................(((((...))))).((.........................))))))))....--.............(((((......)))))........... ( -17.11) >DroYak_CAF1 4595 118 - 1 AUAUGCAGCAAACUUGCCCAUAAAAGCAUCAAAGAUGCAGCGCCCUCUUUUCACAUCUUCCCAACUGCGUUGCAAAAACCAAA--AAAAAAAAACACAAACAGAUGUGUUCAACAUACUA .((((..(((....))).)))).............((((((((.......................)))))))).........--.......((((((......)))))).......... ( -17.90) >consensus AUAUGCAGCAAACUUGCCCAUAAAAGCAUCAAAGAUGCAGCGCCCUCUCUUCACAUCUUCCCAACAGCGUUGCAAAAA__AAAAAAAAAACACACACACACAGAUGUGUUCAACAUAAUA .((((..(((....))).)))).............((((((((.......................))))))))...................(((((......)))))........... (-16.55 = -16.33 + -0.22)

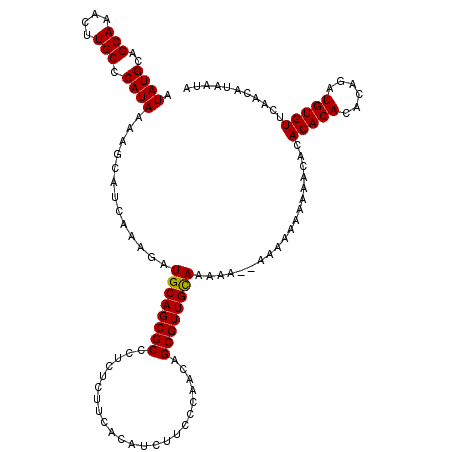

| Location | 6,549,610 – 6,549,730 |
|---|---|
| Length | 120 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 96.11 |
| Mean single sequence MFE | -39.00 |
| Consensus MFE | -37.13 |
| Energy contribution | -37.13 |
| Covariance contribution | -0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.98 |
| Structure conservation index | 0.95 |
| SVM decision value | 0.78 |
| SVM RNA-class probability | 0.849852 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 6549610 120 + 22224390 CCUUUUUACAACGCUGUUGGUAAGAUGUGAAGAGAGGGCGCUGCAUCUUUGAUGCUUUUAUGGGCAAGUUUGCUGCAUAUGAAACUUGUCUAUUGAGCGCGCCUGGUUCUUUCUUCUGGC ((.(((((((((...))).))))))...(((((((((((((.(((((...)))))(((.((((((((((((.(.......))))))))))))).))).))))).....)))))))).)). ( -37.80) >DroSim_CAF1 4614 118 + 1 --UUUUUGCAACGCUGUUGGGAAGAUGUGAAGAGAGGGCGCUGCAUCUUUGAUGCUUUUAUGGGCAAGUUUGCUGCAUAUGAAACUUGUCUAUUGAGCGCGCCUGGUUCUUUCUUCUGGC --(((((.((((...)))).)))))...(((((((((((((.(((((...)))))(((.((((((((((((.(.......))))))))))))).))).))))).....)))))))).... ( -39.90) >DroYak_CAF1 4633 120 + 1 GGUUUUUGCAACGCAGUUGGGAAGAUGUGAAAAGAGGGCGCUGCAUCUUUGAUGCUUUUAUGGGCAAGUUUGCUGCAUAUGAAACUUGUCUAUUGAGCGCGCCUGGUUCUUUCUUCUGGC ........((((...))))((((((...(((....((((((.(((((...)))))(((.((((((((((((.(.......))))))))))))).))).))))))..)))..))))))... ( -39.30) >consensus __UUUUUGCAACGCUGUUGGGAAGAUGUGAAGAGAGGGCGCUGCAUCUUUGAUGCUUUUAUGGGCAAGUUUGCUGCAUAUGAAACUUGUCUAUUGAGCGCGCCUGGUUCUUUCUUCUGGC ...............((..(.((((...(((....((((((.(((((...)))))(((.((((((((((((.(.......))))))))))))).))).))))))..)))..)))))..)) (-37.13 = -37.13 + -0.00)


