
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 6,499,855 – 6,499,950 |
| Length | 95 |
| Max. P | 0.995578 |

| Location | 6,499,855 – 6,499,950 |
|---|---|
| Length | 95 |
| Sequences | 5 |
| Columns | 98 |
| Reading direction | forward |
| Mean pairwise identity | 94.09 |
| Mean single sequence MFE | -30.08 |
| Consensus MFE | -24.80 |
| Energy contribution | -24.80 |
| Covariance contribution | 0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.67 |
| Structure conservation index | 0.82 |
| SVM decision value | 2.59 |
| SVM RNA-class probability | 0.995578 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 6499855 95 + 22224390 GA--UCCAUAAUCUCGAAAAAACCAUCG-AUGGCAUCGAACCGGCUGGAACUGAACCAUCGAUAUUAUCAGUAGCGCCAUCGAUGGUUUGAUAGCACC ..--...............(((((((((-(((((..((...))(((...(((((.............))))))))))))))))))))))......... ( -28.92) >DroSec_CAF1 53783 95 + 1 GU--CCCAUUGUCUCGAAAAAACCAUCG-AUGGCAUCGAACCGGCUGGAACUGAACCAUCGAUAUUAUCAGUAGUGCCAUCGAUGGUUUGAUAGCACA ..--.....(((((.....(((((((((-(((((((....((....)).(((((.............))))).))))))))))))))))...)).))) ( -30.72) >DroSim_CAF1 53122 95 + 1 GU--CCCAUAGUCUCGAAAAAACCAUCG-AUGGCAUCGAACCGGCUGGAACUGAACCAUCGAUAUUAUCAGUAGUGCCAUCGAUGGUUUGAUAGCACC ..--...............(((((((((-(((((((....((....)).(((((.............))))).))))))))))))))))......... ( -30.12) >DroEre_CAF1 53385 98 + 1 GCGACCCAUAGUCUCGAAAAAACCAUCGGAUGGCAUCGAACCGGCUGGAACUGAACCGUCGAUAGUAUCAGUAGCGCCAUCGAUGGUUUGAUAGCACC (((((.....)))......(((((((((.(((((..((...))(((...(((((..(.......)..))))))))))))))))))))))....))... ( -30.30) >DroYak_CAF1 53487 96 + 1 GA-CCCCAUAGUCUCGAAAAAACCAUCG-AUGGCAUCGAACCGGCUGGAACUGAACCAUCGAUAUUAUCAGUAGCGCCAUCGAUGGUUUGAUACCACC ((-(......)))......(((((((((-(((((..((...))(((...(((((.............))))))))))))))))))))))......... ( -30.32) >consensus GA__CCCAUAGUCUCGAAAAAACCAUCG_AUGGCAUCGAACCGGCUGGAACUGAACCAUCGAUAUUAUCAGUAGCGCCAUCGAUGGUUUGAUAGCACC ...................(((((((((.(((((......((....)).(((((.............)))))...))))))))))))))......... (-24.80 = -24.80 + 0.00)


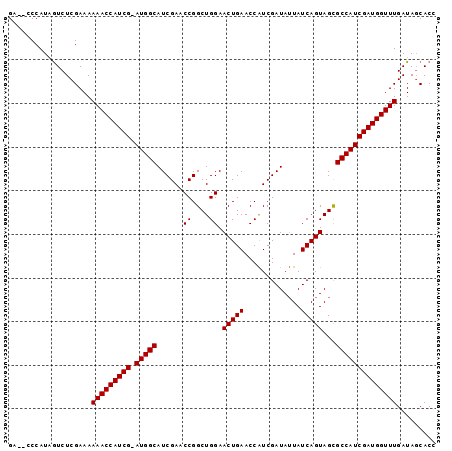
| Location | 6,499,855 – 6,499,950 |
|---|---|
| Length | 95 |
| Sequences | 5 |
| Columns | 98 |
| Reading direction | reverse |
| Mean pairwise identity | 94.09 |
| Mean single sequence MFE | -31.80 |
| Consensus MFE | -26.60 |
| Energy contribution | -26.16 |
| Covariance contribution | -0.44 |
| Combinations/Pair | 1.10 |
| Mean z-score | -2.11 |
| Structure conservation index | 0.84 |
| SVM decision value | 1.94 |
| SVM RNA-class probability | 0.983382 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 6499855 95 - 22224390 GGUGCUAUCAAACCAUCGAUGGCGCUACUGAUAAUAUCGAUGGUUCAGUUCCAGCCGGUUCGAUGCCAU-CGAUGGUUUUUUCGAGAUUAUGGA--UC (((((((((........))))))))).((((...((((((((((...(..((....))..)...)))))-)))))......)).))........--.. ( -30.10) >DroSec_CAF1 53783 95 - 1 UGUGCUAUCAAACCAUCGAUGGCACUACUGAUAAUAUCGAUGGUUCAGUUCCAGCCGGUUCGAUGCCAU-CGAUGGUUUUUUCGAGACAAUGGG--AC (((.((...((((((((((((((.(....).....((((((((((.......)))))..))))))))))-))))))))).....))))).....--.. ( -31.30) >DroSim_CAF1 53122 95 - 1 GGUGCUAUCAAACCAUCGAUGGCACUACUGAUAAUAUCGAUGGUUCAGUUCCAGCCGGUUCGAUGCCAU-CGAUGGUUUUUUCGAGACUAUGGG--AC (((((((((........))))))))).((.(((.((((((((((...(..((....))..)...)))))-)))))(((((...)))))))).))--.. ( -32.90) >DroEre_CAF1 53385 98 - 1 GGUGCUAUCAAACCAUCGAUGGCGCUACUGAUACUAUCGACGGUUCAGUUCCAGCCGGUUCGAUGCCAUCCGAUGGUUUUUUCGAGACUAUGGGUCGC (((((((((........))))))))).........((((((((((.......)))))..)))))((.((((.((((((((...)))))))))))).)) ( -30.70) >DroYak_CAF1 53487 96 - 1 GGUGGUAUCAAACCAUCGAUGGCGCUACUGAUAAUAUCGAUGGUUCAGUUCCAGCCGGUUCGAUGCCAU-CGAUGGUUUUUUCGAGACUAUGGGG-UC (((((((((.(((((((((((...(....)....)))))))))))..(..((....))..)))))))))-).((((((((...))))))))....-.. ( -34.00) >consensus GGUGCUAUCAAACCAUCGAUGGCGCUACUGAUAAUAUCGAUGGUUCAGUUCCAGCCGGUUCGAUGCCAU_CGAUGGUUUUUUCGAGACUAUGGG__AC (((.((...((((((((((((((.(....).....((((((((((.......)))))..)))))))))).))))))))).....)))))......... (-26.60 = -26.16 + -0.44)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:34:11 2006