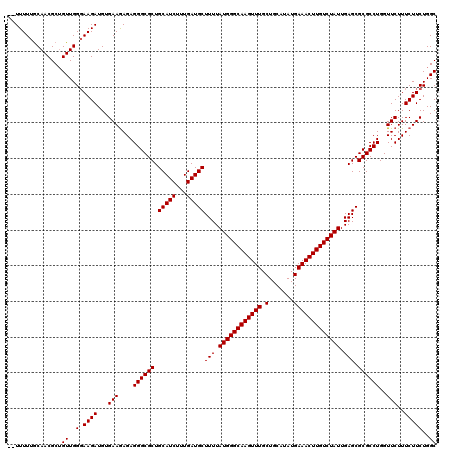
| Location | 6,549,610 – 6,549,730 |
|---|---|
| Length | 120 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 96.11 |
| Mean single sequence MFE | -26.57 |
| Consensus MFE | -24.93 |
| Energy contribution | -25.27 |
| Covariance contribution | 0.33 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.65 |
| Structure conservation index | 0.94 |
| SVM decision value | 0.14 |
| SVM RNA-class probability | 0.602763 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 6549610 120 - 22224390 GCCAGAAGAAAGAACCAGGCGCGCUCAAUAGACAAGUUUCAUAUGCAGCAAACUUGCCCAUAAAAGCAUCAAAGAUGCAGCGCCCUCUCUUCACAUCUUACCAACAGCGUUGUAAAAAGG .((.(((((.((.....(((((..........((((((((.......).))))))).........(((((...))))).))))))).))))).....((((.(((...)))))))...)) ( -26.60) >DroSim_CAF1 4614 118 - 1 GCCAGAAGAAAGAACCAGGCGCGCUCAAUAGACAAGUUUCAUAUGCAGCAAACUUGCCCAUAAAAGCAUCAAAGAUGCAGCGCCCUCUCUUCACAUCUUCCCAACAGCGUUGCAAAAA-- ....(((((((((...(((.(((((.......((((((((.......).))))))).........(((((...))))))))))))).))))....))))).((((...))))......-- ( -26.60) >DroYak_CAF1 4633 120 - 1 GCCAGAAGAAAGAACCAGGCGCGCUCAAUAGACAAGUUUCAUAUGCAGCAAACUUGCCCAUAAAAGCAUCAAAGAUGCAGCGCCCUCUUUUCACAUCUUCCCAACUGCGUUGCAAAAACC ....(((((..(((..(((.(((((.......((((((((.......).))))))).........(((((...)))))))))))))...)))...))))).((((...))))........ ( -26.50) >consensus GCCAGAAGAAAGAACCAGGCGCGCUCAAUAGACAAGUUUCAUAUGCAGCAAACUUGCCCAUAAAAGCAUCAAAGAUGCAGCGCCCUCUCUUCACAUCUUCCCAACAGCGUUGCAAAAA__ ....(((((..(((..(((.(((((.......((((((((.......).))))))).........(((((...)))))))))))))...)))...))))).((((...))))........ (-24.93 = -25.27 + 0.33)

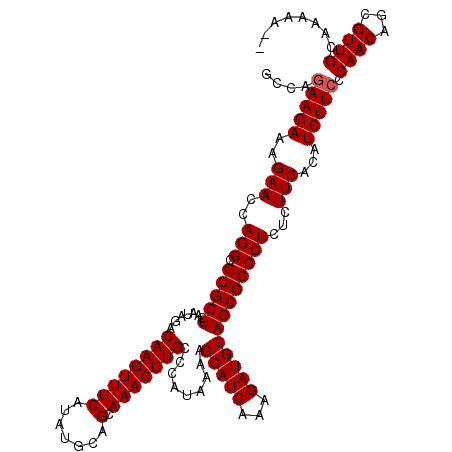

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:34:32 2006